

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147000.4 + phase: 1 /pseudo
(187 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
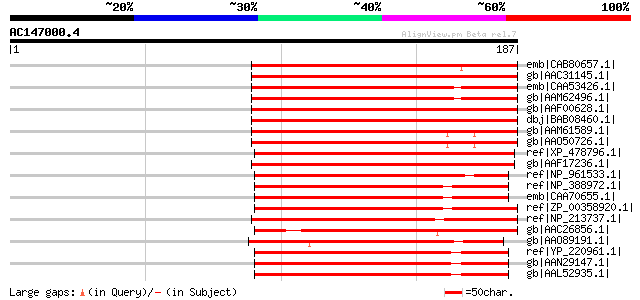
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB80657.1| adenosine-5'-phosphosulfate-kinase [Arabidopsis ... 156 3e-37
gb|AAC31145.1| adenosine-5'-phosphosulfate-kinase [Catharanthus ... 156 3e-37
emb|CAA53426.1| APS kinase [Arabidopsis thaliana] gi|22135773|gb... 152 6e-36
gb|AAM62496.1| putative adenosine phosphosulfate kinase [Arabido... 152 6e-36
gb|AAF00628.1| putative adenylylsulfate kinase [Arabidopsis thal... 142 6e-33
dbj|BAB08460.1| adenylylsulfate kinase-like protein [Arabidopsis... 140 1e-32
gb|AAM61589.1| adenylylsulfate kinase-like protein [Arabidopsis ... 138 7e-32
gb|AAO50726.1| putative adenylylsulfate kinase [Arabidopsis thal... 138 7e-32
ref|XP_478796.1| putative adenosine-5'-phosphosulfate kinase [Or... 135 8e-31
gb|AAF17236.1| adenosine-5'-phosphosulfate kinase [Zea mays] 130 2e-29
ref|NP_961533.1| hypothetical protein MAP2599c [Mycobacterium av... 112 4e-24
ref|NP_388972.1| hypothetical protein BSU10910 [Bacillus subtili... 103 2e-21
emb|CAA70655.1| YisZ [Bacillus subtilis] 103 2e-21
ref|ZP_00358920.1| COG0529: Adenylylsulfate kinase and related k... 100 2e-20
ref|NP_213737.1| sulfate adenylyltransferase [Aquifex aeolicus V... 99 6e-20
gb|AAC26856.1| 5'-adenylylsulfate kinase [Enteromorpha intestina... 98 1e-19
gb|AAO89191.1| CysNC [Rhodococcus sp. DS7] 98 1e-19
ref|YP_220961.1| CysNC, sulfate adenylate transferase, subunit 1... 97 2e-19
gb|AAN29147.1| sulfate adenylate transferase, subunit 1/adenylyl... 97 2e-19
gb|AAL52935.1| SULFATE ADENYLYLTRANSFERASE / ADENYLYLSULFATE KIN... 97 2e-19
>emb|CAB80657.1| adenosine-5'-phosphosulfate-kinase [Arabidopsis thaliana]
gi|4490745|emb|CAB38907.1|
adenosine-5'-phosphosulfate-kinase [Arabidopsis
thaliana] gi|20453397|gb|AAM19937.1| AT4g39940/T5J17_110
[Arabidopsis thaliana] gi|18087563|gb|AAL58913.1|
AT4g39940/T5J17_110 [Arabidopsis thaliana]
gi|15236087|ref|NP_195704.1| adenylylsulfate kinase 2
(AKN2) [Arabidopsis thaliana] gi|2829133|gb|AAC39520.1|
adenosine-5'-phosphosulfate-kinase [Arabidopsis
thaliana] gi|7434263|pir||T06100 adenylyl-sulfate kinase
(EC 2.7.1.25) [validated] - Arabidopsis thaliana
gi|7387808|sp|O49196|KAP2_ARATH Adenylyl-sulfate kinase
2, chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 293
Score = 156 bits (394), Expect = 3e-37
Identities = 71/99 (71%), Positives = 86/99 (86%), Gaps = 1/99 (1%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY++DRDACR+LLP+GDF+EVF+DVPLHVCE+RDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 194 LISPYRRDRDACRSLLPDGDFVEVFMDVPLHVCESRDPKGLYKLARAGKIKGFTGIDDPY 253
Query: 150 EPPCCCEIILQQKGSD-CKSPKDMAETVISYLEKSGHLQ 187
E P CE++L+ G D SP+ MAE +ISYL+ G+L+
Sbjct: 254 EAPVNCEVVLKHTGDDESCSPRQMAENIISYLQNKGYLE 292
