

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147011.10 - phase: 0
(165 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
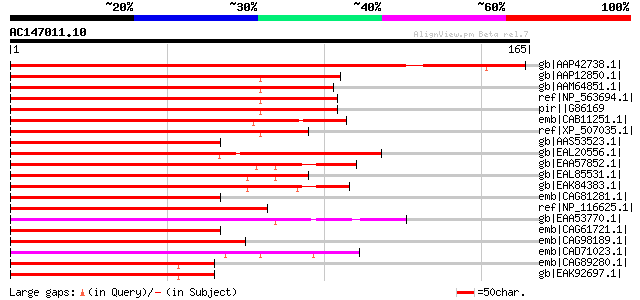
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP42738.1| At5g02270 [Arabidopsis thaliana] gi|7406423|emb|C... 259 1e-68
gb|AAP12850.1| At5g44110 [Arabidopsis thaliana] gi|9759512|dbj|B... 122 4e-27
gb|AAM64851.1| NBD-like protein [Arabidopsis thaliana] 120 2e-26
ref|NP_563694.1| ABC transporter family protein [Arabidopsis tha... 116 2e-25
pir||G86169 hypothetical protein [imported] - Arabidopsis thalia... 116 2e-25
emb|CAB11251.1| SPAC20G4.01 [Schizosaccharomyces pombe] gi|75224... 114 1e-24
ref|XP_507035.1| PREDICTED OJ1116_E04.3 gene product [Oryza sati... 111 6e-24
gb|AAS53523.1| AFR152Cp [Ashbya gossypii ATCC 10895] gi|45198670... 100 1e-20
gb|EAL20556.1| hypothetical protein CNBE4760 [Cryptococcus neofo... 95 6e-19
gb|EAA57852.1| hypothetical protein AN6512.2 [Aspergillus nidula... 94 1e-18
gb|EAL85531.1| ABC transporter, putative [Aspergillus fumigatus ... 94 2e-18
gb|EAK84383.1| hypothetical protein UM03153.1 [Ustilago maydis 5... 93 2e-18
emb|CAG81281.1| unnamed protein product [Yarrowia lipolytica CLI... 91 2e-17
ref|NP_116625.1| Part of the evolutionarily-conserved CCR4-NOT t... 89 3e-17
gb|EAA53770.1| hypothetical protein MG09520.4 [Magnaporthe grise... 89 6e-17
emb|CAG61721.1| unnamed protein product [Candida glabrata CBS138... 84 1e-15
emb|CAG98189.1| unnamed protein product [Kluyveromyces lactis NR... 84 2e-15
emb|CAD71023.1| related to ATP-binding cassette (ABC) transporte... 83 3e-15
emb|CAG89280.1| unnamed protein product [Debaryomyces hansenii C... 82 4e-15
gb|EAK92697.1| potential CCR4-NOT complex associated factor Caf1... 82 5e-15
>gb|AAP42738.1| At5g02270 [Arabidopsis thaliana] gi|7406423|emb|CAB85532.1| ABC
transporter-like protein [Arabidopsis thaliana]
gi|21554262|gb|AAM63337.1| ABC transporter-like protein
[Arabidopsis thaliana] gi|15241744|ref|NP_195847.1| ABC
transporter family protein [Arabidopsis thaliana]
gi|13899119|gb|AAK48981.1| ABC transporter-like protein
[Arabidopsis thaliana] gi|11357153|pir||T48248 ABC
transporter-like protein - Arabidopsis thaliana
Length = 328
Score = 259 bits (663), Expect = 1e-68
Identities = 131/165 (79%), Positives = 149/165 (89%), Gaps = 6/165 (3%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKPF+VLLLDEITVDLDVLARADLLKFLRKEC+ERGATIIYATHIFDGLEDWPT+IV
Sbjct: 169 MGLLKPFKVLLLDEITVDLDVLARADLLKFLRKECEERGATIIYATHIFDGLEDWPTHIV 228
Query: 61 YVAHGRLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQVDG 120
YVA+G+L+LA+P+EKVKETSK SLMRTVE WLRKERDE+R++RKERKA GLPEF+ + +
Sbjct: 229 YVANGKLQLALPMEKVKETSKKSLMRTVESWLRKERDEERKRRKERKANGLPEFETRTEE 288
Query: 121 SRVVGGDPARAPVRVTNNGWAAGRLHSTIA-GEENFLLSSNRVLR 164
SRV G P R+ NNGWAAGRLHST+A GE+NF+LSSNRVLR
Sbjct: 289 SRVTGD-----PARMLNNGWAAGRLHSTVAGGEDNFVLSSNRVLR 328
>gb|AAP12850.1| At5g44110 [Arabidopsis thaliana] gi|9759512|dbj|BAB10978.1|
NBD-like protein [Arabidopsis thaliana]
gi|15240093|ref|NP_199224.1| ABC transporter family
