

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146941.4 + phase: 0
(328 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
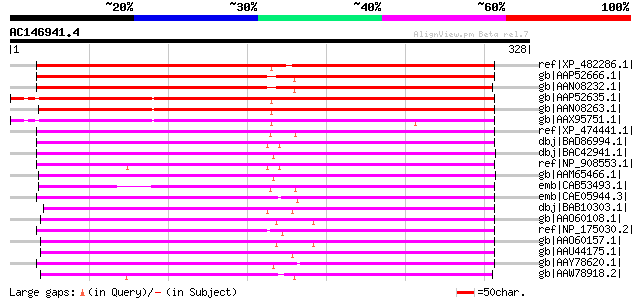
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_482286.1| putative nodulin MtN21 [Oryza sativa (japonica ... 255 1e-66
gb|AAP52666.1| nodulin-like protein [Oryza sativa (japonica cult... 254 2e-66
gb|AAN08232.1| nodulin-like protein [Oryza sativa (japonica cult... 254 2e-66
gb|AAP52635.1| nodulin-like protein [Oryza sativa (japonica cult... 243 4e-63
gb|AAN08263.1| nodulin-like protein [Oryza sativa (japonica cult... 240 4e-62
gb|AAX95751.1| Integral membrane protein DUF6, putative [Oryza s... 234 3e-60
ref|XP_474441.1| OSJNBa0070M12.19 [Oryza sativa (japonica cultiv... 189 7e-47
dbj|BAD86994.1| putative MtN21 [Oryza sativa (japonica cultivar-... 183 7e-45
dbj|BAC42941.1| putative nodulin protein N21 [Arabidopsis thalia... 180 4e-44
ref|NP_908553.1| putative MtN21 [Oryza sativa (japonica cultivar... 179 1e-43
gb|AAM65466.1| putative nodulin protein, N21 [Arabidopsis thaliana] 178 2e-43
emb|CAB53493.1| CAA303720.1 protein [Oryza sativa] 176 8e-43
emb|CAE05944.3| OSJNBb0088C09.3 [Oryza sativa (japonica cultivar... 173 7e-42
dbj|BAB10303.1| nodulin-like protein [Arabidopsis thaliana] gi|1... 169 8e-41
gb|AAO60108.1| nodulin-like protein [Gossypium hirsutum] 169 1e-40
ref|NP_175030.2| integral membrane family protein / nodulin MtN2... 169 1e-40
gb|AAO60157.1| putative nodulin protein [Gossypium hirsutum] 168 2e-40
gb|AAU44175.1| unknown protein [Oryza sativa (japonica cultivar-... 167 4e-40
gb|AAY78620.1| nodulin MtN21 family protein [Arabidopsis thaliana] 167 5e-40
gb|AAW78918.2| nodulin-like protein [Triticum aestivum] 165 1e-39
>ref|XP_482286.1| putative nodulin MtN21 [Oryza sativa (japonica cultivar-group)]
gi|37573001|dbj|BAC98693.1| putative nodulin MtN21
[Oryza sativa (japonica cultivar-group)]
Length = 347
Score = 255 bits (652), Expect = 1e-66
Identities = 118/293 (40%), Positives = 191/293 (64%), Gaps = 7/293 (2%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
SM+ VQL +TG +LS++ + G IF L +Y A V + P A+ FERG+ ++
Sbjct: 31 SMVLVQLFITGQILLSKVSIGGGMLIFVLLAYNSFFAVVFLLPFALIFERGKWRDMDWGA 90
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
IFLN F+G S+ M LYYYG++DT+++Y++ FLN+ P+ TF+ +++FR+E + +
Sbjct: 91 FGWIFLNAFIGYSVPMSLYYYGLKDTTSSYSVIFLNITPLFTFILSLMFRLEAFKLRSIP 150
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFY----IAQYHSFHSVAAHKTQMLRGTLFLIGACF 193
G K +L + GT+ LYKGK + I Q+ + H A T LRGT+ L+G+ F
Sbjct: 151 GVLKIASILLSIGGTMLISLYKGKSLHLWDSIIQHQNEHKSA---TNQLRGTILLVGSSF 207
Query: 194 SYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITIL 253
+++ WF +Q K+++V+P +YW M+ C++ Q+A++G + K AW L WNL L+TI+
Sbjct: 208 TFACWFLIQSKILKVYPYKYWSSMVTCLVGVFQTALVGIILRRDKSAWELGWNLNLVTIV 267
Query: 254 YSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
Y+GAL+TA + L SWA+T +GPTYP+MF+PL++VF ++++LG +T+G+
Sbjct: 268 YTGALATAGKYILNSWAITKRGPTYPTMFSPLSVVFTVVLDSVLLGNDITIGS 320
>gb|AAP52666.1| nodulin-like protein [Oryza sativa (japonica cultivar-group)]
gi|37532154|ref|NP_920379.1| nodulin-like protein [Oryza
sativa (japonica cultivar-group)]
gi|23266295|gb|AAN16334.1| nodulin-like protein [Oryza
sativa (japonica cultivar-group)]
Length = 330
Score = 254 bits (650), Expect = 2e-66
Identities = 118/297 (39%), Positives = 190/297 (63%), Gaps = 13/297 (4%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
