

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146863.4 + phase: 0
(309 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
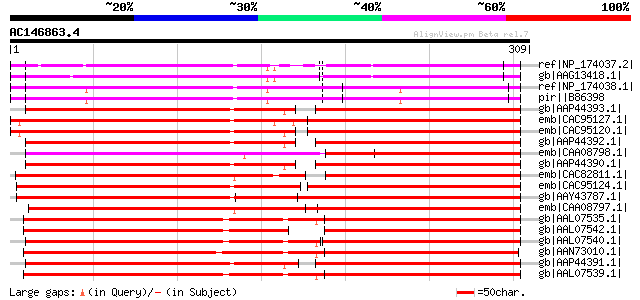
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_174037.2| disease resistance protein (TIR-NBS-LRR class),... 188 2e-46
gb|AAG13418.1| T7N9.23 [Arabidopsis thaliana] 183 6e-45
ref|NP_174038.1| disease resistance protein (TIR-NBS-LRR class),... 172 9e-42
pir||B86398 protein T7N9.24 [imported] - Arabidopsis thaliana gi... 172 9e-42
gb|AAP44393.1| nematode resistance-like protein [Solanum tuberosum] 154 3e-36
emb|CAC95127.1| part I of G08 TIR/NBS/LRR protein [Populus delto... 154 4e-36
emb|CAC95120.1| TIR/NBS/LRR protein [Populus deltoides] 154 4e-36
gb|AAP44392.1| nematode resistance-like protein [Solanum tuberosum] 153 5e-36
emb|CAA08798.1| NL27 [Solanum tuberosum] 153 7e-36
gb|AAP44390.1| nematode resistance protein [Solanum tuberosum] 153 7e-36
emb|CAC82811.1| resistance gene-like [Solanum tuberosum subsp. a... 151 2e-35
emb|CAC95124.1| TIR/NBS/LRR protein [Populus deltoides] 150 3e-35
gb|AAY43787.1| disease resistance protein [(Populus tomentosa x ... 150 6e-35
emb|CAA08797.1| NL25 [Solanum tuberosum] 149 1e-34
gb|AAL07535.1| resistance gene analog PU3 [Helianthus annuus] 148 2e-34
gb|AAL07542.1| resistance gene analog NBS7 [Helianthus annuus] 148 2e-34
gb|AAL07540.1| resistance gene analog NBS5 [Helianthus annuus] 148 2e-34
gb|AAN73010.1| NBS-LRR resistance protein RAS5-1 [Helianthus ann... 147 4e-34
gb|AAP44391.1| nematode resistance-like protein [Solanum tuberosum] 147 5e-34
gb|AAL07539.1| resistance gene analog NBS4 [Helianthus annuus] 146 7e-34
>ref|NP_174037.2| disease resistance protein (TIR-NBS-LRR class), putative
[Arabidopsis thaliana]
Length = 1544
Score = 188 bits (477), Expect = 2e-46
Identities = 109/295 (36%), Positives = 170/295 (56%), Gaps = 24/295 (8%)
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGN-SIASDLIQ 59
++ R + + VF+SF+ D R FT+ L+ L ++++ + +D ++++GN + + L++
Sbjct: 7 VSDQRSRLEWDVFLSFQ-RDARHKFTERLYEVLVKEQVRVWNND-DVERGNHELGASLVE 64
Query: 60 AIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYG 119
A+E S +VV S NYA S WCL ELA + + G+ VLP+FY+V+ +RKQ+G Y
Sbjct: 65 AMEDSVALVVVLSPNYAKSHWCLEELAMLCDLKSSLGRLVLPIFYEVEPCMLRKQNGPYE 124
Query: 120 ESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPS 179
F H KRF + +QR R L ++GNI G+ R +E+E+ + P
Sbjct: 125 MDFEEHSKRFSEEK--IQRWRRALNIIGNIPGFVYR-----SEMESGV-------VSKPH 170
Query: 180 RTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRA 239
R K YDVF+SFRG DTR NF DH AL+ + + FRD+ +++G+ I+ L
Sbjct: 171 RLK------YDVFLSFRGADTRDNFGDHLYKALKDK-VRVFRDNEGMERGDEISSSLKAG 223
Query: 240 IEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQS 294
+E S ++V S+NY+ S WCL EL + + +LP+FY VDPS V+KQS
Sbjct: 224 MEDSAASVIVISRNYSGSRWCLDELAMLCKMKSSLDRRILPIFYHVDPSHVRKQS 278
Score = 127 bits (318), Expect = 5e-28
Identities = 76/187 (40%), Positives = 103/187 (54%), Gaps = 13/187 (6%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG DTR NF DHL+ AL+ K + FRD+ +++G+ I+S L +E S ++
Sbjct: 174 YDVFLSFRGADTRDNFGDHLYKALKDK-VRVFRDNEGMERGDEISSSLKAGMEDSAASVI 232
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
V S+NY+ S WCL ELA + +R+LP+FY VD S VRKQS + F H RF
Sbjct: 233 VISRNYSGSRWCLDELAMLCKMKSSLDRRILPIFYHVDPSHVRKQSDHIKKDFEEHQVRF 292
Query: 130 QDHSNMVQRRRETLQLVGNISGW----DLRD--------KPHHAELENIIEHINILGCKF 177
+ VQ RE L LVGN++G+ D +D K AEL N E +
Sbjct: 293 SEEKEKVQEWREALTLVGNLAGYVCDKDSKDDDMIELVVKRVLAELSNTPEKVGEFIVGL 352
Query: 178 PSRTKDL 184
S KDL
Sbjct: 353 ESPLKDL 359
Score = 77.4 bits (189), Expect = 5e-13
Identities = 40/118 (33%), Positives = 62/118 (51%), Gaps = 1/118 (0%)
Query: 187 INYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVY 246
+ +DVF+SF+ D R FT+ L + + + +D + + L A+E S
Sbjct: 14 LEWDVFLSFQR-DARHKFTERLYEVLVKEQVRVWNNDDVERGNHELGASLVEAMEDSVAL 72
Query: 247 IVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+VV S NYA S WCL EL + G+ VLP+FY+V+P ++KQ+G Y +H
Sbjct: 73 VVVLSPNYAKSHWCLEELAMLCDLKSSLGRLVLPIFYEVEPCMLRKQNGPYEMDFEEH 130
>gb|AAG13418.1| T7N9.23 [Arabidopsis thaliana]
Length = 1560
Score = 183 bits (464), Expect = 6e-45
