

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146862.7 - phase: 0
(198 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
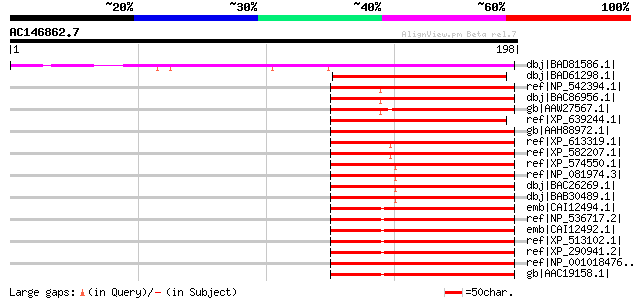
Sequences producing significant alignments: (bits) Value
dbj|BAD81586.1| putative 3' exoribonuclease [Oryza sativa (japon... 153 2e-36
dbj|BAD61298.1| unknown protein [Oryza sativa (japonica cultivar... 88 1e-16
ref|NP_542394.1| hypothetical protein LOC112479 [Homo sapiens] g... 79 1e-13
dbj|BAC86956.1| unnamed protein product [Homo sapiens] 79 1e-13
gb|AAW27567.1| unknown [Schistosoma japonicum] 77 4e-13
ref|XP_639244.1| hypothetical protein DDB0185367 [Dictyostelium ... 77 4e-13
gb|AAH88972.1| LOC503681 protein [Xenopus laevis] 76 5e-13
ref|XP_613319.1| PREDICTED: similar to 4933424N09Rik protein [Bo... 73 5e-12
ref|XP_582207.1| PREDICTED: similar to 4933424N09Rik protein [Bo... 73 5e-12
ref|XP_574550.1| PREDICTED: similar to RIKEN cDNA 4933424N09 [Ra... 72 7e-12
ref|NP_081974.3| hypothetical protein LOC71151 [Mus musculus] gi... 72 9e-12
dbj|BAC26269.1| unnamed protein product [Mus musculus] 72 9e-12
dbj|BAB30489.1| unnamed protein product [Mus musculus] 72 9e-12
emb|CAI12494.1| prion protein interacting protein [Homo sapiens]... 72 1e-11
ref|NP_536717.2| prion protein interacting protein 1 [Mus muscul... 72 1e-11
emb|CAI12492.1| prion protein interacting protein [Homo sapiens] 72 1e-11
ref|XP_513102.1| PREDICTED: similar to prion protein interacting... 72 1e-11
ref|XP_290941.2| PREDICTED: prion protein interacting protein [H... 72 1e-11
ref|NP_001018476.1| hypothetical protein LOC553667 [Danio rerio]... 72 1e-11
gb|AAC19158.1| unknown [Homo sapiens] 72 1e-11
>dbj|BAD81586.1| putative 3' exoribonuclease [Oryza sativa (japonica
cultivar-group)] gi|56783678|dbj|BAD81090.1| putative 3'
exoribonuclease [Oryza sativa (japonica cultivar-group)]
Length = 411
Score = 153 bits (387), Expect = 2e-36
Identities = 96/208 (46%), Positives = 117/208 (56%), Gaps = 25/208 (12%)
Query: 1 MQITCEASLKCLQGKGPPFTFQCNGSSMEVFPELNNEPGNHPSGNVPEPNHRLGSE-FLE 59
MQ+ EA L CL+ P + FPE N + N + +E LE
Sbjct: 1 MQVNKEAPLGCLK---PISQYNLQEQRSNGFPE-----------NSEKKNDSIATERVLE 46
Query: 60 PS---NEFHNKPTYHHDYSTWTACHFHPHKMQQCQMNAFENHFYP------HPVENQFQY 110
S N+ +P +Y + + Q Q +N FY P+E++ QY
Sbjct: 47 ASPLPNQGFFRPVQRTEYYAYPFIYADYQMPGQPQPYNLDNQFYQINRDHSFPIESRVQY 106
Query: 111 APINMVSQGYPRE-QYQEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEAC 169
P M QGYP + Q QEFQ FVVIDFEATCDK+ NPHPQEIIEFPSV+V+S TGQLEA
Sbjct: 107 LPFKMPPQGYPPDAQLQEFQYFVVIDFEATCDKENNPHPQEIIEFPSVLVNSATGQLEAS 166
Query: 170 FQTYVRPTCNQHLSDFCKDLTGIQQIQV 197
FQTYVRP NQ L+DFCK+LTGIQQIQV
Sbjct: 167 FQTYVRPAYNQLLTDFCKELTGIQQIQV 194
>dbj|BAD61298.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 304
Score = 88.2 bits (217), Expect = 1e-16
Identities = 38/68 (55%), Positives = 54/68 (78%)
Query: 127 EFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDFC 186
EF +FVV+DFEATC++ + +PQEIIEFP+V+V + TG+L + F+ YVRP + L+DFC
Sbjct: 26 EFDHFVVVDFEATCERGRRIYPQEIIEFPAVLVDAATGRLVSAFRAYVRPRHHPRLTDFC 85
Query: 187 KDLTGIQQ 194
++LTGI Q
Sbjct: 86 RELTGIAQ 93
>ref|NP_542394.1| hypothetical protein LOC112479 [Homo sapiens]