SM+ VQL +TG +LS++ + G FIFAL +YR L AV + P A+ FERG+ ++
Sbjct: 20 SMVLVQLFITGMIMLSKVSIGGGMFIFALLAYRSLFGAVFILPFALIFERGKWRDMDWRA 79
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
IF N F+G ++ M LY+YG++DT+A+YA+ F+N+IP+ F+ +++FR+E I +
Sbjct: 80 FGWIFFNAFIGYAVPMSLYFYGLKDTTASYAVIFINIIPLFAFILSLMFRLETFEIGSIV 139
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHK--------TQMLRGTLFLI 189
G K VG +L V GT+ LYKGK H ++S+ H+ T LRGT+ L+
Sbjct: 140 GVLKIVGVLLSVGGTMLVSLYKGKSL-----HLWNSILQHQNEPATKTATNQLRGTILLV 194
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
+ F+Y+ W+ +Q K+++V+P +YW M+ C++ Q A +G + K AW+L W+L L
Sbjct: 195 ASSFAYACWYLVQSKVLKVYPYKYWSSMITCLVGGFQVAFVGIILRRHKSAWKLGWDLNL 254
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+T++YSGAL+T + L SW + +GP YP MFNPL++VF +++++G+ +TVG+
Sbjct: 255 VTVVYSGALATGGKYSLNSWVVAKRGPAYPPMFNPLSVVFTVVLDSVLMGDDVTVGS 311
>gb|AAN08232.1| nodulin-like protein [Oryza sativa (japonica cultivar-group)]
Length = 316
Score = 254 bits (649), Expect = 2e-66
Identities = 118/296 (39%), Positives = 189/296 (62%), Gaps = 13/296 (4%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
SM+ VQL +TG +LS++ + G FIFAL +YR L AV + P A+ FERG+ ++
Sbjct: 20 SMVLVQLFITGMIMLSKVSIGGGMFIFALLAYRSLFGAVFILPFALIFERGKWRDMDWRA 79
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
IF N F+G ++ M LY+YG++DT+A+YA+ F+N+IP+ F+ +++FR+E I +
Sbjct: 80 FGWIFFNAFIGYAVPMSLYFYGLKDTTASYAVIFINIIPLFAFILSLMFRLETFEIGSIV 139
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHK--------TQMLRGTLFLI 189
G K VG +L V GT+ LYKGK H ++S+ H+ T LRGT+ L+
Sbjct: 140 GVLKIVGVLLSVGGTMLVSLYKGKSL-----HLWNSILQHQNEPATKTATNQLRGTILLV 194
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
+ F+Y+ W+ +Q K+++V+P +YW M+ C++ Q A +G + K AW+L W+L L
Sbjct: 195 ASSFAYACWYLVQSKVLKVYPYKYWSSMITCLVGGFQVAFVGIILRRHKSAWKLGWDLNL 254
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
+T++YSGAL+T + L SW + +GP YP MFNPL++VF +++++G+ +TVG
Sbjct: 255 VTVVYSGALATGGKYSLNSWVVAKRGPAYPPMFNPLSVVFTVVLDSVLMGDDVTVG 310
>gb|AAP52635.1| nodulin-like protein [Oryza sativa (japonica cultivar-group)]
gi|37532092|ref|NP_920348.1| nodulin-like protein [Oryza
sativa (japonica cultivar-group)]
Length = 356
Score = 243 bits (621), Expect = 4e-63
Identities = 121/311 (38%), Positives = 190/311 (60%), Gaps = 10/311 (3%)
Query: 1 MNFNDVKEWFVLSSILWSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAP 60
M + +KEW L +I M+ +Q+ G+ +L ++++ G F+ L +YR L+ AV V P
Sbjct: 2 MTSSSLKEW--LPAIF--MVMLQIFTAGSLMLVKVVVDSGLFVCTLLTYRYLLGAVLVVP 57
Query: 61 LAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTF 120
A+ FE+G+ K + IF + VG + V LYY G+ DTS YA+NF N++PI F
Sbjct: 58 FAVSFEKGKLKELKLKAFIWIFTSALVGFT-VPGLYYIGLGDTSPGYAINFYNIVPIAAF 116
Query: 121 LTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFY-----IAQYHSFHSVA 175
+ A+LFR E LN+ + G K VGA++CV GT+ LYKGK + I YH +
Sbjct: 117 ILAVLFRKEPLNMRSIVGIIKVVGALVCVGGTIIISLYKGKVLHLWPTNIIGYHPSKAAT 176
Query: 176 AHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD 235
A +RGT+ L +C S + W+ +Q ++++VFP +YW + C + IQ A+IG ++
Sbjct: 177 AFGHHHIRGTILLAISCLSLAVWYTVQAQMLKVFPYKYWSTVATCFVGCIQMAIIGVAMN 236
Query: 236 SSKEAWRLEWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEA 295
K W+L+WN+ L+TI+YS L+TAA F + SW +T +GPTYPSMF ++++F ++
Sbjct: 237 REKATWKLKWNMSLLTIIYSAILNTAAKFVMISWVVTQRGPTYPSMFCAVSVLFTTILDS 296