Identities = 104/295 (35%), Positives = 162/295 (54%), Gaps = 37/295 (12%)
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGN-SIASDLIQ 59
++ R + + VF+SF+ D R FT+ L+ L ++++ + +D ++++GN + + L++
Sbjct: 7 VSDQRSRLEWDVFLSFQ-RDARHKFTERLYEVLVKEQVRVWNND-DVERGNHELGASLVE 64
Query: 60 AIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYG 119
A+E S +VV S NYA S WCL ELA + + G+ VLP+FY+V+ +RKQ+G Y
Sbjct: 65 AMEDSVALVVVLSPNYAKSHWCLEELAMLCDLKSSLGRLVLPIFYEVEPCMLRKQNGPYE 124
Query: 120 ESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPS 179
F H KRF + +QR R L ++GNI G+ R
Sbjct: 125 MDFEEHSKRFSEEK--IQRWRRALNIIGNIPGFVYR------------------------ 158
Query: 180 RTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRA 239
+ YDVF+SFRG DTR NF DH AL+ + + FRD+ +++G+ I+ L
Sbjct: 159 -------LKYDVFLSFRGADTRDNFGDHLYKALKDK-VRVFRDNEGMERGDEISSSLKAG 210
Query: 240 IEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQS 294
+E S ++V S+NY+ S WCL EL + + +LP+FY VDPS V+KQS
Sbjct: 211 MEDSAASVIVISRNYSGSRWCLDELAMLCKMKSSLDRRILPIFYHVDPSHVRKQS 265
Score = 127 bits (318), Expect = 5e-28
Identities = 76/187 (40%), Positives = 103/187 (54%), Gaps = 13/187 (6%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG DTR NF DHL+ AL+ K + FRD+ +++G+ I+S L +E S ++
Sbjct: 161 YDVFLSFRGADTRDNFGDHLYKALKDK-VRVFRDNEGMERGDEISSSLKAGMEDSAASVI 219
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
V S+NY+ S WCL ELA + +R+LP+FY VD S VRKQS + F H RF
Sbjct: 220 VISRNYSGSRWCLDELAMLCKMKSSLDRRILPIFYHVDPSHVRKQSDHIKKDFEEHQVRF 279
Query: 130 QDHSNMVQRRRETLQLVGNISGW----DLRD--------KPHHAELENIIEHINILGCKF 177
+ VQ RE L LVGN++G+ D +D K AEL N E +
Sbjct: 280 SEEKEKVQEWREALTLVGNLAGYVCDKDSKDDDMIELVVKRVLAELSNTPEKVGEFIVGL 339
Query: 178 PSRTKDL 184
S KDL
Sbjct: 340 ESPLKDL 346
Score = 77.4 bits (189), Expect = 5e-13
Identities = 40/118 (33%), Positives = 62/118 (51%), Gaps = 1/118 (0%)
Query: 187 INYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVY 246
+ +DVF+SF+ D R FT+ L + + + +D + + L A+E S
Sbjct: 14 LEWDVFLSFQR-DARHKFTERLYEVLVKEQVRVWNNDDVERGNHELGASLVEAMEDSVAL 72
Query: 247 IVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+VV S NYA S WCL EL + G+ VLP+FY+V+P ++KQ+G Y +H
Sbjct: 73 VVVLSPNYAKSHWCLEELAMLCDLKSSLGRLVLPIFYEVEPCMLRKQNGPYEMDFEEH 130
>ref|NP_174038.1| disease resistance protein (TIR-NBS-LRR class), putative
[Arabidopsis thaliana]
Length = 1556
Score = 172 bits (437), Expect = 9e-42
Identities = 107/301 (35%), Positives = 169/301 (55%), Gaps = 27/301 (8%)
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDD---TNLQKGNSIASDL 57
+++PR + F+SF+ DT NFTD L+ AL ++ + + DD + + + L
Sbjct: 8 VSNPRSRVKWDAFLSFQ-RDTSHNFTDRLYEALVKEELRVWNDDLERVDHDHDHELRPSL 66
Query: 58 IQAIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGG 117
++AIE S F+VV S NYA+S L ELA + + L ++P+FY V+ EV++Q+G
Sbjct: 67 VEAIEDSVAFVVVLSPNYANSHLRLEELAKLCDLKCL----MVPIFYKVEPREVKEQNGP 122
Query: 118 YGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKF 177
+ + F H KRF + +QR + + VGNISG+ + E+E +
Sbjct: 123 FEKDFEEHSKRFGEEK--IQRWKGAMTTVGNISGF-ICGYEIQLEMETGV---------V 170
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAAL-QRRGINAFRDDTKLKKGEFIAPGL 236
P+R K Y VF+SFRG DTR NF + AL +++ + FRD+ ++KG+ I P L
Sbjct: 171 PNRLK------YSVFLSFRGFDTRTNFCERLYIALNEKQNVRVFRDNEGMEKGDKIDPSL 224
Query: 237 FRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGG 296
F AIE S +++ S NYA+S+WCL EL + + ++P+FY V+P +V+KQSG
Sbjct: 225 FEAIEDSAASVIILSTNYANSSWCLDELALLCDLRSSLKRPMIPIFYGVNPEDVRKQSGE 284
Query: 297 Y 297
+
Sbjct: 285 F 285
Score = 117 bits (292), Expect = 6e-25
Identities = 67/195 (34%), Positives = 109/195 (55%), Gaps = 8/195 (4%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKR-IFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
++VF+SFRG DTR NF + L+ AL K+ + FRD+ ++KG+ I L +AIE S +
Sbjct: 176 YSVFLSFRGFDTRTNFCERLYIALNEKQNVRVFRDNEGMEKGDKIDPSLFEAIEDSAASV 235
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
++ S NYA+S+WCL ELA + + + ++P+FY V+ +VRKQSG + + F K
Sbjct: 236 IILSTNYANSSWCLDELALLCDLRSSLKRPMIPIFYGVNPEDVRKQSGEFRKDFEEKAKS 295
Query: 129 FQDHSNMVQRRRETLQLVGNISGW-----DLRDKPHHAELENIIEHINILGCKFPSRTKD 183
F + + +QR + + LVGNI G+ + D E + + I+++ K + ++
Sbjct: 296 FDEET--IQRWKRAMNLVGNIPGYVCTAKTVGDDNEGINREKVDDMIDLVVKKVVAAVRN 353