protein [Arabidopsis thaliana] gi|4427003|gb|AAD20643.1|
NBD-like protein [Arabidopsis thaliana]
Length = 282
Score = 122 bits (306), Expect = 4e-27
Identities = 55/106 (51%), Positives = 83/106 (77%), Gaps = 1/106 (0%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL PF+VLLLDE+TVDLDV+AR DLL+F ++EC++RGATI+YATHIFDGLE W +++
Sbjct: 160 MGLLHPFKVLLLDEVTVDLDVVARMDLLEFFKEECEQRGATIVYATHIFDGLETWASHLA 219
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQRKE 105
Y+ G L+L+ ++++K+ + +L+ VE WLR E +++ +K+
Sbjct: 220 YINGGELKLSAKLDEIKDLKTSPNLLSVVEAWLRSETKVEKKTKKK 265
>gb|AAM64851.1| NBD-like protein [Arabidopsis thaliana]
Length = 282
Score = 120 bits (300), Expect = 2e-26
Identities = 54/104 (51%), Positives = 81/104 (76%), Gaps = 1/104 (0%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL PF+VLLLDE+TVDLDV+AR DLL+F ++EC++RGATI+YATHIFDGLE W +++
Sbjct: 160 MGLLHPFKVLLLDEVTVDLDVVARMDLLEFFKEECEQRGATIVYATHIFDGLETWASHLA 219
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQR 103
Y+ G L+L+ ++++K+ + +L+ VE WLR E +++ +
Sbjct: 220 YINGGELKLSAKLDEIKDLKTSPNLLSVVEAWLRSETKVEKKTK 263
>ref|NP_563694.1| ABC transporter family protein [Arabidopsis thaliana]
Length = 290
Score = 116 bits (291), Expect = 2e-25
Identities = 56/105 (53%), Positives = 77/105 (73%), Gaps = 1/105 (0%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL PF+VLLLDE+TVDLDV+AR DLL+F ++ECD+RGATI+YATHIFDGLE W T++
Sbjct: 161 MGLLHPFKVLLLDEVTVDLDVVARMDLLEFFKEECDQRGATIVYATHIFDGLETWATHLA 220
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQRK 104
Y+ G L + ++E + +L+ VE WLR E ++++K
Sbjct: 221 YIQDGELNRLSKMTDIEELKTSPNLLSVVESWLRSEIKLVKKKKK 265
>pir||G86169 hypothetical protein [imported] - Arabidopsis thaliana
gi|4204313|gb|AAD10694.1| Hypothetical protein
[Arabidopsis thaliana]
Length = 580
Score = 116 bits (291), Expect = 2e-25
Identities = 56/105 (53%), Positives = 77/105 (73%), Gaps = 1/105 (0%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL PF+VLLLDE+TVDLDV+AR DLL+F ++ECD+RGATI+YATHIFDGLE W T++
Sbjct: 451 MGLLHPFKVLLLDEVTVDLDVVARMDLLEFFKEECDQRGATIVYATHIFDGLETWATHLA 510
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKERDEDRRQRK 104
Y+ G L + ++E + +L+ VE WLR E ++++K
Sbjct: 511 YIQDGELNRLSKMTDIEELKTSPNLLSVVESWLRSEIKLVKKKKK 555
>emb|CAB11251.1| SPAC20G4.01 [Schizosaccharomyces pombe] gi|7522450|pir||T38115
probable ATP-dependent transporter - fission yeast
(Schizosaccharomyces pombe) (fragment)
Length = 255
Score = 114 bits (284), Expect = 1e-24
Identities = 61/113 (53%), Positives = 80/113 (69%), Gaps = 7/113 (6%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL+PF+VLLLDE+TVDLDVLARADLL FL++E + R ATI+YATHIFDGL +WPT++V
Sbjct: 116 MGLLRPFEVLLLDEVTVDLDVLARADLLNFLQEETEVRNATIVYATHIFDGLAEWPTHLV 175
Query: 61 YVAHGRLELAMPIEKV------KETSKLSLMRTVEVWLRKERDEDRRQRKERK 107
+++ GR+ PI K T +L+ T WL KE ++R R+E K
Sbjct: 176 HLSLGRIVDYGPISKFGALMTRSSTGNSALLETCLEWL-KEDKKNRGTREEEK 227
>ref|XP_507035.1| PREDICTED OJ1116_E04.3 gene product [Oryza sativa (japonica
cultivar-group)] gi|50915734|ref|XP_468331.1| putative
ATP-dependent transporter [Oryza sativa (japonica
cultivar-group)] gi|47847809|dbj|BAD21584.1| putative