Query: 296 MILGEPLTVGT 306
++LG L+VG+
Sbjct: 297 LLLGHDLSVGS 307
>gb|AAN08263.1| nodulin-like protein [Oryza sativa (japonica cultivar-group)]
Length = 341
Score = 240 bits (613), Expect = 4e-62
Identities = 115/293 (39%), Positives = 181/293 (61%), Gaps = 6/293 (2%)
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
M+ +Q+ G+ +L ++++ G F+ L +YR L+ AV V P A+ FE+G+ K +
Sbjct: 1 MVMLQIFTAGSLMLVKVVVDSGLFVCTLLTYRYLLGAVLVVPFAVSFEKGKLKELKLKAF 60
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNG 138
IF + VG + V LYY G+ DTS YA+NF N++PI F+ A+LFR E LN+ + G
Sbjct: 61 IWIFTSALVGFT-VPGLYYIGLGDTSPGYAINFYNIVPIAAFILAVLFRKEPLNMRSIVG 119
Query: 139 RAKCVGAILCVAGTLAARLYKGKEFY-----IAQYHSFHSVAAHKTQMLRGTLFLIGACF 193
K VGA++CV GT+ LYKGK + I YH + A +RGT+ L +C
Sbjct: 120 IIKVVGALVCVGGTIIISLYKGKVLHLWPTNIIGYHPSKAATAFGHHHIRGTILLAISCL 179
Query: 194 SYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITIL 253
S + W+ +Q ++++VFP +YW + C + IQ A+IG ++ K W+L+WN+ L+TI+
Sbjct: 180 SLAVWYTVQAQMLKVFPYKYWSTVATCFVGCIQMAIIGVAMNREKATWKLKWNMSLLTII 239
Query: 254 YSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
YS L+TAA F + SW +T +GPTYPSMF ++++F ++++LG L+VG+
Sbjct: 240 YSAILNTAAKFVMISWVVTQRGPTYPSMFCAVSVLFTTILDSLLLGHDLSVGS 292
>gb|AAX95751.1| Integral membrane protein DUF6, putative [Oryza sativa (japonica
cultivar-group)]
Length = 369
Score = 234 bits (596), Expect = 3e-60
Identities = 121/325 (37%), Positives = 190/325 (58%), Gaps = 24/325 (7%)
Query: 1 MNFNDVKEWFVLSSILWSMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAP 60
M + +KEW L +I M+ +Q+ G+ +L ++++ G F+ L +YR L+ AV V P
Sbjct: 1 MTSSSLKEW--LPAIF--MVMLQIFTAGSLMLVKVVVDSGLFVCTLLTYRYLLGAVLVVP 56
Query: 61 LAIYFERGQPKNFSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTF 120
A+ FE+G+ K + IF + VG + V LYY G+ DTS YA+NF N++PI F
Sbjct: 57 FAVSFEKGKLKELKLKAFIWIFTSALVGFT-VPGLYYIGLGDTSPGYAINFYNIVPIAAF 115
Query: 121 LTAILFRMENLNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFY-----IAQYHSFHSVA 175
+ A+LFR E LN+ + G K VGA++CV GT+ LYKGK + I YH +
Sbjct: 116 ILAVLFRKEPLNMRSIVGIIKVVGALVCVGGTIIISLYKGKVLHLWPTNIIGYHPSKAAT 175
Query: 176 AHKTQMLRGTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVD 235
A +RGT+ L +C S + W+ +Q ++++VFP +YW + C + IQ A+IG ++
Sbjct: 176 AFGHHHIRGTILLAISCLSLAVWYTVQAQMLKVFPYKYWSTVATCFVGCIQMAIIGVAMN 235
Query: 236 SSKEAWRLEWNLQLITILYS--------------GALSTAAVFCLQSWAMTIKGPTYPSM 281
K W+L+WN+ L+TI+YS L+TAA F + SW +T +GPTYPSM
Sbjct: 236 REKATWKLKWNMSLLTIIYSVTIPVVSDVLNLMQAILNTAAKFVMISWVVTQRGPTYPSM 295
Query: 282 FNPLALVFVAFAEAMILGEPLTVGT 306
F ++++F ++++LG L+VG+
Sbjct: 296 FCAVSVLFTTILDSLLLGHDLSVGS 320
>ref|XP_474441.1| OSJNBa0070M12.19 [Oryza sativa (japonica cultivar-group)]
gi|38345836|emb|CAD41942.2| OSJNBa0070M12.19 [Oryza
sativa (japonica cultivar-group)]
Length = 412
Score = 189 bits (481), Expect = 7e-47
Identities = 107/302 (35%), Positives = 164/302 (53%), Gaps = 13/302 (4%)
Query: 18 SMIFVQLIVTGTQILSRIILV-EGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCE 76
+M+ VQ++ G I ++ +V +G + L +YR L A+ +APLA + ER +
Sbjct: 15 AMVVVQVVFAGVNIFYKLAVVCDGMDMRVLVAYRYLFASAVLAPLAYFVERKNRTKMTWR 74
Query: 77 VLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTW 136
VL F+ G G S+ LY G++ TSAT+A NLIP TF+ A+L R E L I T
Sbjct: 75 VLMLSFVCGLSGGSLAQNLYISGMKLTSATFATAMTNLIPAVTFVLAVLCRYERLAIRTV 134
Query: 137 NGRAKCVGAILCVAGTLAARLYKGKEF--------YIAQYHSFHSVAAHKT----QMLRG 184