>gb|AAC31145.1| adenosine-5'-phosphosulfate-kinase [Catharanthus roseus]
gi|7434264|pir||T08076 adenylyl-sulfate kinase (EC
2.7.1.25) precursor - Madagascar periwinkle
gi|7387809|sp|O49204|KAPS_CATRO Adenylyl-sulfate kinase,
chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 312
Score = 156 bits (394), Expect = 3e-37
Identities = 75/98 (76%), Positives = 85/98 (86%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+K DACR+LLPEGDFIEVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 214 LISPYRKPPDACRSLLPEGDFIEVFMDVPLKVCEARDPKGLYKLARAGKIKGFTGIDDPY 273
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP EI+L QK C SP D+A+ VISYLE++G+L+
Sbjct: 274 EPPLKSEIVLHQKLGMCDSPCDLADIVISYLEENGYLK 311
>emb|CAA53426.1| APS kinase [Arabidopsis thaliana] gi|22135773|gb|AAM91043.1|
At2g14750/F26C24.11 [Arabidopsis thaliana]
gi|3252812|gb|AAC24182.1| putative adenosine
phosphosulfate kinase [Arabidopsis thaliana]
gi|15810038|gb|AAL06946.1| At2g14750/F26C24.11
[Arabidopsis thaliana] gi|450235|gb|AAC50035.1| APS
kinase [Arabidopsis thaliana] gi|1575322|gb|AAC50034.1|
APS kinase [Arabidopsis thaliana] gi|1076283|pir||S47640
adenylyl-sulfate kinase (EC 2.7.1.25) precursor -
Arabidopsis thaliana gi|15225999|ref|NP_179082.1|
adenylylsulfate kinase 1 (AKN1) [Arabidopsis thaliana]
gi|7387811|sp|Q43295|KAP1_ARATH Adenylyl-sulfate kinase
1, chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 276
Score = 152 bits (383), Expect = 6e-36
Identities = 73/98 (74%), Positives = 83/98 (84%), Gaps = 2/98 (2%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ DRDACR+LLPEGDF+EVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 180 LISPYRTDRDACRSLLPEGDFVEVFMDVPLSVCEARDPKGLYKLARAGKIKGFTGIDDPY 239
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI L ++G SP +MAE V+ YL+ G+LQ
Sbjct: 240 EPPLNCEISLGREGG--TSPIEMAEKVVGYLDNKGYLQ 275
>gb|AAM62496.1| putative adenosine phosphosulfate kinase [Arabidopsis thaliana]
Length = 276
Score = 152 bits (383), Expect = 6e-36
Identities = 73/98 (74%), Positives = 83/98 (84%), Gaps = 2/98 (2%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ DRDACR+LLPEGDF+EVF+DVPL VCEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 180 LISPYRTDRDACRSLLPEGDFVEVFMDVPLSVCEARDPKGLYKLARAGKIKGFTGIDDPY 239
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP CEI L ++G SP +MAE V+ YL+ G+LQ
Sbjct: 240 EPPLNCEISLGREGG--TSPIEMAEKVVGYLDNKGYLQ 275
>gb|AAF00628.1| putative adenylylsulfate kinase [Arabidopsis thaliana]
gi|15228666|ref|NP_187040.1| adenylylsulfate kinase,
putative [Arabidopsis thaliana]
Length = 208
Score = 142 bits (357), Expect = 6e-33
Identities = 68/98 (69%), Positives = 78/98 (79%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+KDRDACR ++ FIEVF+++ L +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 109 LISPYRKDRDACREMIQNSSFIEVFMNMSLQLCEARDPKGLYKLARAGKIKGFTGIDDPY 168
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
E P CEI L++K +C SP MAE VISYLE G LQ
