

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147363.1 - phase: 0 /pseudo
(427 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
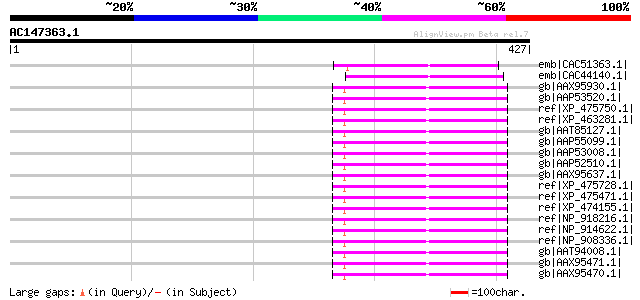
Score E
Sequences producing significant alignments: (bits) Value
emb|CAC51363.1| putative polyprotein [Cicer arietinum] 89 3e-16
emb|CAC44140.1| putative polyprotein [Cicer arietinum] 87 1e-15
gb|AAX95930.1| retrotransposon protein, putative, Ty3-gypsy sub-... 72 4e-11
gb|AAP53520.1| Similar to Sorghum bicolor 22 kDakafirinclusterpo... 70 2e-10
ref|XP_475750.1| putative polyprotein [Oryza sativa (japonica cu... 69 2e-10
ref|XP_463281.1| putative polyprotein [Oryza sativa (japonica cu... 69 2e-10
gb|AAT85127.1| putative polyprotein [Oryza sativa (japonica cult... 69 2e-10
gb|AAP55099.1| putative polyprotein [Oryza sativa (japonica cult... 69 3e-10
gb|AAP53008.1| putative polyprotein [Oryza sativa (japonica cult... 69 3e-10
gb|AAP52510.1| putative polyprotein [Oryza sativa (japonica cult... 69 3e-10
gb|AAX95637.1| Reverse transcriptase (RNA-dependent DNA polymera... 69 3e-10
ref|XP_475728.1| putative polyprotein [Oryza sativa (japonica cu... 69 3e-10
ref|XP_475471.1| putative polyprotein [Oryza sativa (japonica cu... 69 3e-10
ref|XP_474155.1| OSJNBa0060D06.14 [Oryza sativa (japonica cultiv... 69 3e-10
ref|NP_918216.1| putative Sorghum bicolor 22 kDa kafirin cluster... 69 3e-10
ref|NP_914622.1| similar to polyprotein [Oryza sativa (japonica ... 69 3e-10
ref|NP_908336.1| putative polyprotein [Oryza sativa (japonica cu... 69 3e-10
gb|AAT94008.1| putative polyprotein [Oryza sativa (japonica cult... 69 3e-10
gb|AAX95471.1| Retrotransposon gag protein, putative [Oryza sati... 69 3e-10
gb|AAX95470.1| Retrotransposon gag protein, putative [Oryza sati... 69 3e-10
>emb|CAC51363.1| putative polyprotein [Cicer arietinum]
Length = 318
Score = 88.6 bits (218), Expect = 3e-16
Identities = 50/147 (34%), Positives = 76/147 (51%), Gaps = 12/147 (8%)
Query: 267 CKQPKKAPTI-----------GRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATH 315
C PK P + GR++ + + LI C I L A+ D GATH
Sbjct: 108 CTAPKVEPVVNVARATRPTARGRIYCMNAEEGNPSGNLIQRDCEIAGNTLTALFDSGATH 167
Query: 316 CFIAFDCASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLS 375
FIA DC + L L +S + ++VV TP K ++T + CL +++ RD ++L+CLPL
Sbjct: 168 SFIAMDCVNRLKLSVSALPFDLVVSTPDK-TLTVNSACLHCRMTIQNRDLLVNLICLPLQ 226
Query: 376 GMYVILGMNWFE*NHVHINYFSKSVYF 402
+ VILGM+W ++V ++ K V F
Sbjct: 227 SLEVILGMDWMSYHYVILDCARKLVIF 253
>emb|CAC44140.1| putative polyprotein [Cicer arietinum]
Length = 318
Score = 87.0 bits (214), Expect = 1e-15
Identities = 47/130 (36%), Positives = 72/130 (55%), Gaps = 1/130 (0%)
Query: 277 GRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDCASTLGLDMSNMNGE 336
GR++ + + LI C I L A+ D GATH FIA C + L L +S + +
Sbjct: 129 GRIYCVNAEEGNPSGNLIQRDCEIAGNTLTALFDSGATHSFIAMGCVNRLKLSVSALPFD 188
Query: 337 MVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILGMNWFE*NHVHINYF 396
+VV TPAK ++T + CL +++ RDF ++L+CLPL + VILGM+W ++V ++
Sbjct: 189 LVVSTPAK-TLTVNSACLHCRMTIQNRDFLVNLICLPLQSLEVILGMDWMSYHYVILDCA 247
Query: 397 SKSVYFSSVE 406
V F E
Sbjct: 248 RTLVLFPEPE 257
>gb|AAX95930.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 1169
Score = 71.6 bits (174), Expect = 4e-11
Identities = 44/147 (29%), Positives = 78/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 436 KCPKPRRAGPKFVQARVNHASAEEAQSATEVILGTFPVNSTPAVILFDSGATHSFISKRF 495
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 496 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 554
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ +EE+
Sbjct: 555 MDWLTRHRGVIDCASRTIKLTNAKEEV 581
>gb|AAP53520.1| Similar to Sorghum bicolor 22 kDakafirinclusterpolyprotein [Oryza
sativa (japonica cultivar-group)]
gi|37533862|ref|NP_921233.1| Similar to Sorghum bicolor
22 kDakafirinclusterpolyprotein [Oryza sativa (japonica
cultivar-group)] gi|13129427|gb|AAK13085.1| Similar to
Sorghum bicolor 22 kDakafirinclusterpolyprotein [Oryza
sativa]
Length = 1484
Score = 69.7 bits (169), Expect = 2e-10
Identities = 43/147 (29%), Positives = 78/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C++P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 343 KCQKPRRAGPKFVQARVNHASAKEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 402
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 403 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 461
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 462 MDWLTRHRGVIDCASRTIELTNAKGEV 488
>ref|XP_475750.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|48843822|gb|AAT47081.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 69.3 bits (168), Expect = 2e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGSIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>ref|XP_463281.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1455
Score = 69.3 bits (168), Expect = 2e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCSKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>gb|AAT85127.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 69.3 bits (168), Expect = 2e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGSIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>gb|AAP55099.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37537020|ref|NP_922812.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|19225016|gb|AAL86492.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>gb|AAP53008.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37532838|ref|NP_920721.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|16905208|gb|AAL31078.1| putative polyprotein [Oryza