G+AK G +L V G + LYKG E +A + H AA T +++ G
Sbjct: 135 AGQAKVAGTLLGVGGAMLLTLYKGAELNPWHTHLDLVAALEARHPAAAAATGNNDRVIMG 194
Query: 185 TLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLE 244
++ ++G+C Y+ W +Q KL +P Y L CVM+ QSA VD WRL
Sbjct: 195 SMLVVGSCVFYAVWLILQAKLSREYPFHYTSTALMCVMSGAQSAAFALLVDREPARWRLG 254
Query: 245 WNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTV 304
+++L++++YSG L++ + + SW + +GP + S+FNPL LV VA +++L E + V
Sbjct: 255 LDIRLLSVVYSGVLASGVMLVVLSWCVKRRGPLFASVFNPLMLVVVAVLGSLLLDEKMHV 314
Query: 305 GT 306
GT
Sbjct: 315 GT 316
>dbj|BAD86994.1| putative MtN21 [Oryza sativa (japonica cultivar-group)]
gi|57899083|dbj|BAD86902.1| putative MtN21 [Oryza sativa
(japonica cultivar-group)]
Length = 378
Score = 183 bits (464), Expect = 7e-45
Identities = 99/299 (33%), Positives = 162/299 (54%), Gaps = 10/299 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ VQL G ++S++ L G + L +YR ++AAV +AP A YFER + +V
Sbjct: 10 AMVMVQLGFAGMNVVSKLALDTGMSPYVLIAYRNIIAAVFLAPFAYYFERKSGMVITKKV 69
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L +IF + G ++ VLY+ G++ T+ T A N +P TF A FRME++ +
Sbjct: 70 LVQIFFSSIFGATLNQVLYFVGLKSTTPTVACALSNTLPALTFAMAAAFRMESVRLSAAA 129
Query: 138 GRAKCVGAILCVAGTLAARLYKGK--EFYIAQYH--------SFHSVAAHKTQMLRGTLF 187
G+AK G ++CV G++ YKG + + H S + A + G +
Sbjct: 130 GQAKVFGTVVCVGGSMIMPFYKGPLLRLWASPIHWRFAESAASGAAAPAAGGAAVLGDVL 189
Query: 188 LIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNL 247
+I +C +++ WF +Q K+ E F Y + C+MA +Q A + A +D S W+L +++
Sbjct: 190 IILSCAAWAVWFIIQTKMSERFSAPYTSTTIMCLMAGVQCAGVSAAMDRSVAVWKLGFDI 249
Query: 248 QLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+L ++LY G + + F L SW + ++GP + SMF+PL LV VA IL E + VG+
Sbjct: 250 RLYSVLYIGVVGSGIAFALMSWCIQVRGPLFVSMFSPLMLVVVAIVGWAILDEKIHVGS 308
>dbj|BAC42941.1| putative nodulin protein N21 [Arabidopsis thaliana]
gi|28973473|gb|AAO64061.1| putative nodulin protein, N21
[Arabidopsis thaliana] gi|15217505|ref|NP_172409.1|
integral membrane family protein / nodulin MtN21-related
[Arabidopsis thaliana]
Length = 374
Score = 180 bits (457), Expect = 4e-44
Identities = 98/299 (32%), Positives = 153/299 (50%), Gaps = 9/299 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ VQ+ G I S++ + G L +YR + A + P+A + ER + +
Sbjct: 11 AMVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLERKTRPKITLRI 70
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L ++F G + VLY+ G++++S T A NL+P TFL A +FR E + I +
Sbjct: 71 LVQVFFCSITGATGNQVLYFVGLQNSSPTIACALTNLLPAVTFLLAAIFRQETVGIKKAS 130
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYI---------AQYHSFHSVAAHKTQMLRGTLFL 188
G+AK +G ++CV G + Y G I A+ + H ++ + G +
Sbjct: 131 GQAKVIGTLVCVIGAMVLSFYHGHTIGIGESKIHWAYAENITKHGSSSGHSNFFLGPFLI 190
Query: 189 IGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQ 248
+ A S++AWF +Q K+ E F Y +L C+M +IQ I D + W L L+
Sbjct: 191 MAAAVSWAAWFIIQTKMSETFAAPYTSTLLMCLMGSIQCGAIALISDHTISDWSLSSPLR 250
Query: 249 LITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGTY 307
I+ LY+G +++A FCL SWAM KGP Y S+F+PL LV VA +L E L GT+
Sbjct: 251 FISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGTF 309
>ref|NP_908553.1| putative MtN21 [Oryza sativa (japonica cultivar-group)]
Length = 380
Score = 179 bits (454), Expect = 1e-43
Identities = 99/301 (32%), Positives = 162/301 (52%), Gaps = 12/301 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNF--SC 75
+M+ VQL G ++S++ L G + L +YR ++AAV +AP A YFER +