gi|14714721|gb|AAH10503.1| Hypothetical protein MGC16943
[Homo sapiens]
Length = 328
Score = 78.6 bits (192), Expect = 1e-13
Identities = 39/73 (53%), Positives = 55/73 (74%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F +VIDFE+TC D K+ H QEIIEFP+V++++ TGQ+++ FQ YV+P + LS+
Sbjct: 32 QLFDYLIVIDFESTCWNDGKHHHSQEIIEFPAVLLNTSTGQIDSEFQAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCMELTGIKQAQV 104
>dbj|BAC86956.1| unnamed protein product [Homo sapiens]
Length = 279
Score = 78.6 bits (192), Expect = 1e-13
Identities = 39/73 (53%), Positives = 55/73 (74%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F +VIDFE+TC D K+ H QEIIEFP+V++++ TGQ+++ FQ YV+P + LS+
Sbjct: 32 QLFDYLIVIDFESTCWNDGKHHHSQEIIEFPAVLLNTSTGQIDSEFQAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCMELTGIKQAQV 104
>gb|AAW27567.1| unknown [Schistosoma japonicum]
Length = 201
Score = 76.6 bits (187), Expect = 4e-13
Identities = 41/73 (56%), Positives = 52/73 (71%), Gaps = 2/73 (2%)
Query: 126 QEFQNFVVIDFEATCDKD-KNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q+F F+V+DFEATC+KD K P P EIIEFP ++V++ T E+ F YVRPT N LSD
Sbjct: 5 QKFAYFMVLDFEATCEKDIKIPDP-EIIEFPVLMVNAYTLHTESIFHHYVRPTINPVLSD 63
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI Q +
Sbjct: 64 FCTELTGIIQSMI 76
>ref|XP_639244.1| hypothetical protein DDB0185367 [Dictyostelium discoideum]
gi|60467872|gb|EAL65886.1| hypothetical protein
DDB0185367 [Dictyostelium discoideum]
Length = 220
Score = 76.6 bits (187), Expect = 4e-13
Identities = 36/69 (52%), Positives = 51/69 (73%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q+F+ +V+DFEATC+KD QEIIEFPSVI+++ T + + F+ YV+P N +LS F
Sbjct: 19 QKFKYLIVLDFEATCEKDVKFPNQEIIEFPSVIINTETLETVSTFREYVKPIINPNLSKF 78
Query: 186 CKDLTGIQQ 194
C +LTGI+Q
Sbjct: 79 CTELTGIKQ 87
>gb|AAH88972.1| LOC503681 protein [Xenopus laevis]
Length = 687
Score = 76.3 bits (186), Expect = 5e-13
Identities = 37/72 (51%), Positives = 51/72 (70%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q F+ ++IDFE+TC KD QEIIEFP+V+++ G++E+ F TYV+P + LSDF
Sbjct: 33 QFFEYLIIIDFESTCWKDGKHSTQEIIEFPAVLLNVSNGEIESEFHTYVQPQEHPILSDF 92
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q QV
Sbjct: 93 CTELTGINQQQV 104
>ref|XP_613319.1| PREDICTED: similar to 4933424N09Rik protein [Bos taurus]
Length = 825
Score = 72.8 bits (177), Expect = 5e-12
Identities = 36/73 (49%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPH-PQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F ++IDFE+TC D H QEIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 168 QLFDYLIIIDFESTCWNDGKRHRSQEIIEFPAVLLNTSTGEIESEFHAYVQPQEHPILSE 227
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 228 FCVELTGIKQAQV 240
>ref|XP_582207.1| PREDICTED: similar to 4933424N09Rik protein [Bos taurus]
Length = 689
Score = 72.8 bits (177), Expect = 5e-12
Identities = 36/73 (49%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPH-PQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q F ++IDFE+TC D H QEIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLFDYLIIIDFESTCWNDGKRHRSQEIIEFPAVLLNTSTGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q QV
Sbjct: 92 FCVELTGIKQAQV 104
>ref|XP_574550.1| PREDICTED: similar to RIKEN cDNA 4933424N09 [Rattus norvegicus]