Query: 184 LVEINYDVFVSFRGP 198
EI D V P
Sbjct: 354 RPEIVADYTVGLESP 368
Score = 65.1 bits (157), Expect = 3e-09
Identities = 42/121 (34%), Positives = 65/121 (53%), Gaps = 8/121 (6%)
Query: 187 INYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEF---IAPGLFRAIEAS 243
+ +D F+SF+ DT NFTD AL + + + DD + + + P L AIE S
Sbjct: 15 VKWDAFLSFQR-DTSHNFTDRLYEALVKEELRVWNDDLERVDHDHDHELRPSLVEAIEDS 73
Query: 244 QVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
++VV S NYA+S L EL + C K ++P+FY V+P EV++Q+G + +
Sbjct: 74 VAFVVVLSPNYANSHLRLEELAKL--CDLKC--LMVPIFYKVEPREVKEQNGPFEKDFEE 129
Query: 304 H 304
H
Sbjct: 130 H 130
>pir||B86398 protein T7N9.24 [imported] - Arabidopsis thaliana
gi|10121909|gb|AAG13419.1| T7N9.24 [Arabidopsis
thaliana]
Length = 1590
Score = 172 bits (437), Expect = 9e-42
Identities = 107/301 (35%), Positives = 169/301 (55%), Gaps = 27/301 (8%)
Query: 1 MTSPRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDD---TNLQKGNSIASDL 57
+++PR + F+SF+ DT NFTD L+ AL ++ + + DD + + + L
Sbjct: 42 VSNPRSRVKWDAFLSFQ-RDTSHNFTDRLYEALVKEELRVWNDDLERVDHDHDHELRPSL 100
Query: 58 IQAIEGSQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGG 117
++AIE S F+VV S NYA+S L ELA + + L ++P+FY V+ EV++Q+G
Sbjct: 101 VEAIEDSVAFVVVLSPNYANSHLRLEELAKLCDLKCL----MVPIFYKVEPREVKEQNGP 156
Query: 118 YGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKF 177
+ + F H KRF + +QR + + VGNISG+ + E+E +
Sbjct: 157 FEKDFEEHSKRFGEEK--IQRWKGAMTTVGNISGF-ICGYEIQLEMETGV---------V 204
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAAL-QRRGINAFRDDTKLKKGEFIAPGL 236
P+R K Y VF+SFRG DTR NF + AL +++ + FRD+ ++KG+ I P L
Sbjct: 205 PNRLK------YSVFLSFRGFDTRTNFCERLYIALNEKQNVRVFRDNEGMEKGDKIDPSL 258
Query: 237 FRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGG 296
F AIE S +++ S NYA+S+WCL EL + + ++P+FY V+P +V+KQSG
Sbjct: 259 FEAIEDSAASVIILSTNYANSSWCLDELALLCDLRSSLKRPMIPIFYGVNPEDVRKQSGE 318
Query: 297 Y 297
+
Sbjct: 319 F 319
Score = 117 bits (292), Expect = 6e-25
Identities = 67/195 (34%), Positives = 109/195 (55%), Gaps = 8/195 (4%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKR-IFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
++VF+SFRG DTR NF + L+ AL K+ + FRD+ ++KG+ I L +AIE S +
Sbjct: 210 YSVFLSFRGFDTRTNFCERLYIALNEKQNVRVFRDNEGMEKGDKIDPSLFEAIEDSAASV 269
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
++ S NYA+S+WCL ELA + + + ++P+FY V+ +VRKQSG + + F K
Sbjct: 270 IILSTNYANSSWCLDELALLCDLRSSLKRPMIPIFYGVNPEDVRKQSGEFRKDFEEKAKS 329
Query: 129 FQDHSNMVQRRRETLQLVGNISGW-----DLRDKPHHAELENIIEHINILGCKFPSRTKD 183
F + + +QR + + LVGNI G+ + D E + + I+++ K + ++
Sbjct: 330 FDEET--IQRWKRAMNLVGNIPGYVCTAKTVGDDNEGINREKVDDMIDLVVKKVVAAVRN 387
Query: 184 LVEINYDVFVSFRGP 198
EI D V P
Sbjct: 388 RPEIVADYTVGLESP 402
Score = 65.1 bits (157), Expect = 3e-09
Identities = 42/121 (34%), Positives = 65/121 (53%), Gaps = 8/121 (6%)
Query: 187 INYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEF---IAPGLFRAIEAS 243
+ +D F+SF+ DT NFTD AL + + + DD + + + P L AIE S
Sbjct: 49 VKWDAFLSFQR-DTSHNFTDRLYEALVKEELRVWNDDLERVDHDHDHELRPSLVEAIEDS 107
Query: 244 QVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
++VV S NYA+S L EL + C K ++P+FY V+P EV++Q+G + +
Sbjct: 108 VAFVVVLSPNYANSHLRLEELAKL--CDLKC--LMVPIFYKVEPREVKEQNGPFEKDFEE 163
Query: 304 H 304
H
Sbjct: 164 H 164
>gb|AAP44393.1| nematode resistance-like protein [Solanum tuberosum]
Length = 1136
Score = 154 bits (389), Expect = 3e-36
Identities = 81/164 (49%), Positives = 111/164 (67%), Gaps = 5/164 (3%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG D R F DHL+ ALQ+K I TF+DD L+KG I+ +L+ +IE S++ ++
Sbjct: 18 YDVFLSFRGEDVRKTFVDHLYLALQQKCINTFKDDEKLEKGKFISPELVSSIEESRIALI 77
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
+FSKNYA+STWCL EL I+ C + G+ V+PVFYDVD S VRKQ +GE+F+ H RF
Sbjct: 78 IFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFSKHEARF 137
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAE---LENIIEHI 170
Q+ + VQ+ R L+ NISGWDL + + E +E I E I
Sbjct: 138 QE--DKVQKWRAALEEAANISGWDLPNTSNGHEARVMEKIAEDI 179
Score = 144 bits (363), Expect = 3e-33
Identities = 66/122 (54%), Positives = 90/122 (73%)
Query: 183 DLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEA 242