Sbjct: 10 AMVMVQLGFAGMNVVSKLALDTGMSPYVLIAYRNIIAAVFLAPFAYYFERYTKSGMVITK 69
Query: 76 EVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHT 135
+VL +IF + G ++ VLY+ G++ T+ T A N +P TF A FRME++ +
Sbjct: 70 KVLVQIFFSSIFGATLNQVLYFVGLKSTTPTVACALSNTLPALTFAMAAAFRMESVRLSA 129
Query: 136 WNGRAKCVGAILCVAGTLAARLYKGK--EFYIAQYH--------SFHSVAAHKTQMLRGT 185
G+AK G ++CV G++ YKG + + H S + A + G
Sbjct: 130 AAGQAKVFGTVVCVGGSMIMPFYKGPLLRLWASPIHWRFAESAASGAAAPAAGGAAVLGD 189
Query: 186 LFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEW 245
+ +I +C +++ WF +Q K+ E F Y + C+MA +Q A + A +D S W+L +
Sbjct: 190 VLIILSCAAWAVWFIIQTKMSERFSAPYTSTTIMCLMAGVQCAGVSAAMDRSVAVWKLGF 249
Query: 246 NLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
+++L ++LY G + + F L SW + ++GP + SMF+PL LV VA IL E + VG
Sbjct: 250 DIRLYSVLYIGVVGSGIAFALMSWCIQVRGPLFVSMFSPLMLVVVAIVGWAILDEKIHVG 309
Query: 306 T 306
+
Sbjct: 310 S 310
>gb|AAM65466.1| putative nodulin protein, N21 [Arabidopsis thaliana]
Length = 363
Score = 178 bits (452), Expect = 2e-43
Identities = 97/298 (32%), Positives = 151/298 (50%), Gaps = 9/298 (3%)
Query: 19 MIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVL 78
M+ VQ+ G I S++ + G L +YR + A + P+A + ER + +L
Sbjct: 1 MVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLERKTRPKITLRIL 60
Query: 79 TKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNG 138
++F G + VLY+ G++++S T A NL+P TFL A +FR E + I +G
Sbjct: 61 VQVFFCSITGATGNQVLYFVGLQNSSPTIACALTNLLPAVTFLLAAIFRQETVGIKKASG 120
Query: 139 RAKCVGAILCVAGTLAARLYKGKEFYI---------AQYHSFHSVAAHKTQMLRGTLFLI 189
+ K +G ++CV G + Y G I A+ + H ++ + G ++
Sbjct: 121 QTKVIGTLVCVIGAMVLSFYHGHTIGIGESKIHWAYAENITKHGSSSGHSNFFLGPFLIM 180
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
A S++AWF +Q K+ E F Y +L C+M +IQ I D + W L L+
Sbjct: 181 AAAVSWAAWFIIQTKMSETFAAPYTSTLLMCLMGSIQCGAIALISDHTISDWSLSSPLRF 240
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGTY 307
I+ LY+G +++A FCL SWAM KGP Y S+F+PL LV VA +L E L GT+
Sbjct: 241 ISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGTF 298
>emb|CAB53493.1| CAA303720.1 protein [Oryza sativa]
Length = 376
Score = 176 bits (446), Expect = 8e-43
Identities = 103/301 (34%), Positives = 157/301 (51%), Gaps = 34/301 (11%)
Query: 19 MIFVQLIVTGTQILSRIILV-EGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
M+ VQ++ G I ++ +V +G + L +YR L A+ +APLA + ERG
Sbjct: 1 MVVVQVVFAGVNIFYKLAVVCDGMDMRVLVAYRYLFASAVLAPLAYFVERG--------- 51
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
S+ LY G++ TSAT+A NLIP TF+ A+L R E L I T
Sbjct: 52 ------------SLAQNLYISGMKLTSATFATAMTNLIPAVTFVLAVLCRYERLAIRTVA 99
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEF--------YIAQYHSFHSVAAHKT----QMLRGT 185
G+AK G +L V G + LYKG E +A + H AA T +++ G+
Sbjct: 100 GQAKVAGTLLGVGGAMLLTLYKGAELNPWHTHLDLVAALEARHPAAAAATGNNDRVIMGS 159
Query: 186 LFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEW 245
+ ++G+C Y+ W +Q KL +P Y L CVM+ QSA VD WRL
Sbjct: 160 MLVVGSCVFYAVWLILQAKLSREYPFHYTSTALMCVMSGAQSAAFALLVDREPARWRLGL 219
Query: 246 NLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVG 305
+++L++++YSG L++ + + SW + +GP + S+FNPL LV VA +++L E + VG
Sbjct: 220 DIRLLSVVYSGVLASGVMLVVLSWCVKRRGPLFASVFNPLMLVVVAVLGSLLLDEKMHVG 279
Query: 306 T 306
T
Sbjct: 280 T 280
>emb|CAE05944.3| OSJNBb0088C09.3 [Oryza sativa (japonica cultivar-group)]
Length = 356
Score = 173 bits (438), Expect = 7e-42
