

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.9 + phase: 1 /partial
(375 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
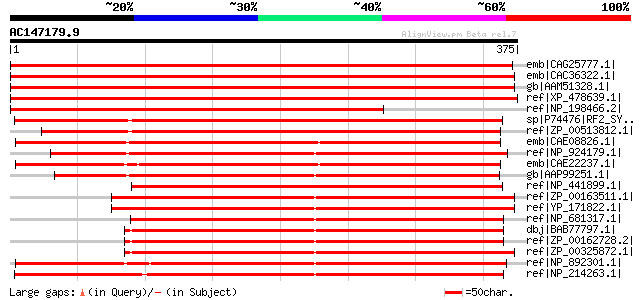
Score E
Sequences producing significant alignments: (bits) Value
emb|CAG25777.1| putative translation releasing factor 2 [Cucumis... 625 e-178
emb|CAC36322.1| translation releasing factor2 [Arabidopsis thali... 613 e-174
gb|AAM51328.1| putative translation releasing factor RF-2 [Arabi... 613 e-174
ref|XP_478639.1| putative translation releasing factor2 [Oryza s... 595 e-169
ref|NP_198466.2| peptide chain release factor, putative [Arabido... 448 e-124
sp|P74476|RF2_SYNY3 Peptide chain release factor 2 (RF-2) 426 e-118
ref|ZP_00513812.1| Peptide chain release factor 2 [Crocosphaera ... 411 e-113
emb|CAE08826.1| peptide chain release factor RF-2 [Synechococcus... 395 e-108
ref|NP_924179.1| peptide chain release factor [Gloeobacter viola... 393 e-108
emb|CAE22237.1| peptide-chain-release factor RF-2 [Prochlorococc... 391 e-107
gb|AAP99251.1| Protein chain release factor B [Prochlorococcus m... 390 e-107
ref|NP_441899.1| peptide chain release factor [Synechocystis sp.... 386 e-106
ref|ZP_00163511.1| COG1186: Protein chain release factor B [Syne... 384 e-105
ref|YP_171822.1| peptide chain release factor RF-2 [Synechococcu... 382 e-105
ref|NP_681317.1| peptide chain release factor 2 [Thermosynechoco... 379 e-103
dbj|BAB77797.1| peptide chain release factor [Nostoc sp. PCC 712... 378 e-103
ref|ZP_00162728.2| COG1186: Protein chain release factor B [Anab... 377 e-103
ref|ZP_00325872.1| COG1186: Protein chain release factor B [Tric... 377 e-103
ref|NP_892301.1| peptide chain release factor RF-2 [Prochlorococ... 367 e-100
ref|NP_214263.1| peptide chain release factor RF-2 [Aquifex aeol... 327 4e-88
>emb|CAG25777.1| putative translation releasing factor 2 [Cucumis sativus]
Length = 453
Score = 625 bits (1612), Expect = e-178
Identities = 309/372 (83%), Positives = 349/372 (93%)
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
FY+LRK+VE S+RV+EIR S+GL L+QELA LE +A+ +SFWDDR+KAQ+ L + DV
Sbjct: 80 FYTLRKEVETTSERVEEIRNSAGLLQLDQELADLESKAADNSFWDDRSKAQKVLMAMTDV 139
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSG 120
K+KIK+L D+KTQVE+AETIV LTEEM+SVD GL EEA+ +IK+LNK++D+FEL++LLSG
Sbjct: 140 KDKIKMLTDFKTQVEEAETIVKLTEEMDSVDVGLLEEATKIIKDLNKALDQFELSELLSG 199
Query: 121 PYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATI 180
PYDKEGAVISI+AGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKS GEEAGIKSATI
Sbjct: 200 PYDKEGAVISISAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSAGEEAGIKSATI 259
Query: 181 EVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLE 240
E+EGRYAYGY+SGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLP+ES+NVE+PEEDLE
Sbjct: 260 EIEGRYAYGYISGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPDESMNVELPEEDLE 319
Query: 241 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAE 300
ISFSRAGGKGGQNVNKVETAVRITHIPTGVT+RCTEERSQLANKIKALSRLKAKLLVIAE
Sbjct: 320 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTVRCTEERSQLANKIKALSRLKAKLLVIAE 379
Query: 301 EQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKS 360
EQRA+E KQIRGD VKAEWGQQIRNYVFHPYKLVKDVRTG+ET DI SV+DGEL+PFIK+
Sbjct: 380 EQRASEIKQIRGDAVKAEWGQQIRNYVFHPYKLVKDVRTGYETSDIVSVMDGELEPFIKA 439
Query: 361 YLKHKYSMTMST 372
YLK+KYS+ +ST
Sbjct: 440 YLKYKYSIALST 451
>emb|CAC36322.1| translation releasing factor2 [Arabidopsis thaliana]
gi|30692913|ref|NP_851097.1| peptide chain release
factor, putative [Arabidopsis thaliana]
Length = 455
Score = 613 bits (1581), Expect = e-174
Identities = 300/373 (80%), Positives = 344/373 (91%)
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
FY+LRKDVEIAS RV+EIR S+ LQ LEQE+ LE +A+ +SFWDDR KAQ+TLS+L D+
Sbjct: 80 FYTLRKDVEIASARVEEIRASANLQQLEQEITNLESKATDTSFWDDRTKAQETLSSLNDL 139
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSG 120
K++++LL+++KT VEDAETIV LTEEM+S D L EEA +IKELNKS+D+FELTQLLSG
Sbjct: 140 KDRMRLLSEFKTMVEDAETIVKLTEEMDSTDVSLLEEAMGIIKELNKSLDKFELTQLLSG 199
Query: 121 PYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATI 180
PYDKEGAV+ ITAGAGGTDAQDWADMLLRMY+RWGEKQRYKT+VVE S GEEAGIKSAT+
Sbjct: 200 PYDKEGAVVYITAGAGGTDAQDWADMLLRMYMRWGEKQRYKTKVVEMSNGEEAGIKSATL 259
Query: 181 EVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLE 240