+++ +YDVF+SFRG D R F DH ALQ++ IN F+DD KL+KG+FI+P L +IE
Sbjct: 12 EIIRWSYDVFLSFRGEDVRKTFVDHLYLALQQKCINTFKDDEKLEKGKFISPELVSSIEE 71
Query: 243 SQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALS 302
S++ +++FSKNYA+STWCL EL I+ C G+ V+PVFYDVDPS V+KQ +G+A S
Sbjct: 72 SRIALIIFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFS 131
Query: 303 KH 304
KH
Sbjct: 132 KH 133
>emb|CAC95127.1| part I of G08 TIR/NBS/LRR protein [Populus deltoides]
Length = 226
Score = 154 bits (388), Expect = 4e-36
Identities = 88/187 (47%), Positives = 120/187 (64%), Gaps = 12/187 (6%)
Query: 1 MTSP-----RKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIAS 55
MT P R + D+ VF+SFRG DTR FTDHL+ AL + I+TFRDD L +G I+
Sbjct: 1 MTEPESSRSRPEGDYDVFLSFRGEDTRKTFTDHLYTALVQAGIYTFRDDDELPRGEEISY 60
Query: 56 DLIQAIEGSQVFIVVFSKNYASSTWCLRELAYILNC-SVLYGKRVLPVFYDVDLSEVRKQ 114
L++AI+ S++ IVVFSK YASS WCL EL IL C + G+ VLP+FYD+D S VRKQ
Sbjct: 61 HLLRAIQESKISIVVFSKGYASSRWCLNELVEILKCKNGKTGQIVLPIFYDIDPSYVRKQ 120
Query: 115 SGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRD--KPHHAELENII--EHI 170
+G + E+F H +RF++ +V+ R+ L GN+SGW+L D H A+ +I E +
Sbjct: 121 NGSFAEAFVKHEERFEE--TLVKEWRKALAEAGNLSGWNLNDMANGHEAKFIQVIIKEVL 178
Query: 171 NILGCKF 177
N L K+
Sbjct: 179 NKLDPKY 185
Score = 136 bits (343), Expect = 7e-31
Identities = 70/128 (54%), Positives = 88/128 (68%), Gaps = 1/128 (0%)
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLF 237
P ++ E +YDVF+SFRG DTR FTDH AL + GI FRDD +L +GE I+ L
Sbjct: 4 PESSRSRPEGDYDVFLSFRGEDTRKTFTDHLYTALVQAGIYTFRDDDELPRGEEISYHLL 63
Query: 238 RAIEASQVYIVVFSKNYASSTWCLRELEYILHCSK-KHGKHVLPVFYDVDPSEVQKQSGG 296
RAI+ S++ IVVFSK YASS WCL EL IL C K G+ VLP+FYD+DPS V+KQ+G
Sbjct: 64 RAIQESKISIVVFSKGYASSRWCLNELVEILKCKNGKTGQIVLPIFYDIDPSYVRKQNGS 123
Query: 297 YGDALSKH 304
+ +A KH
Sbjct: 124 FAEAFVKH 131
>emb|CAC95120.1| TIR/NBS/LRR protein [Populus deltoides]
Length = 216
Score = 154 bits (388), Expect = 4e-36
Identities = 84/179 (46%), Positives = 117/179 (64%), Gaps = 11/179 (6%)
Query: 1 MTSP-----RKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIAS 55
MT P R + + VF+SFRG DTR FTDHL+ AL + I TFRDD L +G I+
Sbjct: 37 MTEPESSRSRPEGTYDVFLSFRGEDTRHTFTDHLYTALIQAGIHTFRDDDELPRGEEISD 96
Query: 56 DLIQAIEGSQVFIVVFSKNYASSTWCLRELAYILNCS-VLYGKRVLPVFYDVDLSEVRKQ 114
LI+AI+ S++ IVVFSK YASS WCL EL IL C G+ VLP+FYD+D S+VRKQ
Sbjct: 97 HLIRAIQESKISIVVFSKGYASSRWCLNELVEILKCKRKKTGQIVLPIFYDIDPSDVRKQ 156
Query: 115 SGGYGESFNYHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAE---LENIIEHI 170
+G + E+F H +RF++ +V+ R+ L+ GN+SGW+L D + E ++ II+++
Sbjct: 157 NGSFAEAFVKHEERFEE--KLVKEWRKALEEAGNLSGWNLNDMANGHEAKFIQGIIKNV 213
Score = 142 bits (357), Expect = 2e-32
Identities = 71/128 (55%), Positives = 90/128 (69%), Gaps = 1/128 (0%)
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLF 237
P ++ E YDVF+SFRG DTR FTDH AL + GI+ FRDD +L +GE I+ L
Sbjct: 40 PESSRSRPEGTYDVFLSFRGEDTRHTFTDHLYTALIQAGIHTFRDDDELPRGEEISDHLI 99
Query: 238 RAIEASQVYIVVFSKNYASSTWCLRELEYILHCS-KKHGKHVLPVFYDVDPSEVQKQSGG 296
RAI+ S++ IVVFSK YASS WCL EL IL C KK G+ VLP+FYD+DPS+V+KQ+G
Sbjct: 100 RAIQESKISIVVFSKGYASSRWCLNELVEILKCKRKKTGQIVLPIFYDIDPSDVRKQNGS 159
Query: 297 YGDALSKH 304
+ +A KH
Sbjct: 160 FAEAFVKH 167
>gb|AAP44392.1| nematode resistance-like protein [Solanum tuberosum]
Length = 1136
Score = 153 bits (387), Expect = 5e-36
Identities = 81/164 (49%), Positives = 111/164 (67%), Gaps = 5/164 (3%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG D R F DHL+ ALQ+K I TF+DD L+KG I+ +L+ +IE S++ ++
Sbjct: 18 YDVFLSFRGEDVRKTFVDHLYLALQQKCINTFKDDEKLEKGKFISPELMSSIEESRIALI 77
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
+FSKNYA+STWCL EL I+ C + G+ V+PVFYDVD S VRKQ +GE+F+ H RF
Sbjct: 78 IFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFSKHEARF 137
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAE---LENIIEHI 170
Q+ + VQ+ R L+ NISGWDL + + E +E I E I
Sbjct: 138 QE--DKVQKWRAALEEAANISGWDLPNTSNGHEARVMEKIAEDI 179
Score = 144 bits (364), Expect = 2e-33
Identities = 66/122 (54%), Positives = 90/122 (73%)
Query: 183 DLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEA 242
+++ +YDVF+SFRG D R F DH ALQ++ IN F+DD KL+KG+FI+P L +IE
Sbjct: 12 EIIRWSYDVFLSFRGEDVRKTFVDHLYLALQQKCINTFKDDEKLEKGKFISPELMSSIEE 71