Sbjct: 169 ESPLNCEIELKEKEGECPSPVAMAEEVISYLEDKGFLQ 206
>dbj|BAB08460.1| adenylylsulfate kinase-like protein [Arabidopsis thaliana]
Length = 290
Score = 140 bits (354), Expect = 1e-32
Identities = 66/98 (67%), Positives = 81/98 (82%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 183 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 242
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
E P CE+ + S S +MA+ V+SYL+++G+L+
Sbjct: 243 EAPLDCEVHIISNFSSSSSLCEMADIVVSYLDQNGYLK 280
>gb|AAM61589.1| adenylylsulfate kinase-like protein [Arabidopsis thaliana]
Length = 305
Score = 138 bits (348), Expect = 7e-32
Identities = 69/113 (61%), Positives = 84/113 (74%), Gaps = 15/113 (13%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 183 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 242
Query: 150 EPPCCCEIILQ--------QKGSDCKSPK-------DMAETVISYLEKSGHLQ 187
E P CEI++Q S SP +MA+ V+SYL+++G+L+
Sbjct: 243 EAPLDCEIVIQNSRDKGLSSSSSSSSSPSSSSSSLCEMADIVVSYLDQNGYLK 295
>gb|AAO50726.1| putative adenylylsulfate kinase [Arabidopsis thaliana]
gi|28393175|gb|AAO42019.1| putative adenylylsulfate
kinase [Arabidopsis thaliana]
gi|18425199|ref|NP_569050.1| adenylylsulfate kinase,
putative [Arabidopsis thaliana]
Length = 310
Score = 138 bits (348), Expect = 7e-32
Identities = 69/113 (61%), Positives = 84/113 (74%), Gaps = 15/113 (13%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISPY+ +R ACRALLP+GDFIEVF+DVPLHVCEARDPKGLYK ARAGKIKGFTG+DDPY
Sbjct: 188 LISPYRIERAACRALLPQGDFIEVFMDVPLHVCEARDPKGLYKRARAGKIKGFTGVDDPY 247
Query: 150 EPPCCCEIILQ--------QKGSDCKSPK-------DMAETVISYLEKSGHLQ 187
E P CEI++Q S SP +MA+ V+SYL+++G+L+
Sbjct: 248 EAPLDCEIVIQNSRDKGLSSSSSSSSSPSSSSSSLCEMADIVVSYLDQNGYLK 300
>ref|XP_478796.1| putative adenosine-5'-phosphosulfate kinase [Oryza sativa (japonica
cultivar-group)] gi|34393551|dbj|BAC83149.1| putative
adenosine-5'-phosphosulfate kinase [Oryza sativa
(japonica cultivar-group)]
Length = 345
Score = 135 bits (339), Expect = 8e-31
Identities = 62/96 (64%), Positives = 78/96 (80%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISPY++DR++CRALL +G FIEVF+++PL +CE+RDPKGLYKLARAGKIKGFTGIDDPYE
Sbjct: 248 ISPYRRDRESCRALLSDGSFIEVFLNMPLELCESRDPKGLYKLARAGKIKGFTGIDDPYE 307
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHL 186
P EI +++ C SP DMA V++YLE+ G L
Sbjct: 308 SPLNSEIEIKEVDGVCPSPSDMAGQVVTYLEEKGFL 343
>gb|AAF17236.1| adenosine-5'-phosphosulfate kinase [Zea mays]
Length = 288
Score = 130 bits (327), Expect = 2e-29
Identities = 60/97 (61%), Positives = 77/97 (78%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
LISP+++DR++CRALL + FIEVF+++ L +CEARDPKGLYKLARAGKIKGFTGIDDPY
Sbjct: 190 LISPHRRDRESCRALLSDSSFIEVFLNMSLELCEARDPKGLYKLARAGKIKGFTGIDDPY 249
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHL 186
E P CEI +++ C P +MA V++YLE+ G L
Sbjct: 250 EAPLNCEIEIKEVDGVCPPPAEMAGQVVTYLEEKGFL 286
>ref|NP_961533.1| hypothetical protein MAP2599c [Mycobacterium avium subsp.