E+EGRYAYGY+SGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEE++ +EIPEEDL+
Sbjct: 260 EIEGRYAYGYISGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEEAVGIEIPEEDLD 319
Query: 241 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAE 300
ISF+RAGGKGGQNVNKVETAVRITHIPTGV +RCTEERSQLANK +AL RLKAKL+VIAE
Sbjct: 320 ISFTRAGGKGGQNVNKVETAVRITHIPTGVAVRCTEERSQLANKTRALIRLKAKLMVIAE 379
Query: 301 EQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKS 360
EQRATE K+IRGD VKAEWGQQIRNYVFHPYKLVKDVRTGHET DITSV+DG+LDPFIK+
Sbjct: 380 EQRATEIKEIRGDAVKAEWGQQIRNYVFHPYKLVKDVRTGHETSDITSVMDGDLDPFIKA 439
Query: 361 YLKHKYSMTMSTS 373
YLKHKY++ M+++
Sbjct: 440 YLKHKYTLAMASA 452
>gb|AAM51328.1| putative translation releasing factor RF-2 [Arabidopsis thaliana]
gi|15810391|gb|AAL07083.1| putative translation
releasing factor RF-2 [Arabidopsis thaliana]
gi|21536944|gb|AAM61285.1| translation releasing factor
RF-2 [Arabidopsis thaliana] gi|8777302|dbj|BAA96892.1|
translation releasing factor RF-2 [Arabidopsis thaliana]
gi|30692908|ref|NP_851096.1| peptide chain release
factor, putative [Arabidopsis thaliana]
Length = 456
Score = 613 bits (1581), Expect = e-174
Identities = 300/373 (80%), Positives = 344/373 (91%)
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
FY+LRKDVEIAS RV+EIR S+ LQ LEQE+ LE +A+ +SFWDDR KAQ+TLS+L D+
Sbjct: 81 FYTLRKDVEIASARVEEIRASANLQQLEQEITNLESKATDTSFWDDRTKAQETLSSLNDL 140
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSG 120
K++++LL+++KT VEDAETIV LTEEM+S D L EEA +IKELNKS+D+FELTQLLSG
Sbjct: 141 KDRMRLLSEFKTMVEDAETIVKLTEEMDSTDVSLLEEAMGIIKELNKSLDKFELTQLLSG 200
Query: 121 PYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATI 180
PYDKEGAV+ ITAGAGGTDAQDWADMLLRMY+RWGEKQRYKT+VVE S GEEAGIKSAT+
Sbjct: 201 PYDKEGAVVYITAGAGGTDAQDWADMLLRMYMRWGEKQRYKTKVVEMSNGEEAGIKSATL 260
Query: 181 EVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLE 240
E+EGRYAYGY+SGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEE++ +EIPEEDL+
Sbjct: 261 EIEGRYAYGYISGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEEAVGIEIPEEDLD 320
Query: 241 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAE 300
ISF+RAGGKGGQNVNKVETAVRITHIPTGV +RCTEERSQLANK +AL RLKAKL+VIAE
Sbjct: 321 ISFTRAGGKGGQNVNKVETAVRITHIPTGVAVRCTEERSQLANKTRALIRLKAKLMVIAE 380
Query: 301 EQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKS 360
EQRATE K+IRGD VKAEWGQQIRNYVFHPYKLVKDVRTGHET DITSV+DG+LDPFIK+
Sbjct: 381 EQRATEIKEIRGDAVKAEWGQQIRNYVFHPYKLVKDVRTGHETSDITSVMDGDLDPFIKA 440
Query: 361 YLKHKYSMTMSTS 373
YLKHKY++ M+++
Sbjct: 441 YLKHKYTLAMASA 453
>ref|XP_478639.1| putative translation releasing factor2 [Oryza sativa (japonica
cultivar-group)] gi|33146529|dbj|BAC79675.1| putative
translation releasing factor2 [Oryza sativa (japonica
cultivar-group)]
Length = 460
Score = 595 bits (1535), Expect = e-169
Identities = 296/375 (78%), Positives = 336/375 (88%)
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
FY+LRKDVE+A RV E+R+S+GL LE+E+A LE++++ SS WDD +KAQ+ L L +V
Sbjct: 85 FYALRKDVELAVARVSEVRQSAGLDQLEEEIASLEKKSADSSLWDDPSKAQEILVALTEV 144
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSG 120
K+++KLLND K QVE+AETIV LTEE++S+D GL EEAS +IK LNK++D FE+TQLLSG
Sbjct: 145 KDRVKLLNDLKLQVEEAETIVKLTEELDSIDTGLLEEASKIIKALNKALDNFEMTQLLSG 204
Query: 121 PYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATI 180
PYDKEGAVI+ITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKS GEEAGIKSATI
Sbjct: 205 PYDKEGAVINITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSPGEEAGIKSATI 264
Query: 181 EVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLE 240
E+EGRYAYGYLSGEKGTHRIVRQSPFN+KGLRQTSF+GVEVMPLLPEES++VEIPEEDLE
Sbjct: 265 ELEGRYAYGYLSGEKGTHRIVRQSPFNAKGLRQTSFAGVEVMPLLPEESMDVEIPEEDLE 324
Query: 241 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAE 300
ISF+RAGGKGGQNVNKVETAVR+ HIPTG+ +RCTEERSQLANKIKALSRLKAKLLVIAE
Sbjct: 325 ISFTRAGGKGGQNVNKVETAVRMVHIPTGIAVRCTEERSQLANKIKALSRLKAKLLVIAE 384
Query: 301 EQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKS 360
EQRA+E KQIRGD VKAEWGQQIRNYVFHPYKLVKDVRT ET DIT V+DGELD FI++
Sbjct: 385 EQRASEIKQIRGDAVKAEWGQQIRNYVFHPYKLVKDVRTACETSDITGVMDGELDTFIRA 444
Query: 361 YLKHKYSMTMSTSGV 375
YLK+K S V
Sbjct: 445 YLKYKLSAAAEEQSV 459
>ref|NP_198466.2| peptide chain release factor, putative [Arabidopsis thaliana]
Length = 391
Score = 448 bits (1152), Expect = e-124