Query: 243 SQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALS 302
S++ +++FSKNYA+STWCL EL I+ C G+ V+PVFYDVDPS V+KQ +G+A S
Sbjct: 72 SRIALIIFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFS 131
Query: 303 KH 304
KH
Sbjct: 132 KH 133
>emb|CAA08798.1| NL27 [Solanum tuberosum]
Length = 821
Score = 153 bits (386), Expect = 7e-36
Identities = 83/211 (39%), Positives = 125/211 (58%), Gaps = 14/211 (6%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG DTR FT HL+ L+ + IFTF+DD L+ G+SI +L++AIE SQV ++
Sbjct: 20 YDVFLSFRGVDTRRTFTSHLYEGLKNRGIFTFQDDKRLENGDSIPEELLKAIEESQVALI 79
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
+FSKNYA+S WCL EL I+ C G+ V+P+FYDVD SEVRKQ+ + E+F H ++
Sbjct: 80 IFSKNYATSRWCLNELVKIMECKEEKGQIVIPIFYDVDPSEVRKQTKSFAEAFTEHESKY 139
Query: 130 QDHSNMVQR---RRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRTKDLVE 186
+ +Q+ R L ++ G+D+ ++ +++I++HI++L S K+LV
Sbjct: 140 ANDIEGMQKVKGWRTALSDAADLKGYDISNRIESDYIQHIVDHISVLCKGSLSYIKNLV- 198
Query: 187 INYDVFVSFRGPDTRFNFTDHFCAALQRRGI 217
G DT F A LQ G+
Sbjct: 199 ----------GIDTHFKNIRSLLAELQMSGV 219
Score = 134 bits (336), Expect = 4e-30
Identities = 61/116 (52%), Positives = 84/116 (71%)
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF+SFRG DTR FT H L+ RGI F+DD +L+ G+ I L +AIE SQV ++
Sbjct: 20 YDVFLSFRGVDTRRTFTSHLYEGLKNRGIFTFQDDKRLENGDSIPEELLKAIEESQVALI 79
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+FSKNYA+S WCL EL I+ C ++ G+ V+P+FYDVDPSEV+KQ+ + +A ++H
Sbjct: 80 IFSKNYATSRWCLNELVKIMECKEEKGQIVIPIFYDVDPSEVRKQTKSFAEAFTEH 135
>gb|AAP44390.1| nematode resistance protein [Solanum tuberosum]
Length = 1136
Score = 153 bits (386), Expect = 7e-36
Identities = 80/164 (48%), Positives = 111/164 (66%), Gaps = 5/164 (3%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG D R F DHL+ AL++K I TF+DD L+KG I+ +L+ +IE S++ ++
Sbjct: 18 YDVFLSFRGEDVRKTFVDHLYLALEQKCINTFKDDEKLEKGKFISPELVSSIEESRIALI 77
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
+FSKNYA+STWCL EL I+ C + G+ V+PVFYDVD S VRKQ +GE+F+ H RF
Sbjct: 78 IFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFSKHEARF 137
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAE---LENIIEHI 170
Q+ + VQ+ R L+ NISGWDL + + E +E I E I
Sbjct: 138 QE--DKVQKWRAALEEAANISGWDLPNTANGHEARVMEKIAEDI 179
Score = 144 bits (364), Expect = 2e-33
Identities = 66/122 (54%), Positives = 90/122 (73%)
Query: 183 DLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEA 242
D++ +YDVF+SFRG D R F DH AL+++ IN F+DD KL+KG+FI+P L +IE
Sbjct: 12 DIIRWSYDVFLSFRGEDVRKTFVDHLYLALEQKCINTFKDDEKLEKGKFISPELVSSIEE 71
Query: 243 SQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALS 302
S++ +++FSKNYA+STWCL EL I+ C G+ V+PVFYDVDPS V+KQ +G+A S
Sbjct: 72 SRIALIIFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFS 131
Query: 303 KH 304
KH
Sbjct: 132 KH 133
>emb|CAC82811.1| resistance gene-like [Solanum tuberosum subsp. andigena]
Length = 1126
Score = 151 bits (382), Expect = 2e-35
Identities = 78/176 (44%), Positives = 114/176 (64%), Gaps = 6/176 (3%)
Query: 4 PRKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEG 63
P++K + VF+SFRG DTR NFT HL+ L + IFTF DD L+ G+S++ +L++AI+
Sbjct: 17 PQRKYKYDVFLSFRGKDTRRNFTSHLYERLDNRGIFTFLDDKRLENGDSLSKELVKAIKE 76
Query: 64 SQVFIVVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFN 123
SQV +++FSKNYA+S WCL E+ I+ C G+ V+PVFYDVD S+VRKQ+ + E+F
Sbjct: 77 SQVAVIIFSKNYATSRWCLNEVVKIMECKEENGQLVIPVFYDVDPSDVRKQTKSFAEAFA 136
Query: 124 YHGKRFQDH---SNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCK 176
H R++D VQR R L ++ G+D+R++ E E I E +N + K
Sbjct: 137 EHESRYKDDVEGMQKVQRWRTALSEAADLKGYDIRER---IESECIGELVNEISPK 189
Score = 132 bits (331), Expect = 2e-29
Identities = 59/116 (50%), Positives = 85/116 (72%)
Query: 189 YDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIV 248
YDVF+SFRG DTR NFT H L RGI F DD +L+ G+ ++ L +AI+ SQV ++
Sbjct: 23 YDVFLSFRGKDTRRNFTSHLYERLDNRGIFTFLDDKRLENGDSLSKELVKAIKESQVAVI 82
Query: 249 VFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+FSKNYA+S WCL E+ I+ C +++G+ V+PVFYDVDPS+V+KQ+ + +A ++H
Sbjct: 83 IFSKNYATSRWCLNEVVKIMECKEENGQLVIPVFYDVDPSDVRKQTKSFAEAFAEH 138
>emb|CAC95124.1| TIR/NBS/LRR protein [Populus deltoides]
Length = 1147
Score = 150 bits (380), Expect = 3e-35