ATP-dependent transporter [Oryza sativa (japonica
cultivar-group)]
Length = 292
Score = 111 bits (278), Expect = 6e-24
Identities = 55/96 (57%), Positives = 69/96 (71%), Gaps = 1/96 (1%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL P++VLLLDEITVDLDV+ R DLL F ++EC++R ATI+YATHIFDGLE W T+I
Sbjct: 161 MGLLHPYKVLLLDEITVDLDVVTRMDLLDFFKEECEQREATIVYATHIFDGLESWATDIA 220
Query: 61 YVAHGRLELAMPIEKVKE-TSKLSLMRTVEVWLRKE 95
Y+ G L + V+E S +L+ VE WLR E
Sbjct: 221 YIQEGELRKSAKYSDVEELKSAKNLLSVVESWLRSE 256
>gb|AAS53523.1| AFR152Cp [Ashbya gossypii ATCC 10895] gi|45198670|ref|NP_985699.1|
AFR152Cp [Eremothecium gossypii]
Length = 270
Score = 100 bits (249), Expect = 1e-20
Identities = 46/67 (68%), Positives = 56/67 (82%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKPF++LLLDE+TVDLDV+AR LL FLR E + R +IIYATHIFDGL DWP +V
Sbjct: 152 MGLLKPFKLLLLDEVTVDLDVVARTKLLHFLRLETERRECSIIYATHIFDGLADWPDRVV 211
Query: 61 YVAHGRL 67
++AHGR+
Sbjct: 212 HIAHGRV 218
>gb|EAL20556.1| hypothetical protein CNBE4760 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57227416|gb|AAW43875.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58268052|ref|XP_571182.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 268
Score = 95.1 bits (235), Expect = 6e-19
Identities = 56/121 (46%), Positives = 75/121 (61%), Gaps = 4/121 (3%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGL+ + VLLLDE+TVDLDVL RADL+ FL E + RGATI+YATHIFDGL+ +PT I
Sbjct: 147 MGLMGEWDVLLLDEVTVDLDVLVRADLIDFLISESETRGATIVYATHIFDGLKKFPTKIC 206
Query: 61 YVAHG---RLELAMPIEKVKETSKLSLMRTVEVWLRKERDEDRRQRKERKAAGLPEFDKQ 117
++ G R +A P K + +L WLR++RD R++ E+ P+ D
Sbjct: 207 HIQLGSTPRGLIAWP-PADKGLQEETLFELALGWLREDRDLRRQKENEKGRTRGPKTDVS 265
Query: 118 V 118
V
Sbjct: 266 V 266
>gb|EAA57852.1| hypothetical protein AN6512.2 [Aspergillus nidulans FGSC A4]
gi|67540684|ref|XP_664116.1| hypothetical protein
AN6512_2 [Aspergillus nidulans FGSC A4]
gi|49098378|ref|XP_410649.1| hypothetical protein
AN6512.2 [Aspergillus nidulans FGSC A4]
Length = 230
Score = 94.0 bits (232), Expect = 1e-18
Identities = 56/115 (48%), Positives = 74/115 (63%), Gaps = 9/115 (7%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL+P+QVLLLDEITVDLD+L+R++ L FL++E + R TI+YATHI D L WPT++V
Sbjct: 100 MGLLRPWQVLLLDEITVDLDLLSRSNFLSFLKRETETRPCTIVYATHILDNLAHWPTHLV 159
Query: 61 YVAHGRLELAMPIEKVK----ETSKLS-LMRTVEVWLRKERDEDRRQRKERKAAG 110
++ G + IEK K ETS+ S L V WL+ ED + R R G
Sbjct: 160 HMHLGNVRQWGAIEKFKEEAPETSENSQLGEIVLKWLK----EDLQARGPRNVRG 210
>gb|EAL85531.1| ABC transporter, putative [Aspergillus fumigatus Af293]
Length = 280
Score = 93.6 bits (231), Expect = 2e-18
Identities = 50/100 (50%), Positives = 71/100 (71%), Gaps = 5/100 (5%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL+P+QVLLLDEITVDLD+L+R++ L FL++E + R TI+YATHI D L WPT++V
Sbjct: 149 MGLLRPWQVLLLDEITVDLDLLSRSNFLSFLKRETETRPCTIVYATHILDNLSQWPTHLV 208
Query: 61 YVAHGRLELAMPIE----KVKETSKLS-LMRTVEVWLRKE 95
++ G + P+E +V ETS+ S L V WL+++
Sbjct: 209 HMHLGNVRQWGPMENFKSEVTETSENSQLGELVLKWLKED 248
>gb|EAK84383.1| hypothetical protein UM03153.1 [Ustilago maydis 521]