Identities = 219/276 (79%), Positives = 253/276 (91%)
Query: 1 FYSLRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADV 60
FY+LRKDVEIAS RV+EIR S+ LQ LEQE+ LE +A+ +SFWDDR KAQ+TLS+L D+
Sbjct: 81 FYTLRKDVEIASARVEEIRASANLQQLEQEITNLESKATDTSFWDDRTKAQETLSSLNDL 140
Query: 61 KEKIKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSG 120
K++++LL+++KT VEDAETIV LTEEM+S D L EEA +IKELNKS+D+FELTQLLSG
Sbjct: 141 KDRMRLLSEFKTMVEDAETIVKLTEEMDSTDVSLLEEAMGIIKELNKSLDKFELTQLLSG 200
Query: 121 PYDKEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATI 180
PYDKEGAV+ ITAGAGGTDAQDWADMLLRMY+RWGEKQRYKT+VVE S GEEAGIKSAT+
Sbjct: 201 PYDKEGAVVYITAGAGGTDAQDWADMLLRMYMRWGEKQRYKTKVVEMSNGEEAGIKSATL 260
Query: 181 EVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLE 240
E+EGRYAYGY+SGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEE++ +EIPEEDL+
Sbjct: 261 EIEGRYAYGYISGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEEAVGIEIPEEDLD 320
Query: 241 ISFSRAGGKGGQNVNKVETAVRITHIPTGVTLRCTE 276
ISF+RAGGKGGQNVNKVETAVRITHIPTGV +RCT+
Sbjct: 321 ISFTRAGGKGGQNVNKVETAVRITHIPTGVAVRCTD 356
>sp|P74476|RF2_SYNY3 Peptide chain release factor 2 (RF-2)
Length = 372
Score = 426 bits (1094), Expect = e-118
Identities = 210/361 (58%), Positives = 279/361 (77%), Gaps = 2/361 (0%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
L++++E+ S R+ + ++ L L+ ++ LE+ A+ FWDD +AQQ L TL + K +
Sbjct: 8 LKRNLELISSRLGQTQDYLDLPGLKAKVQDLEQCAAQPDFWDDTDQAQQILQTLNETKSQ 67
Query: 64 IKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYD 123
++ ++ Q +D++ IV L E + D+ L EA + +++L K +DR+EL QLLSGPYD
Sbjct: 68 LEQWGIWQQQWQDSQAIVELLELED--DQALLTEAETTLEQLQKELDRWELQQLLSGPYD 125
Query: 124 KEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVE 183
+GA ++I AGAGGTDAQDWA+MLLRMY RW EKQ YK + E S G+EAG+KS T+E+E
Sbjct: 126 AKGATLTINAGAGGTDAQDWAEMLLRMYTRWSEKQGYKVHLAEISEGDEAGLKSVTLEIE 185
Query: 184 GRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISF 243
GRYAYGYL EKGTHR+VR SPFN+ G RQTSF+GVEVMPLL EE+++++IP++DL+IS
Sbjct: 186 GRYAYGYLKSEKGTHRLVRISPFNANGKRQTSFAGVEVMPLLGEEAISLDIPDKDLDIST 245
Query: 244 SRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQR 303
SRAGGKGGQNVNKVETAVRI H+PTG+ +RCT+ERSQL NK KAL+ LKAKLL++ EEQR
Sbjct: 246 SRAGGKGGQNVNKVETAVRIVHLPTGLAVRCTQERSQLQNKEKALAILKAKLLIVLEEQR 305
Query: 304 ATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
A +IRGD+V+A WG QIRNYVFHPY+LVKD+RT ET D+ V+DGEL FI++YL+
Sbjct: 306 AQAIAEIRGDMVEAAWGTQIRNYVFHPYQLVKDLRTNVETTDVGGVMDGELSDFIEAYLR 365
Query: 364 H 364
H
Sbjct: 366 H 366
>ref|ZP_00513812.1| Peptide chain release factor 2 [Crocosphaera watsonii WH 8501]
gi|67857776|gb|EAM53015.1| Peptide chain release factor
2 [Crocosphaera watsonii WH 8501]
Length = 358
Score = 411 bits (1057), Expect = e-113
Identities = 203/340 (59%), Positives = 259/340 (75%), Gaps = 2/340 (0%)
Query: 24 LQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVML 83
L L+ + LE ++ FWD AQ TL L D+K +++ + Q++D + I L
Sbjct: 14 LPALKAVIKDLENTSAQPQFWDQPETAQTTLQELNDLKSQLEQYQQWCLQLDDTKAIAEL 73
Query: 84 TEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDW 143
E + D L +EA I +LN ++DR+EL QLLSG YD +GAV++I AGAGGTDAQDW
Sbjct: 74 LELED--DATLRQEAEDNIIQLNHNLDRWELQQLLSGIYDTKGAVLTINAGAGGTDAQDW 131
Query: 144 ADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQ 203
A MLLRMY RWGE+Q YK + E S G+EAGIKSAT+E+EGRYA+GYL GEKGTHR+VR
Sbjct: 132 AQMLLRMYTRWGEQQGYKVHLTELSEGDEAGIKSATLEIEGRYAFGYLKGEKGTHRLVRI 191
Query: 204 SPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRI 263
SPFN+ G RQTSF+G+E+MP L E+ L V+IP++DL+IS SR+GGKGGQNVNKVETAVR+
Sbjct: 192 SPFNANGKRQTSFAGIEIMPALEEDDLKVDIPDKDLDISTSRSGGKGGQNVNKVETAVRV 251
Query: 264 THIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQI 323
H+PTG+ +RCT+ERSQL NK KAL+ LKAKLL+IAEEQRA E +IRGD+V+A WG QI
Sbjct: 252 VHVPTGIAVRCTQERSQLQNKEKALALLKAKLLIIAEEQRAQEIAEIRGDIVEAAWGTQI 311
Query: 324 RNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
RNYVFHPY++VKD+RT ET D+ V+DG LD FI++YL+
Sbjct: 312 RNYVFHPYQMVKDLRTNIETTDVNGVMDGSLDEFIEAYLR 351
>emb|CAE08826.1| peptide chain release factor RF-2 [Synechococcus sp. WH 8102]
gi|33866841|ref|NP_898400.1| peptide chain release
factor RF-2 [Synechococcus sp. WH 8102]
Length = 374
Score = 395 bits (1014), Expect = e-108
Identities = 202/359 (56%), Positives = 263/359 (72%), Gaps = 3/359 (0%)