sativa]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>gb|AAP52510.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37531842|ref|NP_920223.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|22725995|gb|AAN04995.1| retrotransposon protein,
putative, Ty3-gypsy sub-class [Oryza sativa (japonica
cultivar-group)]
Length = 1413
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 342 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 401
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 402 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 460
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 461 MDWLARHRGVIDCASRTIKLTNAKGEV 487
>gb|AAX95637.1| Reverse transcriptase (RNA-dependent DNA polymerase), putative
[Oryza sativa (japonica cultivar-group)]
gi|50918021|ref|XP_469407.1| putative gag-pol
polyprotein [Oryza sativa (japonica cultivar-group)]
gi|28273360|gb|AAO38446.1| putative gag-pol polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 1229
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 75 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 134
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 135 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 193
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 194 MDWLARHRGVIDCASRTIKLTNAKGEV 220
>ref|XP_475728.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50080333|gb|AAT69667.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>ref|XP_475471.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50080316|gb|AAT69650.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1458
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>ref|XP_474155.1| OSJNBa0060D06.14 [Oryza sativa (japonica cultivar-group)]
gi|38345904|emb|CAE03548.2| OSJNBa0060D06.14 [Oryza
sativa (japonica cultivar-group)]
Length = 1458
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>ref|NP_918216.1| putative Sorghum bicolor 22 kDa kafirin cluster polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>ref|NP_914622.1| similar to polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLARHRGVIDCASRTIKLTNAKGEV 515
>ref|NP_908336.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|52353682|gb|AAU44248.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|52353613|gb|AAU44179.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|20804440|dbj|BAB92137.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|15128451|dbj|BAB62635.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1524
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 370 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 429
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 430 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 488
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 489 MDWLTRHRGVIDCASRTIKLTNAKGEV 515
>gb|AAT94008.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|51038165|gb|AAT93968.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1500
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 346 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 405
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 406 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 464
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 465 MDWLTRHRGVIDCASRTIKLTNAKGEV 491
>gb|AAX95471.1| Retrotransposon gag protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1441
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 371 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 430
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 431 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 489
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 490 MDWLTRHRGVIDCASRTIKLTNAKGEV 516
>gb|AAX95470.1| Retrotransposon gag protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1441
Score = 68.9 bits (167), Expect = 3e-10
Identities = 43/147 (29%), Positives = 77/147 (52%), Gaps = 4/147 (2%)
Query: 266 QCKQPKKAP---TIGRVFALTGTQTENEDRLIIGTCYINNTPLVAIIDIGATHCFIAFDC 322
+C +P++A RV + + ++ +I+GT +N+TP V + D GATH FI+
Sbjct: 371 KCPKPRRAGPKFVQARVNHASAEEAQSAPEVILGTFPVNSTPAVILFDSGATHSFISKRF 430
Query: 323 ASTLGLDMSNMNGEMVVETPAKGSVTTSLVCLKSSLSMFGRDFEMDLVCLPLSGMYVILG 382
A GL + + M V TP G +TT+ C ++ + G F +L+ L + VILG
Sbjct: 431 AGAHGLSLVKLKIPMRVHTPG-GGMTTTHYCPSVTVEIQGLIFPANLILLESKDLDVILG 489
Query: 383 MNWFE*NHVHINYFSKSVYFSSVEEEI 409
M+W + I+ S+++ ++ + E+
Sbjct: 490 MDWLTRHRGVIDCASRTIKLTNAKGEV 516
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.343 0.151 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 595,817,441
Number of Sequences: 2540612
Number of extensions: 21896417
Number of successful extensions: 71556
Number of sequences better than 10.0: 423
Number of HSP's better than 10.0 without gapping: 352
Number of HSP's successfully gapped in prelim test: 71
Number of HSP's that attempted gapping in prelim test: 70803
Number of HSP's gapped (non-prelim): 437
length of query: 427
length of database: 863,360,394
effective HSP length: 131
effective length of query: 296
effective length of database: 530,540,222
effective search space: 157039905712
effective search space used: 157039905712
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 76 (33.9 bits)
Medicago: description of AC147363.1