Identities = 98/300 (32%), Positives = 161/300 (53%), Gaps = 12/300 (4%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ +QL+ + Q+ ++ L +G L +YR + AA + P+A ER + + +V
Sbjct: 11 AMVSLQLLFSALQVFIKLALNDGMDARVLVAYRFMFAATFLCPIAFLRERKKRPPLTMKV 70
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
+ ++FL G G S+ LY I+ TSAT+ NL P TFL AIL R+E L +
Sbjct: 71 VLQLFLCGLFGFSINQNLYVLAIKLTSATFITAISNLTPATTFLLAILTRLETLKLKKPA 130
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHKTQ-----------MLRGTL 186
G+AK +G ++ + G + YKG + + H AH T+ + G+
Sbjct: 131 GQAKLLGTLVGMGGAMLLTFYKGPKIMVLDQLP-HPKFAHLTENPQSHPISTGNQIIGSF 189
Query: 187 FLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWN 246
I +CF+Y+ W +Q K+ +V+P Y + C+ A+QS V+ CV E WRL N
Sbjct: 190 LGIISCFTYATWLVIQAKVSKVYPCHYSIAAMVCLFGALQSTVMALCVHRDMEHWRLGLN 249
Query: 247 LQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
++L + Y+G +++ + F L SW + KGP + S+F+PL L+FVA ++IL E L +G+
Sbjct: 250 IRLYSSAYAGLIASGSAFPLLSWCLRKKGPLFISVFSPLMLIFVALMSSIILNEALHLGS 309
>dbj|BAB10303.1| nodulin-like protein [Arabidopsis thaliana]
gi|15238227|ref|NP_201275.1| nodulin MtN21 family
protein [Arabidopsis thaliana]
Length = 359
Score = 169 bits (429), Expect = 8e-41
Identities = 93/297 (31%), Positives = 155/297 (51%), Gaps = 12/297 (4%)
Query: 22 VQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLTKI 81
+Q+I T ++S+ + G F YR A + +APLA +FER S KI
Sbjct: 15 IQVIYTIMFLISKAVFNGGMNTFVFVFYRQAFATIFLAPLAFFFERKSAPPLSFVTFIKI 74
Query: 82 FLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGRAK 141
F+ G+++ + L + TSAT A +P TF A+LF ME L + + G AK
Sbjct: 75 FMLSLFGVTLSLDLNGIALSYTSATLAAATTASLPAITFFLALLFGMERLKVKSIQGTAK 134
Query: 142 CVGAILCVAGTLAARLYKGK--------EFYIAQYHSFHSVAAH----KTQMLRGTLFLI 189
VG +C+ G + +YKG FY Q H + H T L+G + +I
Sbjct: 135 LVGITVCMGGVIILAIYKGPLLKLPLCPHFYHGQEHPHRNNPGHVSGGSTSWLKGCVLMI 194
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
+ + W +Q ++++V+P + + L C++++IQS VI ++ AW+L WNL+L
Sbjct: 195 TSNILWGLWLVLQGRVLKVYPSKLYFTTLHCLLSSIQSFVIAIALERDISAWKLGWNLRL 254
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ ++Y G + T + LQSW + +GP + SMF PL+L+F + A++L E +++G+
Sbjct: 255 VAVIYCGFIVTGVAYYLQSWVIEKRGPVFLSMFTPLSLLFTLLSSAILLCEIISLGS 311
>gb|AAO60108.1| nodulin-like protein [Gossypium hirsutum]
Length = 376
Score = 169 bits (428), Expect = 1e-40
Identities = 94/296 (31%), Positives = 153/296 (50%), Gaps = 9/296 (3%)
Query: 20 IFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLT 79
+ +Q+ G I S++ L G L +YR + A + +AP A + ER + +L
Sbjct: 12 VLLQVGYAGMNITSKLALESGMKPLILVAYRQIFATLAIAPFAFFLERKTRPKLTMPILF 71
Query: 80 KIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGR 139
+IFL G + V Y+ G+++T+AT A N++P TFL A + R E + I +G+
Sbjct: 72 QIFLCSLTGATANQVFYFVGLQNTTATIACALNNVLPAATFLLAAICRQEAVGIKKASGQ 131
Query: 140 AKCVGAILCVAGTLAARLYKGKEFYIAQ---YHSFHSVAAHKTQMLRGTLFLIG------ 190
AK +G ++CV G + Y G I + + ++ + A+ + G+ F +G
Sbjct: 132 AKVIGTLVCVGGAMLLSFYHGHIIGIGESSIHWNYANKMANSSPSPSGSNFFLGPFLVMA 191
Query: 191 ACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLI 250
+ +++ WF +Q + + FP Y L C MA+I+ +IG D AW L +++LI
Sbjct: 192 SAVAWALWFIIQGQTSKSFPAPYTSTTLMCFMASIECTIIGIFSDPKPSAWSLSSSMRLI 251
Query: 251 TILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
LY+G + A FC+ SW + +GP Y S+F+PL LV VA +L E L VGT
Sbjct: 252 AALYAGIICNAVAFCVMSWCIQKRGPLYVSVFSPLLLVIVAILSWALLREKLYVGT 307