Query: 5 RKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKI 64
++D+ + R+ ++ + L+ LE+ A+ FWDD+ AQ+ + L +VK ++
Sbjct: 7 KRDLSELTDRLGHAQDCLDVPALKARQQDLEQLAAQPDFWDDQQAAQKQMRRLDEVKAQL 66
Query: 65 KLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDK 124
+ L D+ V+DA+ + L E D+ + EA + +L + +DR+EL +LLSG YDK
Sbjct: 67 QQLADWGGAVDDAKATLELYEL--EPDEEMLTEAQEGLNQLRQGLDRWELERLLSGDYDK 124
Query: 125 EGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEG 184
EGAV++I AGAGGTDAQDWA MLLRMY RW E K V E S GEEAGIKS TIEVEG
Sbjct: 125 EGAVLTINAGAGGTDAQDWAQMLLRMYTRWAEDHGMKVTVDELSEGEEAGIKSCTIEVEG 184
Query: 185 RYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFS 244
RYAYGYL EKGTHR+VR SPFN+ RQTSF+G+EVMP + EE ++++IPE+DLE++ S
Sbjct: 185 RYAYGYLRNEKGTHRLVRISPFNANDKRQTSFAGIEVMPKIDEE-VDIDIPEKDLEVTTS 243
Query: 245 RAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRA 304
R+GG GGQNVNKVETAVRI HIPTG+ +RCT+ERSQL NK KA++ LKAKLLVIA+EQRA
Sbjct: 244 RSGGAGGQNVNKVETAVRILHIPTGLAVRCTQERSQLQNKEKAMALLKAKLLVIAQEQRA 303
Query: 305 TEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
E IRGD+V+A WG QIRNYVFHPY++VKD+RT ET D+ +V+DG LDPFI + L+
Sbjct: 304 AEIADIRGDIVEAAWGNQIRNYVFHPYQMVKDLRTSEETNDVQAVMDGALDPFIDASLR 362
>ref|NP_924179.1| peptide chain release factor [Gloeobacter violaceus PCC 7421]
gi|35211797|dbj|BAC89174.1| peptide chain release factor
[Gloeobacter violaceus PCC 7421]
Length = 354
Score = 393 bits (1009), Expect = e-108
Identities = 188/338 (55%), Positives = 259/338 (76%), Gaps = 3/338 (0%)
Query: 31 LAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLTEEMESV 90
+A LE++AS FWDD+A AQ+ L L ++K+ ++ ++ + DAE ++ L +E +
Sbjct: 20 IAALEQQASAPDFWDDQAAAQKALQNLNNLKQPLEQFGRWRNLLSDAEVVLELVQE--TG 77
Query: 91 DKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLRM 150
D+ L EA I+ L +DR+EL QLLSGPYD+ GA++SI AGAGGTDAQDWA+MLLRM
Sbjct: 78 DESLLPEAEGNIEALVGELDRWELEQLLSGPYDERGAILSINAGAGGTDAQDWAEMLLRM 137
Query: 151 YVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKG 210
Y RW E+Q Y+ ++E S GEEAG+KSAT+E++GRYAYGYLS EKGTHR+VR SPFN+
Sbjct: 138 YTRWAERQGYQAHLLELSEGEEAGVKSATLEIDGRYAYGYLSAEKGTHRLVRISPFNAND 197
Query: 211 LRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGV 270
RQTSF+GVEVMP+L ++S+ V+I +D+EI R+GGKGGQNVNKVET VR+TH PTG+
Sbjct: 198 KRQTSFAGVEVMPVL-DDSVQVDIRPDDIEIDTFRSGGKGGQNVNKVETGVRVTHKPTGI 256
Query: 271 TLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYVFHP 330
++RCT+ER+QL N+ KA+ LKA+LLV+ EE+R IRG+ V A WG QIR+YVFHP
Sbjct: 257 SVRCTQERAQLQNRNKAMEMLKARLLVLEEEKRQQHLSDIRGEAVDAAWGNQIRSYVFHP 316
Query: 331 YKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHKYSM 368
Y++VKD+RT ET D+T V++G+++PFI+ YL+ ++ +
Sbjct: 317 YQMVKDLRTEEETSDVTGVMNGKIEPFIERYLRRQHKL 354
>emb|CAE22237.1| peptide-chain-release factor RF-2 [Prochlorococcus marinus str. MIT
9313] gi|33864328|ref|NP_895888.1| peptide-chain-release
factor RF-2 [Prochlorococcus marinus str. MIT 9313]
Length = 375
Score = 391 bits (1004), Expect = e-107
Identities = 199/359 (55%), Positives = 267/359 (73%), Gaps = 3/359 (0%)
Query: 5 RKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKI 64
++D+ + R+ + ++ + L LE+ A+ FW+D+ AQ+ + L +VK ++
Sbjct: 8 KRDLSELTDRLGQAQDCLDVPALNARQQDLEQLAAQPDFWNDQQNAQKQMRQLDEVKAQL 67
Query: 65 KLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDK 124
+ +++ V DA+ + L + +E D+ L EA S +K+L + +DR+EL +LLSG YDK
Sbjct: 68 TQITLWRSAVNDAQATLELYD-LEPDDEML-SEAQSGLKQLRQGLDRWELERLLSGEYDK 125
Query: 125 EGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEG 184
EGAV+SI AGAGGTDAQDWA MLLRMY RW E V E S GEEAGIKSAT+E+EG
Sbjct: 126 EGAVLSINAGAGGTDAQDWAQMLLRMYTRWAEDHDMTVTVDELSEGEEAGIKSATVEIEG 185
Query: 185 RYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFS 244
RYAYGYL EKGTHR+VR SPFN+ G RQTSF+GVE+MP + +E + ++IPE+DLE++ S
Sbjct: 186 RYAYGYLRNEKGTHRLVRISPFNANGKRQTSFAGVEIMPKIDQE-VELDIPEKDLEVTTS 244
Query: 245 RAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRA 304
R+GG GGQNVNKVETAVRI HIPTG+ +RCT+ERSQL NK KAL+ LKAKL+VIA+EQRA
Sbjct: 245 RSGGAGGQNVNKVETAVRILHIPTGLAVRCTQERSQLQNKEKALALLKAKLMVIAQEQRA 304
Query: 305 TEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
E IRGD+V+A WG QIRNYVFHPY++VKD+RT ET D+ +V++G+LD FI++ L+
Sbjct: 305 AEIADIRGDIVEAAWGNQIRNYVFHPYQMVKDLRTAEETTDVQAVMNGDLDRFIQTLLR 363
>gb|AAP99251.1| Protein chain release factor B [Prochlorococcus marinus subsp.