gi|49073041|ref|XP_400768.1| hypothetical protein
UM03153.1 [Ustilago maydis 521]
Length = 555
Score = 93.2 bits (230), Expect = 2e-18
Identities = 55/125 (44%), Positives = 77/125 (61%), Gaps = 21/125 (16%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGL++P+ +LLLDE+TVDLDV RADLL FL++E +ER ATIIYATHIFDGL+ +PT++
Sbjct: 403 MGLMEPWDLLLLDEVTVDLDVQVRADLLDFLQQETEERHATIIYATHIFDGLQSFPTHLA 462
Query: 61 YVAHGRLELAMPIE----------------KVKETSKLSLMRTVEV-WLRKERDEDRRQR 103
++ G +PI+ +T LS + V + WLR ED+ R
Sbjct: 463 HMRLGTTTTTLPIDWPATNAADIAALPPTAAADKTRDLSPLLNVSLAWLR----EDKVVR 518
Query: 104 KERKA 108
E++A
Sbjct: 519 AEQEA 523
>emb|CAG81281.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50551231|ref|XP_503089.1| hypothetical protein
[Yarrowia lipolytica]
Length = 253
Score = 90.5 bits (223), Expect = 2e-17
Identities = 40/67 (59%), Positives = 54/67 (79%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLL P++ LLLDE+TVDLDVLAR +LL FL++E + R ATI+YATHIFDGL WPT++
Sbjct: 141 MGLLVPWETLLLDEVTVDLDVLARTNLLNFLKEETETRQATIVYATHIFDGLNQWPTHVA 200
Query: 61 YVAHGRL 67
+++ +
Sbjct: 201 HMSQSTM 207
>ref|NP_116625.1| Part of the evolutionarily-conserved CCR4-NOT transcriptional
regulatory complex involved in controlling mRNA
initiation, elongation, and degradation; putative ABC
ATPase; interacts with Ssn2p, Ssn3p, and Ssn8p; Caf16p
[Saccharomyces cerevisiae]
gi|1175936|sp|P43569|YFC8_YEAST Probable ATP-dependent
transporter YFL028C gi|836726|dbj|BAA09210.1| YFL028C
[Saccharomyces cerevisiae]
Length = 289
Score = 89.4 bits (220), Expect = 3e-17
Identities = 42/82 (51%), Positives = 61/82 (74%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKP++VLLLDE+TVDLDV+ARA LL+FL+ E + R +++YATHIFDGL WP +
Sbjct: 161 MGLLKPWRVLLLDEVTVDLDVIARARLLEFLKWETETRRCSVVYATHIFDGLAKWPNQVY 220
Query: 61 YVAHGRLELAMPIEKVKETSKL 82
++ G++ + +K E S++
Sbjct: 221 HMKSGKIVDNLDYQKDVEFSEV 242
>gb|EAA53770.1| hypothetical protein MG09520.4 [Magnaporthe grisea 70-15]
gi|39961344|ref|XP_364675.1| hypothetical protein
MG09520.4 [Magnaporthe grisea 70-15]
Length = 290
Score = 88.6 bits (218), Expect = 6e-17
Identities = 55/132 (41%), Positives = 77/132 (57%), Gaps = 9/132 (6%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGL++P+ VLLLDEITVDLDVL+R+ L +L++E + R AT++YATHI D L WPT++V
Sbjct: 150 MGLVRPWTVLLLDEITVDLDVLSRSGFLDWLKRETESRDATVVYATHILDNLHAWPTHLV 209
Query: 61 YVAHGRLELAMPIEKVKETSKLS------LMRTVEVWLRKERDEDRRQRKERKAAGLPEF 114
++ G ++ P E S L V WLR++ +DR R + K A PE
Sbjct: 210 HMHLGTVKEWGPAADFLEAQGASKDGNSQLGHLVLGWLREDL-KDRGPRSQHKRA--PEG 266
Query: 115 DKQVDGSRVVGG 126
G +GG
Sbjct: 267 KSYGAGGIELGG 278
>emb|CAG61721.1| unnamed protein product [Candida glabrata CBS138]
gi|50292651|ref|XP_448758.1| unnamed protein product
[Candida glabrata]
Length = 294
Score = 84.0 bits (206), Expect = 1e-15
Identities = 38/67 (56%), Positives = 52/67 (76%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKP++VLLLDE+TVDLDV+AR LL FL+ E R +++YATHIFDGL WP ++
Sbjct: 166 MGLLKPWRVLLLDEVTVDLDVIARMRLLDFLKWETTNRKCSVVYATHIFDGLAAWPDYVL 225
Query: 61 YVAHGRL 67
++ G++
Sbjct: 226 HMQSGKI 232