>ref|NP_175030.2| integral membrane family protein / nodulin MtN21-related
[Arabidopsis thaliana] gi|44917563|gb|AAS49106.1|
At1g43650 [Arabidopsis thaliana]
gi|51970578|dbj|BAD43981.1| nodulin-like protein
[Arabidopsis thaliana]
Length = 343
Score = 169 bits (427), Expect = 1e-40
Identities = 88/292 (30%), Positives = 157/292 (53%), Gaps = 4/292 (1%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+FVQ++ G +LS++ + +GT F YR AA+ ++P A + E + S +
Sbjct: 10 AMVFVQIVYAGMPLLSKVAISQGTNPFVFVFYRQAFAALALSPFAFFLESSKSSPLSFIL 69
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L KIF G+++ + LYY I +T+AT+A N IP TF+ A+LFR+E + + +
Sbjct: 70 LLKIFFISLCGLTLSLNLYYVAIENTTATFAAATTNAIPSITFVLALLFRLETVTLKKSH 129
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSF---HSVAAHKTQMLRGTLFLIGACFS 194
G AK G+++ + G L KG I Y+S + ++G++ ++ A
Sbjct: 130 GVAKVTGSMVGMLGALVFAFVKGPSL-INHYNSSTIPNGTVPSTKNSVKGSITMLAANTC 188
Query: 195 YSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLITILY 254
+ W MQ K+++ +P + + LQC+ + IQSAV V+ + W++E+ L L+++ Y
Sbjct: 189 WCLWIIMQSKVMKEYPAKLRLVALQCLFSCIQSAVWAVAVNRNPSVWKIEFGLPLLSMAY 248
Query: 255 SGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
G + T + LQ WA+ KGP + +++ PLAL+ + + E +G+
Sbjct: 249 CGIMVTGLTYWLQVWAIEKKGPVFTALYTPLALILTCIVSSFLFKETFYLGS 300
>gb|AAO60157.1| putative nodulin protein [Gossypium hirsutum]
Length = 374
Score = 168 bits (426), Expect = 2e-40
Identities = 94/296 (31%), Positives = 152/296 (50%), Gaps = 9/296 (3%)
Query: 20 IFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLT 79
+ +Q+ G I S++ L G L +YR + A + +AP A + ER + +L
Sbjct: 12 VLLQVGYAGMNITSKLALESGMKPLILVAYRQIFATLAIAPFAFFLERKTRPKLTMPILF 71
Query: 80 KIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGR 139
+IFL G + V Y+ G+++T+AT A N++P TFL A + R E + I +G+
Sbjct: 72 QIFLCSLTGATANQVFYFVGLQNTTATIACALNNVLPAATFLLAAICRQEAVGIKKASGQ 131
Query: 140 AKCVGAILCVAGTLAARLYKGKEFYIAQ---YHSFHSVAAHKTQMLRGTLFLIG------ 190
AK +G + CV G + Y G I + + ++ + A+ + G+ F +G
Sbjct: 132 AKVIGTLACVGGAMLLSFYHGHIIGIGESSIHWNYANKMANSSPSPSGSNFFLGPFLVMA 191
Query: 191 ACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQLI 250
+ +++ WF +Q + + FP Y L C MA+I+ +IG D AW L +++LI
Sbjct: 192 SAVAWALWFIIQGQTSKSFPAPYTSTTLMCFMASIECTIIGIFSDPKPSAWSLSSSMRLI 251
Query: 251 TILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
LY+G + A FC+ SW + +GP Y S+F+PL LV VA +L E L VGT
Sbjct: 252 AALYAGIICNAVAFCVMSWCIQKRGPLYVSVFSPLLLVIVAILSWALLREKLYVGT 307
>gb|AAU44175.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 384
Score = 167 bits (423), Expect = 4e-40
Identities = 91/298 (30%), Positives = 152/298 (50%), Gaps = 11/298 (3%)
Query: 20 IFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEVLT 79
I +Q+I TG ++S+ +G F YR A + + PLAI ER S + T
Sbjct: 11 IIIQVIYTGLYVVSKAAFDQGMNTFVFIFYRQAAATLLLLPLAIILERRNAPAMSLRLFT 70
Query: 80 KIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWNGR 139
K+F+ +G ++ M +Y ++ TSAT A N +P+ TF A+L R+E + + T +G
Sbjct: 71 KLFMYALLGNTITMNMYNVSLKYTSATVASATSNSVPVVTFFLAVLLRLEVIRLRTLSGV 130
Query: 140 AKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAH----------KTQMLRGTLFLI 189
AK G LC+AG L LY G +H S H + + ++GT ++
Sbjct: 131 AKAAGVALCLAGVLVIALYAGPAISPLNHHRALSGGVHGAESSVGTGTRARWMKGTFLML 190
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGAC-VDSSKEAWRLEWNLQ 248
+ ++S W +Q L++ +P + ++QC ++ +QS ++ A V + AWRL +
Sbjct: 191 LSNTTWSLWIVLQASLLKEYPNKLLATLIQCALSTLQSLLLAAAVVRADPAAWRLRLDAG 250