marinus str. CCMP1375] gi|33239657|ref|NP_874599.1|
Protein chain release factor B [Prochlorococcus marinus
subsp. marinus str. CCMP1375]
Length = 356
Score = 390 bits (1003), Expect = e-107
Identities = 197/329 (59%), Positives = 255/329 (76%), Gaps = 3/329 (0%)
Query: 34 LEEEASCSSFWDDRAKAQQTLSTLADVKEKIKLLNDYKTQVEDAETIVMLTEEMESVDKG 93
LE+ A+ FW+D+ KA++ + L +VK+++ L ++ +EDA+ + L E D
Sbjct: 18 LEQIAAQPDFWNDQQKARKQMRQLDEVKDQLTKLANWNAAIEDAKVSLELYEL--EPDNE 75
Query: 94 LYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLRMYVR 153
+ EEA + +L +D +EL +LLSGPYDKEGAV++I AGAGGTDAQDWA MLLRMY+R
Sbjct: 76 MIEEAQIGLCQLKTELDLWELERLLSGPYDKEGAVLAINAGAGGTDAQDWAQMLLRMYMR 135
Query: 154 WGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKGLRQ 213
W E K + E S GEEAGIKSATIE+EGRYAYGYL EKGTHR+VR SPFN+ G RQ
Sbjct: 136 WAESHGMKVILSELSEGEEAGIKSATIEIEGRYAYGYLQNEKGTHRLVRISPFNANGKRQ 195
Query: 214 TSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGVTLR 273
TSF+G+EVMP L EE +++EIPE+DLE++ SR+GG GGQNVNKVETAVRI HIPTG+ +R
Sbjct: 196 TSFAGIEVMPQL-EEDVHLEIPEKDLEVTTSRSGGAGGQNVNKVETAVRILHIPTGLAVR 254
Query: 274 CTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYVFHPYKL 333
CT+ERSQL NK KA++ LKAKL++IA+EQRA E IRGD+V+A WG QIRNYVFHPY++
Sbjct: 255 CTQERSQLQNKEKAMALLKAKLMIIAQEQRAAEIADIRGDIVEAAWGNQIRNYVFHPYQM 314
Query: 334 VKDVRTGHETPDITSVIDGELDPFIKSYL 362
VKD+R+G E+ D+ SV++G+LD FI+S L
Sbjct: 315 VKDLRSGTESNDVESVMNGDLDLFIQSLL 343
>ref|NP_441899.1| peptide chain release factor [Synechocystis sp. PCC 6803]
gi|1653665|dbj|BAA18577.1| peptide chain release factor
[Synechocystis sp. PCC 6803]
Length = 288
Score = 386 bits (992), Expect = e-106
Identities = 184/274 (67%), Positives = 228/274 (83%)
Query: 91 DKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLRM 150
D+ L EA + +++L K +DR+EL QLLSGPYD +GA ++I AGAGGTDAQDWA+MLLRM
Sbjct: 9 DQALLTEAETTLEQLQKELDRWELQQLLSGPYDAKGATLTINAGAGGTDAQDWAEMLLRM 68
Query: 151 YVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSKG 210
Y RW EKQ YK + E S G+EAG+KS T+E+EGRYAYGYL EKGTHR+VR SPFN+ G
Sbjct: 69 YTRWSEKQGYKVHLAEISEGDEAGLKSVTLEIEGRYAYGYLKSEKGTHRLVRISPFNANG 128
Query: 211 LRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTGV 270
RQTSF+GVEVMPLL EE+++++IP++DL+IS SRAGGKGGQNVNKVETAVRI H+PTG+
Sbjct: 129 KRQTSFAGVEVMPLLGEEAISLDIPDKDLDISTSRAGGKGGQNVNKVETAVRIVHLPTGL 188
Query: 271 TLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYVFHP 330
+RCT+ERSQL NK KAL+ LKAKLL++ EEQRA +IRGD+V+A WG QIRNYVFHP
Sbjct: 189 AVRCTQERSQLQNKEKALAILKAKLLIVLEEQRAQAIAEIRGDMVEAAWGTQIRNYVFHP 248
Query: 331 YKLVKDVRTGHETPDITSVIDGELDPFIKSYLKH 364
Y+LVKD+RT ET D+ V+DGEL FI++YL+H
Sbjct: 249 YQLVKDLRTNVETTDVGGVMDGELSDFIEAYLRH 282
>ref|ZP_00163511.1| COG1186: Protein chain release factor B [Synechococcus elongatus
PCC 7942]
Length = 304
Score = 384 bits (985), Expect = e-105
Identities = 189/298 (63%), Positives = 236/298 (78%), Gaps = 1/298 (0%)
Query: 76 DAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGA 135
DA+ +L D L+EEA + ++ L + +D++EL QLLSGPYD EGA++SI AGA
Sbjct: 7 DADITAILELLAIEPDPALFEEAETNVQRLQRELDQWELQQLLSGPYDAEGAILSINAGA 66
Query: 136 GGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEK 195
GGTDAQDWA+MLLRMY RW E Q YK R+ E S GEEAGIKS T+E+EG+YAYGYL EK
Sbjct: 67 GGTDAQDWAEMLLRMYTRWAEDQGYKVRITELSEGEEAGIKSVTLEIEGKYAYGYLKAEK 126
Query: 196 GTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVN 255
GTHR+VR SPFN+ RQTSF+GVEVMP+L + +++EIPE+DLEI+ SRAGGKGGQNVN