Identities = 79/170 (46%), Positives = 111/170 (64%), Gaps = 3/170 (1%)
Query: 5 RKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGS 64
R + + VF+SFRG DTR FTDHL+ AL + I TFRDD L +G I+ ++AI+ S
Sbjct: 34 RPEGAYDVFLSFRGEDTRKTFTDHLYTALVQAGIHTFRDDDELPRGEEISDHFLRAIQES 93
Query: 65 QVFIVVFSKNYASSTWCLRELAYILNCSV-LYGKRVLPVFYDVDLSEVRKQSGGYGESFN 123
++ I VFSK YASS WCL EL IL C G+ VLP+FYD+D S+VRKQ+G + E+F
Sbjct: 94 KISIAVFSKGYASSRWCLNELVEILKCKKRKTGQIVLPIFYDIDPSDVRKQNGSFAEAFV 153
Query: 124 YHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINIL 173
H +RF++ +V+ R+ L+ GN+SGW+L D + E + I E I ++
Sbjct: 154 KHEERFEE--KLVKEWRKALEEAGNLSGWNLNDMANGHEAKFIKEIIKVV 201
Score = 137 bits (346), Expect = 3e-31
Identities = 69/128 (53%), Positives = 88/128 (67%), Gaps = 1/128 (0%)
Query: 178 PSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLF 237
P ++ E YDVF+SFRG DTR FTDH AL + GI+ FRDD +L +GE I+
Sbjct: 28 PESSRSRPEGAYDVFLSFRGEDTRKTFTDHLYTALVQAGIHTFRDDDELPRGEEISDHFL 87
Query: 238 RAIEASQVYIVVFSKNYASSTWCLRELEYILHCSK-KHGKHVLPVFYDVDPSEVQKQSGG 296
RAI+ S++ I VFSK YASS WCL EL IL C K K G+ VLP+FYD+DPS+V+KQ+G
Sbjct: 88 RAIQESKISIAVFSKGYASSRWCLNELVEILKCKKRKTGQIVLPIFYDIDPSDVRKQNGS 147
Query: 297 YGDALSKH 304
+ +A KH
Sbjct: 148 FAEAFVKH 155
>gb|AAY43787.1| disease resistance protein [(Populus tomentosa x P. bolleana) x P.
tomentosa var. truncata]
Length = 428
Score = 150 bits (378), Expect = 6e-35
Identities = 82/168 (48%), Positives = 111/168 (65%), Gaps = 3/168 (1%)
Query: 5 RKKNDHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGS 64
R + + VF+SFRG DTR FTDHL+ AL + I TFRDD L +G I+ L++AIE S
Sbjct: 47 RPEGAYDVFLSFRGEDTRKTFTDHLYTALVQAGIRTFRDDDELPRGEEISHHLLRAIEES 106
Query: 65 QVFIVVFSKNYASSTWCLRELAYILNC-SVLYGKRVLPVFYDVDLSEVRKQSGGYGESFN 123
++ IVVFSK YASS WCL EL IL C + G+ VLP+F+D+D S+VRKQ+ + E+F
Sbjct: 107 RISIVVFSKGYASSRWCLNELVEILKCKNRKTGQIVLPIFFDIDPSDVRKQTASFAEAFV 166
Query: 124 YHGKRFQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHIN 171
H +R Q+ +VQ R+ L+ GN+SGW+L D + E + I E IN
Sbjct: 167 KHEERSQE--KLVQEWRKALKEAGNLSGWNLNDMANGHEAKFIKEIIN 212
Score = 140 bits (352), Expect = 6e-32
Identities = 78/171 (45%), Positives = 106/171 (61%), Gaps = 4/171 (2%)
Query: 135 MVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRTKDLVEINYDVFVS 194
M + +R+ + GN S R K A+L + ++ P ++ E YDVF+S
Sbjct: 1 MQKEKRKQSKDEGNDSSLRKRRK---ADLSKPVSFVSTAAKTEPESSRSRPEGAYDVFLS 57
Query: 195 FRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYIVVFSKNY 254
FRG DTR FTDH AL + GI FRDD +L +GE I+ L RAIE S++ IVVFSK Y
Sbjct: 58 FRGEDTRKTFTDHLYTALVQAGIRTFRDDDELPRGEEISHHLLRAIEESRISIVVFSKGY 117
Query: 255 ASSTWCLRELEYILHC-SKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
ASS WCL EL IL C ++K G+ VLP+F+D+DPS+V+KQ+ + +A KH
Sbjct: 118 ASSRWCLNELVEILKCKNRKTGQIVLPIFFDIDPSDVRKQTASFAEAFVKH 168
>emb|CAA08797.1| NL25 [Solanum tuberosum]
Length = 533
Score = 149 bits (376), Expect = 1e-34
Identities = 76/175 (43%), Positives = 112/175 (63%), Gaps = 3/175 (1%)
Query: 12 VFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIVVF 71
VF+SFRG DTR FT HLF L+ + IFTF+DD L+KG+SI +L++AIE SQV +V+F
Sbjct: 23 VFLSFRGDDTRKTFTSHLFEGLKHRGIFTFQDDKRLEKGDSIPEELLKAIEESQVALVIF 82
Query: 72 SKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRFQD 131
SKNYA+S WCL EL I+ C + + V+PVFYDVD S+VR Q+G + E+F+ H R++D
Sbjct: 83 SKNYATSRWCLNELVKIMECKEVKKQIVMPVFYDVDPSDVRHQTGSFAEAFSKHKSRYKD 142
Query: 132 H---SNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRTKD 183
MVQ R L ++SG ++ + + +++ ++ CK S + +
Sbjct: 143 DVDGMQMVQGWRTALSAAADLSGTNVPGRIESECIRELVDAVSSKLCKTSSSSSE 197
Score = 133 bits (335), Expect = 6e-30
Identities = 64/128 (50%), Positives = 87/128 (67%)
Query: 177 FPSRTKDLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGL 236
+ S +++ Y VF+SFRG DTR FT H L+ RGI F+DD +L+KG+ I L
Sbjct: 9 YASDSQNCTHWKYHVFLSFRGDDTRKTFTSHLFEGLKHRGIFTFQDDKRLEKGDSIPEEL 68
Query: 237 FRAIEASQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGG 296
+AIE SQV +V+FSKNYA+S WCL EL I+ C + + V+PVFYDVDPS+V+ Q+G
Sbjct: 69 LKAIEESQVALVIFSKNYATSRWCLNELVKIMECKEVKKQIVMPVFYDVDPSDVRHQTGS 128
Query: 297 YGDALSKH 304
+ +A SKH
Sbjct: 129 FAEAFSKH 136
>gb|AAL07535.1| resistance gene analog PU3 [Helianthus annuus]