Query: 249 LITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
L+ + Y+G + T F LQ+W + KGP + +M NPL VF F + L E + +G+
Sbjct: 251 LLAVAYTGFVVTGVSFYLQAWCIEKKGPVFLAMSNPLCFVFTIFCSSFFLAEIVHLGS 308
>gb|AAY78620.1| nodulin MtN21 family protein [Arabidopsis thaliana]
Length = 355
Score = 167 bits (422), Expect = 5e-40
Identities = 95/297 (31%), Positives = 157/297 (51%), Gaps = 9/297 (3%)
Query: 18 SMIFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKNFSCEV 77
+M+ VQ I G IL +I + +GT + L +YR+ A + + PLA+ F+R + F+ +
Sbjct: 6 AMVAVQFIFAGMFILFKITVDDGTNLKVLVAYRLSFATIFMLPLALIFQRKKRPEFTWRL 65
Query: 78 LTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMENLNIHTWN 137
L F++G +G ++ +LY G+ TSAT++ + P+ T + ++FRME L + +
Sbjct: 66 LLLAFVSGLLGAAIPNILYLPGMARTSATFSAASSIISPLITLVLGLVFRMETLRLGSNE 125
Query: 138 GRAKCVGAILCVAGTLAARLYKGKEFYI--------AQYHSFHSVAAHKTQMLRGTLFLI 189
GRAK VG +L G L YKG E +I H+ + H +L G L ++
Sbjct: 126 GRAKLVGTLLGACGALVFVFYKGIEIHIWSTHVDLLKGSHTGRATTNHHVSIL-GVLMVL 184
Query: 190 GACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRLEWNLQL 249
G+ S S W +Q K+ + YW L + ++ +I C D E W+L W++ L
Sbjct: 185 GSNVSTSLWLLLQAKIGKELGGLYWNTSLMNGVGSLVCVIIALCSDHDWEQWQLGWDINL 244
Query: 250 ITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLTVGT 306
+ LYSG + + V L +W + KGP + ++F+P+ LV VA + L EPL +G+
Sbjct: 245 LATLYSGIVVSGMVVPLVAWCIATKGPLFVTVFSPIRLVIVALIGSFALEEPLHLGS 301
>gb|AAW78918.2| nodulin-like protein [Triticum aestivum]
Length = 312
Score = 165 bits (418), Expect = 1e-39
Identities = 92/302 (30%), Positives = 152/302 (49%), Gaps = 19/302 (6%)
Query: 20 IFVQLIVTGTQILSRIILVEGTFIFALTSYRVLVAAVCVAPLAIYFERGQPKN------- 72
+ +Q I TG ++S+ G + YR+ A + P+A+ +
Sbjct: 13 VAIQAIYTGLFVVSKAAFDSGINTYVFIFYRLAAATALLLPIALIDSTCRRSRSTTATPA 72
Query: 73 --FSCEVLTKIFLNGFVGMSMVMVLYYYGIRDTSATYALNFLNLIPICTFLTAILFRMEN 130
SC +L K+FL +G + + +Y ++ TSAT N +P+ TFL A+L RME
Sbjct: 73 PALSCRLLFKLFLYALLGNTFTLNMYNVSLKQTSATVGSAATNSMPVATFLLAVLLRMEA 132
Query: 131 LNIHTWNGRAKCVGAILCVAGTLAARLYKGKEFYIAQYHSFHSVAAHK-------TQMLR 183
+ + + +G K G LC+AG L Y G ++ V AHK + ++
Sbjct: 133 VKLRSRSGLGKLAGVALCLAGVLVIAFYAGPSIRPLAHNP---VFAHKPKSVSSGAEWIK 189
Query: 184 GTLFLIGACFSYSAWFFMQVKLVEVFPLRYWGIMLQCVMAAIQSAVIGACVDSSKEAWRL 243
GT LI AC ++S W +QV L++ +P + LQC+ A+QS V+ V+ W+L
Sbjct: 190 GTFLLILACATWSLWIVLQVPLLKEYPNKLMATALQCMFGALQSFVVAVVVERDFTKWKL 249
Query: 244 EWNLQLITILYSGALSTAAVFCLQSWAMTIKGPTYPSMFNPLALVFVAFAEAMILGEPLT 303
++ L+ +LYS L T A+ LQ+W ++GP + +M++PLAL+F F + LGE +
Sbjct: 250 GLDIGLLAVLYSAFLGTGALMYLQAWCAEMRGPVFVAMWSPLALIFTIFCSSFFLGEAVH 309
Query: 304 VG 305
+G
Sbjct: 310 LG 311
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.332 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 510,472,401
Number of Sequences: 2540612
Number of extensions: 19297163
Number of successful extensions: 54090
Number of sequences better than 10.0: 319
Number of HSP's better than 10.0 without gapping: 194
Number of HSP's successfully gapped in prelim test: 125
Number of HSP's that attempted gapping in prelim test: 53553
Number of HSP's gapped (non-prelim): 403
length of query: 328
length of database: 863,360,394
effective HSP length: 128
effective length of query: 200
effective length of database: 538,162,058
effective search space: 107632411600
effective search space used: 107632411600
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146941.4