Sbjct: 127 GTHRLVRISPFNANDKRQTSFAGVEVMPVL-DTDIDLEIPEKDLEITTSRAGGKGGQNVN 185
Query: 256 KVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVV 315
KVETAVRI H+PTG+ +RCTEERSQL NK KAL LKAKLLV+ +EQ+A E +IRGD+V
Sbjct: 186 KVETAVRIVHLPTGLAVRCTEERSQLQNKEKALVILKAKLLVVLQEQKAKEVAEIRGDMV 245
Query: 316 KAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHKYSMTMSTS 373
+A WG QIRNYVFHPY+LVKD+RTG ET I V++G+L+PFI++YL+ M + T+
Sbjct: 246 EAAWGNQIRNYVFHPYQLVKDLRTGVETKAIADVMNGDLNPFIQAYLRQDNQMLIETT 303
>ref|YP_171822.1| peptide chain release factor RF-2 [Synechococcus elongatus PCC
6301] gi|56686080|dbj|BAD79302.1| peptide chain release
factor RF-2 [Synechococcus elongatus PCC 6301]
Length = 304
Score = 382 bits (982), Expect = e-105
Identities = 188/298 (63%), Positives = 236/298 (79%), Gaps = 1/298 (0%)
Query: 76 DAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGA 135
DA+ +L D L+EEA + ++ L + +D++EL QLLSGPYD EGA++SI AGA
Sbjct: 7 DADITAILELLAIEPDPALFEEAETNVQRLQRELDQWELQQLLSGPYDAEGAILSINAGA 66
Query: 136 GGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEK 195
GGTDAQDWA+MLLRMY RW E Q YK R+ E S GEEAGIKS T+E+EG+YAYGYL EK
Sbjct: 67 GGTDAQDWAEMLLRMYTRWAEDQGYKVRITELSEGEEAGIKSVTLEIEGKYAYGYLKAEK 126
Query: 196 GTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVN 255
GTHR+VR SPFN+ RQTSF+GVEVMP+L + +++EIPE+DLEI+ SRAGGKGGQNVN
Sbjct: 127 GTHRLVRISPFNANDKRQTSFAGVEVMPVL-DTDIDLEIPEKDLEITTSRAGGKGGQNVN 185
Query: 256 KVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVV 315
KVETAVRI H+PTG+ +RCTEERSQL NK KAL L+AKLLV+ +EQ+A E +IRGD+V
Sbjct: 186 KVETAVRIVHLPTGLAVRCTEERSQLQNKEKALVILRAKLLVVLQEQKAKEVAEIRGDMV 245
Query: 316 KAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHKYSMTMSTS 373
+A WG QIRNYVFHPY+LVKD+RTG ET I V++G+L+PFI++YL+ M + T+
Sbjct: 246 EAAWGNQIRNYVFHPYQLVKDLRTGVETKAIADVMNGDLNPFIQAYLRQDNQMLIETT 303
>ref|NP_681317.1| peptide chain release factor 2 [Thermosynechococcus elongatus BP-1]
gi|22294248|dbj|BAC08079.1| peptide chain release factor
2 [Thermosynechococcus elongatus BP-1]
Length = 290
Score = 379 bits (972), Expect = e-103
Identities = 186/276 (67%), Positives = 228/276 (82%), Gaps = 1/276 (0%)
Query: 90 VDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWADMLLR 149
+D+GL EA++L+KEL+ ++R+EL QLLSGPYDK A+++I AGAGGTDAQDWA+MLLR
Sbjct: 12 LDEGLLTEATTLLKELDTQLNRWELEQLLSGPYDKNNAIVTINAGAGGTDAQDWAEMLLR 71
Query: 150 MYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSPFNSK 209
MY RW E+ Y+T + E S GEEAGIKSAT+E+ GRYAYGYL EKGTHR+VR SPFN+
Sbjct: 72 MYTRWAERHGYQTHLAELSEGEEAGIKSATLEIRGRYAYGYLRSEKGTHRLVRISPFNAN 131
Query: 210 GLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITHIPTG 269
G RQTSF+GV+VMP L ++S+ VEIPE DLEI+ SR+GGKGGQNVNKVETAVRI H PTG
Sbjct: 132 GKRQTSFAGVDVMPEL-DQSVTVEIPESDLEITTSRSGGKGGQNVNKVETAVRIVHKPTG 190
Query: 270 VTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRNYVFH 329
+ +RCT+ERSQL NK KAL+ LKAKLLVIA+EQ+A E IRG+ V+A WG QIRNYVFH
Sbjct: 191 LAVRCTQERSQLQNKEKALAILKAKLLVIAQEQKAKEIADIRGEAVEAAWGNQIRNYVFH 250
Query: 330 PYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHK 365
PY+LVKD+RT ET I V+DG LDPFI++YL+ +
Sbjct: 251 PYQLVKDLRTEVETTAIQDVMDGNLDPFIQAYLRQQ 286
>dbj|BAB77797.1| peptide chain release factor [Nostoc sp. PCC 7120]
gi|17227769|ref|NP_484317.1| peptide chain release
factor [Nostoc sp. PCC 7120] gi|25299700|pir||AI1840
peptide chain release factor [imported] - Nostoc sp.