Length = 770
Score = 148 bits (374), Expect = 2e-34
Identities = 77/183 (42%), Positives = 118/183 (64%), Gaps = 9/183 (4%)
Query: 9 DHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
+H VF+SFRG DTR +F DHL+ AL ++ I T++DD L +G I L++AI+ S++ +
Sbjct: 82 NHDVFLSFRGEDTRNSFVDHLYAALVQQGIQTYKDDQTLPRGERIGPALLKAIQESRIAV 141
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
VVFS+NYA S+WCL ELA+I+ C G+ V+P+FY VD S+VRKQ G YG++F H +
Sbjct: 142 VVFSQNYADSSWCLDELAHIMECMDTRGQIVIPIFYFVDPSDVRKQKGKYGKAFRKHKR- 200
Query: 129 FQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRT----KDL 184
++ V+ R+ L+ GN+SGW + + H A+ I E + + + P+ + KDL
Sbjct: 201 --ENKQKVESWRKALEKAGNLSGWVINENSHEAKC--IKEIVATISSRLPTLSTNVNKDL 256
Query: 185 VEI 187
+ I
Sbjct: 257 IGI 259
Score = 138 bits (347), Expect = 2e-31
Identities = 63/117 (53%), Positives = 85/117 (71%)
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
N+DVF+SFRG DTR +F DH AAL ++GI ++DD L +GE I P L +AI+ S++ +
Sbjct: 82 NHDVFLSFRGEDTRNSFVDHLYAALVQQGIQTYKDDQTLPRGERIGPALLKAIQESRIAV 141
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VVFS+NYA S+WCL EL +I+ C G+ V+P+FY VDPS+V+KQ G YG A KH
Sbjct: 142 VVFSQNYADSSWCLDELAHIMECMDTRGQIVIPIFYFVDPSDVRKQKGKYGKAFRKH 198
>gb|AAL07542.1| resistance gene analog NBS7 [Helianthus annuus]
Length = 259
Score = 148 bits (373), Expect = 2e-34
Identities = 72/158 (45%), Positives = 108/158 (67%), Gaps = 3/158 (1%)
Query: 9 DHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
++ VF+SFRG DTR +F DHL+ AL+++ I+T++DD L +G SI L++AI+ S+V +
Sbjct: 21 NYDVFLSFRGDDTRKSFVDHLYTALEQRGIYTYKDDETLPRGESIGPALLKAIQESRVAV 80
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
+VFSKNYA S+WCL ELA+I+ C G+ V+P+FY VD S+VRKQ G YGE+F H +
Sbjct: 81 IVFSKNYADSSWCLDELAHIMECMDTRGQIVMPIFYHVDPSDVRKQKGKYGEAFTKHER- 139
Query: 129 FQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENI 166
++ V+ R+ L+ G +SGW + D + +L I
Sbjct: 140 --ENKLKVESWRKALEKAGKLSGWVINDIENSKDLIGI 175
Score = 145 bits (366), Expect = 1e-33
Identities = 65/117 (55%), Positives = 87/117 (73%)
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
NYDVF+SFRG DTR +F DH AL++RGI ++DD L +GE I P L +AI+ S+V +
Sbjct: 21 NYDVFLSFRGDDTRKSFVDHLYTALEQRGIYTYKDDETLPRGESIGPALLKAIQESRVAV 80
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
+VFSKNYA S+WCL EL +I+ C G+ V+P+FY VDPS+V+KQ G YG+A +KH
Sbjct: 81 IVFSKNYADSSWCLDELAHIMECMDTRGQIVMPIFYHVDPSDVRKQKGKYGEAFTKH 137
>gb|AAL07540.1| resistance gene analog NBS5 [Helianthus annuus]
Length = 285
Score = 148 bits (373), Expect = 2e-34
Identities = 76/180 (42%), Positives = 116/180 (64%), Gaps = 9/180 (5%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
H VF+SFRG DTR NF DHL+ L ++ I T++DD L++G SI L++AI+ S++ ++
Sbjct: 47 HEVFLSFRGEDTRKNFVDHLYKDLVQQGIQTYKDDETLRRGESIRPALLKAIQESRIAVI 106
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
VFS+NYA S+WCL EL I+ C G+ V+P+FY VD S+VRKQ+G YG++F H K
Sbjct: 107 VFSENYADSSWCLDELQQIIECMDTNGQIVIPIFYHVDPSDVRKQNGKYGKAFRKHKK-- 164
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRT----KDLV 185
++ V+ R+ L+ GN+SGW + + H A+ I E + + + P+ + KDL+
Sbjct: 165 -ENKQKVESWRKALEKAGNLSGWVINENSHEAKC--IKEIVGTISSRLPTLSTNVNKDLI 221
Score = 135 bits (339), Expect = 2e-30
Identities = 59/118 (50%), Positives = 86/118 (72%)
Query: 187 INYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVY 246
+ ++VF+SFRG DTR NF DH L ++GI ++DD L++GE I P L +AI+ S++
Sbjct: 45 LKHEVFLSFRGEDTRKNFVDHLYKDLVQQGIQTYKDDETLRRGESIRPALLKAIQESRIA 104
Query: 247 IVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
++VFS+NYA S+WCL EL+ I+ C +G+ V+P+FY VDPS+V+KQ+G YG A KH
Sbjct: 105 VIVFSENYADSSWCLDELQQIIECMDTNGQIVIPIFYHVDPSDVRKQNGKYGKAFRKH 162
>gb|AAN73010.1| NBS-LRR resistance protein RAS5-1 [Helianthus annuus]
Length = 448
Score = 147 bits (371), Expect = 4e-34
Identities = 77/183 (42%), Positives = 118/183 (64%), Gaps = 9/183 (4%)
Query: 9 DHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
+H VF+SFRG DTR +F DHL+ AL ++ I ++DD L +G I L++AI+ S++ +
Sbjct: 83 NHDVFLSFRGEDTRNSFVDHLYAALAQQGIQAYKDDETLPRGERIGPALLKAIQESRIAV 142
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
VVFS+NYA S+WCL ELA+I+ C G+ V+P+FY VD S+VRKQ G YG++F KR