>emb|CAG98189.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50310917|ref|XP_455481.1| unnamed protein product
[Kluyveromyces lactis]
Length = 273
Score = 83.6 bits (205), Expect = 2e-15
Identities = 38/75 (50%), Positives = 54/75 (71%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGLLKPF+VLLLDE+TVDLDV+AR L +FL +E +R +++YATHIFDGL W I+
Sbjct: 151 MGLLKPFKVLLLDEVTVDLDVVARDRLFQFLDRETRDRQCSVVYATHIFDGLAPWADEII 210
Query: 61 YVAHGRLELAMPIEK 75
++ G + + ++
Sbjct: 211 HLQDGTITRTLDYQR 225
>emb|CAD71023.1| related to ATP-binding cassette (ABC) transporter [Neurospora
crassa] gi|32405648|ref|XP_323437.1| hypothetical
protein [Neurospora crassa] gi|28922388|gb|EAA31623.1|
hypothetical protein [Neurospora crassa]
Length = 297
Score = 82.8 bits (203), Expect = 3e-15
Identities = 50/124 (40%), Positives = 70/124 (56%), Gaps = 13/124 (10%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGLEDWPTNIV 60
MGL++P+ VLLLDEITVDLDV +RA L +LR+E + R T++YATHI D L WPT++V
Sbjct: 147 MGLIRPWTVLLLDEITVDLDVWSRAQFLGWLRRETETRECTVVYATHILDNLASWPTHLV 206
Query: 61 YVAHGRL-------ELAMPIEKVKE-----TSKLSLMRTVEVWLRKE-RDEDRRQRKERK 107
++ G + E+ +E K T L V WLR + ++ R + R
Sbjct: 207 HMHLGTVKGWGKAEEMLKEVEDGKNVIEGVTGNSRLGELVLRWLRDDLKERGPRSQHRRG 266
Query: 108 AAGL 111
GL
Sbjct: 267 PEGL 270
>emb|CAG89280.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50424673|ref|XP_460926.1| unnamed protein product
[Debaryomyces hansenii]
Length = 321
Score = 82.4 bits (202), Expect = 4e-15
Identities = 39/66 (59%), Positives = 51/66 (77%), Gaps = 1/66 (1%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGL-EDWPTNI 59
MGLLKP+++LLLDE+TVDLDV+ R DLL FL+KEC R +IYATHIFDGL + W +
Sbjct: 161 MGLLKPWKLLLLDEVTVDLDVVVRQDLLNFLKKECRLRNCCVIYATHIFDGLGKHWCDRL 220
Query: 60 VYVAHG 65
+++ G
Sbjct: 221 IHLNEG 226
>gb|EAK92697.1| potential CCR4-NOT complex associated factor Caf16p [Candida
albicans SC5314] gi|46433220|gb|EAK92668.1| potential
CCR4-NOT complex associated factor Caf16p [Candida
albicans SC5314]
Length = 320
Score = 82.0 bits (201), Expect = 5e-15
Identities = 35/66 (53%), Positives = 52/66 (78%), Gaps = 1/66 (1%)
Query: 1 MGLLKPFQVLLLDEITVDLDVLARADLLKFLRKECDERGATIIYATHIFDGL-EDWPTNI 59
MGL+KP+++LLLDE+T+DLDV+ R+ LL +L++EC ER T++YATHIFDGL W +
Sbjct: 161 MGLVKPWKLLLLDEVTIDLDVVVRSKLLNYLKQECQERNCTVVYATHIFDGLGHKWCDRV 220
Query: 60 VYVAHG 65
+++ G
Sbjct: 221 IHIDAG 226
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.139 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 270,134,882
Number of Sequences: 2540612
Number of extensions: 10854669
Number of successful extensions: 47300
Number of sequences better than 10.0: 2644
Number of HSP's better than 10.0 without gapping: 1488
Number of HSP's successfully gapped in prelim test: 1156
Number of HSP's that attempted gapping in prelim test: 45337
Number of HSP's gapped (non-prelim): 2887
length of query: 165
length of database: 863,360,394
effective HSP length: 118
effective length of query: 47
effective length of database: 563,568,178
effective search space: 26487704366
effective search space used: 26487704366
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC147011.10