(strain PCC 7120)
Length = 292
Score = 378 bits (971), Expect = e-103
Identities = 186/280 (66%), Positives = 232/280 (82%), Gaps = 2/280 (0%)
Query: 86 EMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWAD 145
E+E+ D+ L +EA S I +LN+ +D++EL QLLSGPYD EGAV++I AGAGGTDAQDWA
Sbjct: 6 ELET-DEALLQEAESTITKLNRDLDQWELQQLLSGPYDAEGAVLTINAGAGGTDAQDWAF 64
Query: 146 MLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSP 205
MLLRMY RW E Q YK + E+S G+EAGIKSAT+E+ GRYAYGYL E GTHR+VR SP
Sbjct: 65 MLLRMYTRWAEDQGYKVALAEESEGDEAGIKSATLEITGRYAYGYLRSETGTHRLVRISP 124
Query: 206 FNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITH 265
FN+ G RQTSF+GVEVMP L + ++ ++IP++DLEI+ +R+GGKGGQNVNKVETAVR+ H
Sbjct: 125 FNANGKRQTSFAGVEVMPQL-DNTVQLDIPDKDLEITTTRSGGKGGQNVNKVETAVRVVH 183
Query: 266 IPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRN 325
+PTG+ +RCTEERSQL NK KAL+RLKAKLLVIA+EQ+A E QIRGD+V+A WG QIRN
Sbjct: 184 LPTGIAVRCTEERSQLQNKEKALARLKAKLLVIAQEQKAQEIAQIRGDMVEASWGNQIRN 243
Query: 326 YVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHK 365
YVFHPY++VKD+RT ET I V++GELD FI+SYL+ +
Sbjct: 244 YVFHPYQMVKDLRTNVETTAIADVMNGELDMFIQSYLRQE 283
>ref|ZP_00162728.2| COG1186: Protein chain release factor B [Anabaena variabilis ATCC
29413]
Length = 292
Score = 377 bits (969), Expect = e-103
Identities = 185/280 (66%), Positives = 232/280 (82%), Gaps = 2/280 (0%)
Query: 86 EMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWAD 145
E+E+ D+ L +EA S I +LN+ +D++EL QLLSGPYD EGAV++I AGAGGTDAQDWA
Sbjct: 6 ELET-DEALLQEAESTITKLNRDLDQWELQQLLSGPYDAEGAVLTINAGAGGTDAQDWAF 64
Query: 146 MLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSP 205
MLLRMY RW E Q YK + E+S G+EAGIKSAT+E+ GRYAYGYL E GTHR+VR SP
Sbjct: 65 MLLRMYTRWAEDQGYKVALAEESEGDEAGIKSATLEITGRYAYGYLRSETGTHRLVRISP 124
Query: 206 FNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITH 265
FN+ G RQTSF+GVEVMP L + ++ ++IP++DLEI+ +R+GGKGGQNVNKVETAVR+ H
Sbjct: 125 FNANGKRQTSFAGVEVMPQL-DNTVQLDIPDKDLEITTTRSGGKGGQNVNKVETAVRVVH 183
Query: 266 IPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRN 325
+PTG+ +RCTEERSQL NK KAL+RLKAKLLVIA+EQ+A E QIRGD+V+A WG QIRN
Sbjct: 184 LPTGIAVRCTEERSQLQNKEKALARLKAKLLVIAQEQKAQEIAQIRGDMVEASWGNQIRN 243
Query: 326 YVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHK 365
YVFHPY++VKD+RT ET I V++GELD FI++YL+ +
Sbjct: 244 YVFHPYQMVKDLRTNAETTAIADVMNGELDMFIQAYLRQE 283
>ref|ZP_00325872.1| COG1186: Protein chain release factor B [Trichodesmium erythraeum
IMS101]
Length = 288
Score = 377 bits (968), Expect = e-103
Identities = 185/288 (64%), Positives = 237/288 (82%), Gaps = 2/288 (0%)
Query: 86 EMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDKEGAVISITAGAGGTDAQDWAD 145
E+E+ D+ L EA+ I LN+ +D++EL QLLSG YD+ GAV++I AGAGGTDAQDWA+
Sbjct: 2 ELET-DESLLAEANLTIANLNQELDQWELQQLLSGLYDQNGAVVTINAGAGGTDAQDWAE 60
Query: 146 MLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEGRYAYGYLSGEKGTHRIVRQSP 205
MLLRMY RWGE+ +YK ++ E S G+EAG+KSAT+E+ GRYAYGYL EKGTHR+VR SP
Sbjct: 61 MLLRMYTRWGEEHKYKVQLAEISEGDEAGLKSATLEIIGRYAYGYLKAEKGTHRLVRISP 120
Query: 206 FNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFSRAGGKGGQNVNKVETAVRITH 265
FN+ G RQTSF+GVEVMP+L + S+ ++IPE+DLEI+ SR+GGKGGQNVNKVETAVR+ H
Sbjct: 121 FNANGKRQTSFAGVEVMPIL-DHSVELDIPEKDLEITTSRSGGKGGQNVNKVETAVRVVH 179
Query: 266 IPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRATEFKQIRGDVVKAEWGQQIRN 325
IPTG+++RCT+ERSQL NK KAL+ LKA+LL+IA+EQRA E +IRGD+V+A WG QIRN
Sbjct: 180 IPTGLSVRCTQERSQLQNKDKALALLKARLLIIAQEQRAQEINEIRGDMVEAAWGNQIRN 239
Query: 326 YVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKHKYSMTMSTS 373
YVFHPY++VKD+RT ET I V++GELD FI++YL+ + M S S
Sbjct: 240 YVFHPYQMVKDLRTQKETTAINDVMNGELDQFIEAYLRQENQMVESQS 287
>ref|NP_892301.1| peptide chain release factor RF-2 [Prochlorococcus marinus subsp.