paratuberculosis str. k10] gi|41397055|gb|AAS04916.1|
hypothetical protein MAP2599c [Mycobacterium avium
subsp. paratuberculosis str. k10]
Length = 230
Score = 112 bits (281), Expect = 4e-24
Identities = 56/94 (59%), Positives = 69/94 (72%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISPY +DRDA RA L +GDF E+FID P+ +CE RDPKGLYK ARAG+IKGFTGIDDPYE
Sbjct: 122 ISPYVRDRDAIRATLDDGDFQEIFIDTPIEICEKRDPKGLYKKARAGEIKGFTGIDDPYE 181
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P E+ L D ++ +AE VI++LE+ G
Sbjct: 182 APPRPELRLDGAAKDAET---LAEEVIAHLERVG 212
>ref|NP_388972.1| hypothetical protein BSU10910 [Bacillus subtilis subsp. subtilis
str. 168] gi|2633427|emb|CAB12931.1| yisZ [Bacillus
subtilis subsp. subtilis str. 168]
gi|7434259|pir||A69839 adenylylsulfate kinase homolog
yisZ - Bacillus subtilis
gi|7387594|sp|O06735|CYSC2_BACSU Probable
adenylyl-sulfate kinase (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)
Length = 199
Score = 103 bits (257), Expect = 2e-21
Identities = 50/94 (53%), Positives = 66/94 (70%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP+++DRD RAL P+G+F E+++ PLHVCE RDPKGLYK AR G+IK FTGID PYE
Sbjct: 107 ISPFREDRDMVRALFPKGEFFEIYVKCPLHVCEQRDPKGLYKKARNGEIKHFTGIDSPYE 166
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P + I++ SD S D A+ +I+ L+ G
Sbjct: 167 APLSPDFIIE---SDQTSISDGADLIINALQNRG 197
>emb|CAA70655.1| YisZ [Bacillus subtilis]
Length = 197
Score = 103 bits (257), Expect = 2e-21
Identities = 50/94 (53%), Positives = 66/94 (70%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP+++DRD RAL P+G+F E+++ PLHVCE RDPKGLYK AR G+IK FTGID PYE
Sbjct: 105 ISPFREDRDMVRALFPKGEFFEIYVKCPLHVCEQRDPKGLYKKARNGEIKHFTGIDSPYE 164
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
P + I++ SD S D A+ +I+ L+ G
Sbjct: 165 APLSPDFIIE---SDQTSISDGADLIINALQNRG 195
>ref|ZP_00358920.1| COG0529: Adenylylsulfate kinase and related kinases [Chloroflexus
aurantiacus]
Length = 278
Score = 100 bits (249), Expect = 2e-20
Identities = 48/97 (49%), Positives = 65/97 (66%), Gaps = 3/97 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
+SPY+ R+ CR ++ FIEVF+D PL VCE RD KGLY AR G+++GFTGIDDPYE
Sbjct: 184 VSPYRAARNECRTMIGSDQFIEVFVDTPLEVCEQRDVKGLYAKARRGELRGFTGIDDPYE 243
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
PP E+ L + +P++ A +I YLE+ G L+
Sbjct: 244 PPVNPELTLT---TTDVTPEENARKIIRYLEEKGFLE 277
>ref|NP_213737.1| sulfate adenylyltransferase [Aquifex aeolicus VF5]
gi|2983561|gb|AAC07134.1| sulfate adenylyltransferase
[Aquifex aeolicus VF5] gi|7428530|pir||C70393 probable
adenylyl-sulfate kinase (EC 2.7.1.25) - Aquifex aeolicus
gi|7388241|sp|O67174|SATC_AQUAE Probable bifunctional
SAT/APS kinase [Includes: Sulfate adenylyltransferase
(Sulfate adenylate transferase) (SAT) (ATP-sulfurylase);
Adenylyl-sulfate kinase (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase)]
Length = 546
Score = 99.0 bits (245), Expect = 6e-20
Identities = 46/98 (46%), Positives = 67/98 (67%), Gaps = 3/98 (3%)
Query: 90 LISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPY 149
L+SPY+ R+ R ++ EG FIEVF+D P+ VCE RD KGLYK A+ G IKGFTG+DDPY
Sbjct: 451 LVSPYRSARNQVRNMMEEGKFIEVFVDAPVEVCEERDVKGLYKKAKEGLIKGFTGVDDPY 510
Query: 150 EPPCCCEIILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
EPP E+ + + +P++ A ++ +L+K G ++
Sbjct: 511 EPPVAPEV---RVDTTKLTPEESALKILEFLKKEGFIK 545
>gb|AAC26856.1| 5'-adenylylsulfate kinase [Enteromorpha intestinalis]
Length = 271
Score = 98.2 bits (243), Expect = 1e-19
Identities = 49/99 (49%), Positives = 67/99 (67%), Gaps = 7/99 (7%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
I+P + R+ C GDF+E ++ +P+ +CE RDPKGLYK ARAG +KGFTGIDDPYE