Sbjct: 143 VVFSQNYADSSWCLDELAHIMECMDTRGQIVIPIFYFVDPSDVRKQKGKYGKAFR---KR 199
Query: 129 FQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRT----KDL 184
+++ V+ R+ L+ GN+SGW + + H A+ I E + + + P+ + KDL
Sbjct: 200 KRENRQKVESWRKALEKAGNLSGWVINENSHEAKC--IKEIVATISSRLPTLSTNVNKDL 257
Query: 185 VEI 187
+ I
Sbjct: 258 IGI 260
Score = 137 bits (344), Expect = 5e-31
Identities = 63/116 (54%), Positives = 85/116 (72%)
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
N+DVF+SFRG DTR +F DH AAL ++GI A++DD L +GE I P L +AI+ S++ +
Sbjct: 83 NHDVFLSFRGEDTRNSFVDHLYAALAQQGIQAYKDDETLPRGERIGPALLKAIQESRIAV 142
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSK 303
VVFS+NYA S+WCL EL +I+ C G+ V+P+FY VDPS+V+KQ G YG A K
Sbjct: 143 VVFSQNYADSSWCLDELAHIMECMDTRGQIVIPIFYFVDPSDVRKQKGKYGKAFRK 198
>gb|AAP44391.1| nematode resistance-like protein [Solanum tuberosum]
Length = 1121
Score = 147 bits (370), Expect = 5e-34
Identities = 76/166 (45%), Positives = 112/166 (66%), Gaps = 5/166 (3%)
Query: 10 HTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFIV 69
+ VF+SFRG + R F DHL+ AL++K I TF+DD L+KG I+ +L+ +IE S++ ++
Sbjct: 18 YDVFLSFRGENVRKTFVDHLYLALEQKCINTFKDDEKLEKGKFISPELMSSIEESRIALI 77
Query: 70 VFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKRF 129
+FSKNYA+STWCL EL I+ C + G+ V+PVFYDVD S VR+Q +GE+F+ H RF
Sbjct: 78 IFSKNYANSTWCLDELTKIIECKNVKGQIVVPVFYDVDPSTVRRQKNIFGEAFSKHEARF 137
Query: 130 QDHSNMVQRRRETLQLVGNISGWDLRDKPHHAE---LENIIEHINI 172
++ + V++ R L+ NISGWDL + + E +E I E I +
Sbjct: 138 EE--DKVKKWRAALEEAANISGWDLPNTSNGHEARVIEKITEDIMV 181
Score = 140 bits (353), Expect = 5e-32
Identities = 63/122 (51%), Positives = 90/122 (73%)
Query: 183 DLVEINYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEA 242
+++ +YDVF+SFRG + R F DH AL+++ IN F+DD KL+KG+FI+P L +IE
Sbjct: 12 EIIRWSYDVFLSFRGENVRKTFVDHLYLALEQKCINTFKDDEKLEKGKFISPELMSSIEE 71
Query: 243 SQVYIVVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALS 302
S++ +++FSKNYA+STWCL EL I+ C G+ V+PVFYDVDPS V++Q +G+A S
Sbjct: 72 SRIALIIFSKNYANSTWCLDELTKIIECKNVKGQIVVPVFYDVDPSTVRRQKNIFGEAFS 131
Query: 303 KH 304
KH
Sbjct: 132 KH 133
>gb|AAL07539.1| resistance gene analog NBS4 [Helianthus annuus]
Length = 279
Score = 146 bits (369), Expect = 7e-34
Identities = 77/183 (42%), Positives = 117/183 (63%), Gaps = 9/183 (4%)
Query: 9 DHTVFVSFRGTDTRFNFTDHLFGALQRKRIFTFRDDTNLQKGNSIASDLIQAIEGSQVFI 68
+H VF+SFRG DTR +F DHL+ AL ++ I T++DD L +G I L++AI+ S++ +
Sbjct: 21 NHDVFLSFRGEDTRNSFVDHLYVALAQQGILTYKDDETLPRGERIGPTLLKAIQESRIAL 80
Query: 69 VVFSKNYASSTWCLRELAYILNCSVLYGKRVLPVFYDVDLSEVRKQSGGYGESFNYHGKR 128
VVFS+NYA S+WCL ELA+I+ C G+ V P+FY VD S+VRKQ G YG++F H +
Sbjct: 81 VVFSENYADSSWCLDELAHIMECVDTRGQIVEPIFYFVDPSDVRKQKGKYGKAFRKHKR- 139
Query: 129 FQDHSNMVQRRRETLQLVGNISGWDLRDKPHHAELENIIEHINILGCKFPSRT----KDL 184
++ + V+ R+ L+ GN+SGW + + H A I E + + + P+ + KDL
Sbjct: 140 --ENKHKVESWRKALEKAGNLSGWVINENSHEARC--IKEIVATISSRLPTLSTNVNKDL 195
Query: 185 VEI 187
+ I
Sbjct: 196 IGI 198
Score = 132 bits (332), Expect = 1e-29
Identities = 62/117 (52%), Positives = 83/117 (69%)
Query: 188 NYDVFVSFRGPDTRFNFTDHFCAALQRRGINAFRDDTKLKKGEFIAPGLFRAIEASQVYI 247
N+DVF+SFRG DTR +F DH AL ++GI ++DD L +GE I P L +AI+ S++ +
Sbjct: 21 NHDVFLSFRGEDTRNSFVDHLYVALAQQGILTYKDDETLPRGERIGPTLLKAIQESRIAL 80
Query: 248 VVFSKNYASSTWCLRELEYILHCSKKHGKHVLPVFYDVDPSEVQKQSGGYGDALSKH 304
VVFS+NYA S+WCL EL +I+ C G+ V P+FY VDPS+V+KQ G YG A KH
Sbjct: 81 VVFSENYADSSWCLDELAHIMECVDTRGQIVEPIFYFVDPSDVRKQKGKYGKAFRKH 137
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.139 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 535,948,910
Number of Sequences: 2540612
Number of extensions: 22189428
Number of successful extensions: 42042
Number of sequences better than 10.0: 477
Number of HSP's better than 10.0 without gapping: 400
Number of HSP's successfully gapped in prelim test: 77
Number of HSP's that attempted gapping in prelim test: 40367
Number of HSP's gapped (non-prelim): 985
length of query: 309
length of database: 863,360,394
effective HSP length: 127
effective length of query: 182
effective length of database: 540,702,670
effective search space: 98407885940
effective search space used: 98407885940
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146863.4