pastoris str. CCMP1986] gi|33633682|emb|CAE18639.1|
peptide chain release factor RF-2 [Prochlorococcus
marinus subsp. pastoris str. CCMP1986]
Length = 373
Score = 367 bits (941), Expect = e-100
Identities = 186/363 (51%), Positives = 263/363 (72%), Gaps = 3/363 (0%)
Query: 5 RKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEKI 64
+KD+ ++R+ ++ + L + +LE+ A+ FW ++A A++ + L DV++++
Sbjct: 8 KKDLSELTKRLGNAQDCLDVPNLVAKKKELEQIAAQPEFWGNQADAKKHMLNLDDVRDQL 67
Query: 65 KLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYDK 124
+LL+ +K + DAE + L +E ++ + E L+ L +++D++E +LL G YDK
Sbjct: 68 QLLDKWKMFISDAEASLELYS-LEPEEEMIVESHQGLVT-LRENLDKWEFQRLLCGEYDK 125
Query: 125 EGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVEG 184
E A++SI AGAGGTDAQDW ++LLRMY RW + + + S GEEAGIKS T EV G
Sbjct: 126 EAAIVSINAGAGGTDAQDWVEILLRMYSRWVDNNGMSLEITDLSDGEEAGIKSVTFEVNG 185
Query: 185 RYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISFS 244
+YAYGYL EKGTHR+VR SPFN+ G RQTSF+GVEVMP L ++S++++IP++DLEI+ S
Sbjct: 186 KYAYGYLQNEKGTHRLVRISPFNANGKRQTSFAGVEVMPKL-DQSISLDIPDKDLEITTS 244
Query: 245 RAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQRA 304
R+GG GGQNVNKVETAVRI H+P+G+ +RCT+ERSQL NK KA+ LKAKLLVIA+EQRA
Sbjct: 245 RSGGAGGQNVNKVETAVRIVHLPSGIAVRCTQERSQLQNKEKAMLLLKAKLLVIAKEQRA 304
Query: 305 TEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLKH 364
+E I+GD+V+A WG QIRNYVFHPY++VKD+RT ET D+ +V++G LD FI L+
Sbjct: 305 SEISDIKGDIVEAAWGNQIRNYVFHPYQMVKDLRTMKETNDLDAVLNGGLDQFIHELLRM 364
Query: 365 KYS 367
S
Sbjct: 365 NIS 367
>ref|NP_214263.1| peptide chain release factor RF-2 [Aquifex aeolicus VF5]
gi|2984119|gb|AAC07656.1| peptide chain release factor
RF-2 [Aquifex aeolicus VF5] gi|7443581|pir||E70458
translation releasing factor RF-2 - Aquifex aeolicus
gi|6225943|sp|O67695|RF2_AQUAE Peptide chain release
factor 2 (RF-2)
Length = 373
Score = 327 bits (838), Expect = 4e-88
Identities = 168/362 (46%), Positives = 243/362 (66%), Gaps = 4/362 (1%)
Query: 4 LRKDVEIASQRVKEIRESSGLQLLEQELAKLEEEASCSSFWDDRAKAQQTLSTLADVKEK 63
L+ VE +R++++++ + LE EL +L+++ S +FW+D+ KA+Q + V+E
Sbjct: 6 LKGKVEELRKRLEDVKKILSPEKLESELKELDQKMSEPNFWEDQEKAKQVIQRRKWVEET 65
Query: 64 IKLLNDYKTQVEDAETIVMLTEEMESVDKGLYEEASSLIKELNKSIDRFELTQLLSGPYD 123
+ L + + V+D E +V +T E ++ + +E IKE+ +++ EL LSG D
Sbjct: 66 LNKLKNLEKSVKDLEELVEITSEEDTETWAMMDEE---IKEVERTLRELELKTYLSGEMD 122
Query: 124 KEGAVISITAGAGGTDAQDWADMLLRMYVRWGEKQRYKTRVVEKSLGEEAGIKSATIEVE 183
+ A ++I AGAGGT+A DWADML RMY RW EK+ Y+ +++ + + AGIKS T+ V+
Sbjct: 123 AKNAYLTIQAGAGGTEACDWADMLFRMYKRWAEKKGYEVELIDITPDDVAGIKSVTVLVK 182
Query: 184 GRYAYGYLSGEKGTHRIVRQSPFNSKGLRQTSFSGVEVMPLLPEESLNVEIPEEDLEISF 243
G YAYGYL GE+G HR+VR SPF++ R TSF+ V VMP + +E + +EI EDL+I
Sbjct: 183 GPYAYGYLKGEQGVHRLVRISPFDANARRHTSFAAVSVMPQI-DEDIKIEIKPEDLKIET 241
Query: 244 SRAGGKGGQNVNKVETAVRITHIPTGVTLRCTEERSQLANKIKALSRLKAKLLVIAEEQR 303
RA G GGQ VNK +TAVRITHIPTG+T+ C +ERSQ NK KAL LKAKL + ++
Sbjct: 242 FRASGAGGQYVNKTDTAVRITHIPTGITVSCQQERSQYQNKRKALELLKAKLYQLEMKKL 301
Query: 304 ATEFKQIRGDVVKAEWGQQIRNYVFHPYKLVKDVRTGHETPDITSVIDGELDPFIKSYLK 363
+ KQ G+ WG QIR+YVFHPYKL+KD+RTG+ET ++ +V+DGE+D FI+SYLK
Sbjct: 302 EEKKKQYEGEKTDIGWGHQIRSYVFHPYKLIKDLRTGYETGNVEAVMDGEIDEFIESYLK 361
Query: 364 HK 365
K
Sbjct: 362 WK 363
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.131 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 580,348,238
Number of Sequences: 2540612
Number of extensions: 23450015
Number of successful extensions: 72859
Number of sequences better than 10.0: 999
Number of HSP's better than 10.0 without gapping: 792
Number of HSP's successfully gapped in prelim test: 207
Number of HSP's that attempted gapping in prelim test: 70867
Number of HSP's gapped (non-prelim): 1127
length of query: 375
length of database: 863,360,394
effective HSP length: 129
effective length of query: 246
effective length of database: 535,621,446
effective search space: 131762875716
effective search space used: 131762875716
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC147179.9