Sbjct: 147 IAPTRPVRERCA-----GDFVECYMKIPIELCEQRDPKGLYKKARAGLMKGFTGIDDPYE 201
Query: 151 PPCCCE--IILQQKGSDCKSPKDMAETVISYLEKSGHLQ 187
P E I ++++GSD SP+ MA+ + YLE G L+
Sbjct: 202 EPLEPELTITVREEGSDMNSPEAMAKQIFDYLEAKGFLK 240
>gb|AAO89191.1| CysNC [Rhodococcus sp. DS7]
Length = 614
Score = 97.8 bits (242), Expect = 1e-19
Identities = 45/96 (46%), Positives = 66/96 (67%), Gaps = 5/96 (5%)
Query: 89 CLISPYQKDRDACRALLPEGD--FIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGID 146
CLISPY++DRD RA F+EV++D P+ CEARDPKG+Y ARAG+IKGFTG+D
Sbjct: 522 CLISPYREDRDRVRAAHEAAGLKFVEVYVDTPIEQCEARDPKGMYAKARAGEIKGFTGVD 581
Query: 147 DPYEPPCCCEIILQQKGSDCKSPKDMAETVISYLEK 182
DPYE P E++++ + +P ++A ++ L++
Sbjct: 582 DPYEAPASAELVIRPEDG---TPTELALRIMEVLDR 614
>ref|YP_220961.1| CysNC, sulfate adenylate transferase, subunit 1/adenylylsulfate
kinase [Brucella abortus biovar 1 str. 9-941]
gi|62195300|gb|AAX73600.1| CysNC, sulfate adenylate
transferase, subunit 1/adenylylsulfate kinase [Brucella
abortus biovar 1 str. 9-941]
Length = 644
Score = 97.4 bits (241), Expect = 2e-19
Identities = 48/94 (51%), Positives = 63/94 (66%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++++RD RA LPEG+FIE+FID P+ C ARDPKGLY A G+IK FTGID PYE
Sbjct: 542 ISPFREERDQVRARLPEGEFIEIFIDTPIEECMARDPKGLYAQALRGEIKAFTGIDSPYE 601
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
PP E+ L G ++ ++ V +YL + G
Sbjct: 602 PPVAPELRLNTAG---RTVDELIAEVENYLAERG 632
>gb|AAN29147.1| sulfate adenylate transferase, subunit 1/adenylylsulfate kinase
[Brucella suis 1330] gi|23501105|ref|NP_697232.1|
sulfate adenylate transferase, subunit 1/adenylylsulfate
kinase [Brucella suis 1330]
Length = 644
Score = 97.4 bits (241), Expect = 2e-19
Identities = 48/94 (51%), Positives = 63/94 (66%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++++RD RA LPEG+FIE+FID P+ C ARDPKGLY A G+IK FTGID PYE
Sbjct: 542 ISPFREERDQVRARLPEGEFIEIFIDTPIEECMARDPKGLYAQALRGEIKAFTGIDSPYE 601
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
PP E+ L G ++ ++ V +YL + G
Sbjct: 602 PPVAPELRLNTAG---RTVDELIAEVENYLAERG 632
>gb|AAL52935.1| SULFATE ADENYLYLTRANSFERASE / ADENYLYLSULFATE KINASE [Brucella
melitensis 16M] gi|17988037|ref|NP_540671.1| SULFATE
ADENYLYLTRANSFERASE / ADENYLYLSULFATE KINASE [Brucella
melitensis 16M] gi|25299444|pir||AD3471 adenylyl-sulfate
kinase (EC 2.7.1.25) [imported] - Brucella melitensis
(strain 16M)
Length = 644
Score = 97.4 bits (241), Expect = 2e-19
Identities = 48/94 (51%), Positives = 63/94 (66%), Gaps = 3/94 (3%)
Query: 91 ISPYQKDRDACRALLPEGDFIEVFIDVPLHVCEARDPKGLYKLARAGKIKGFTGIDDPYE 150
ISP++++RD RA LPEG+FIE+FID P+ C ARDPKGLY A G+IK FTGID PYE
Sbjct: 542 ISPFREERDQVRARLPEGEFIEIFIDTPIEECMARDPKGLYAQALRGEIKAFTGIDSPYE 601
Query: 151 PPCCCEIILQQKGSDCKSPKDMAETVISYLEKSG 184
PP E+ L G ++ ++ V +YL + G
Sbjct: 602 PPVAPELRLNTAG---RTVDELIAEVENYLAERG 632
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.340 0.150 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 303,883,356
Number of Sequences: 2540612
Number of extensions: 11580667
Number of successful extensions: 26026
Number of sequences better than 10.0: 294
Number of HSP's better than 10.0 without gapping: 292
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 25670
Number of HSP's gapped (non-prelim): 301
length of query: 187
length of database: 863,360,394
effective HSP length: 120
effective length of query: 67
effective length of database: 558,486,954
effective search space: 37418625918
effective search space used: 37418625918
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 71 (32.0 bits)
Medicago: description of AC147000.4