Length = 257
Score = 72.4 bits (176), Expect = 7e-12
Identities = 34/73 (46%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLYDYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q+QV
Sbjct: 92 FCTELTGIKQVQV 104
>ref|NP_081974.3| hypothetical protein LOC71151 [Mus musculus]
gi|60688299|gb|AAH90961.1| RIKEN cDNA 4933424N09 [Mus
musculus]
Length = 688
Score = 72.0 bits (175), Expect = 9e-12
Identities = 34/73 (46%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLYAYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q+QV
Sbjct: 92 FCTELTGIKQVQV 104
>dbj|BAC26269.1| unnamed protein product [Mus musculus]
Length = 688
Score = 72.0 bits (175), Expect = 9e-12
Identities = 34/73 (46%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLYAYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q+QV
Sbjct: 92 FCTELTGIKQVQV 104
>dbj|BAB30489.1| unnamed protein product [Mus musculus]
Length = 274
Score = 72.0 bits (175), Expect = 9e-12
Identities = 34/73 (46%), Positives = 52/73 (70%), Gaps = 1/73 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQ-EIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSD 184
Q + +V+DFE+TC D H EIIEFP+V++++ TG++E+ F YV+P + LS+
Sbjct: 32 QLYAYLIVVDFESTCWNDGKHHSSPEIIEFPAVLLNTATGEIESEFHAYVQPQEHPILSE 91
Query: 185 FCKDLTGIQQIQV 197
FC +LTGI+Q+QV
Sbjct: 92 FCTELTGIKQVQV 104
>emb|CAI12494.1| prion protein interacting protein [Homo sapiens]
gi|56203068|emb|CAI23196.1| prion protein interacting
protein [Homo sapiens]
Length = 337
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 141 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 199
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 200 CTELTGIIQAMV 211
>ref|NP_536717.2| prion protein interacting protein 1 [Mus musculus]
gi|26350187|dbj|BAC38733.1| unnamed protein product [Mus
musculus]
Length = 337
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 141 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 199
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 200 CTELTGIIQAMV 211
>emb|CAI12492.1| prion protein interacting protein [Homo sapiens]
Length = 250
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 139 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 197
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 198 CTELTGIIQAMV 209
>ref|XP_513102.1| PREDICTED: similar to prion protein interacting protein 1 [Pan
troglodytes]
Length = 424
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 141 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 199
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 200 CTELTGIIQAMV 211
>ref|XP_290941.2| PREDICTED: prion protein interacting protein [Homo sapiens]
Length = 462
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 266 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 324
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 325 CTELTGIIQAMV 336
>ref|NP_001018476.1| hypothetical protein LOC553667 [Danio rerio]
gi|63100865|gb|AAH95637.1| Hypothetical protein
LOC553667 [Danio rerio]
Length = 555
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 50/72 (68%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q F ++IDFE+TC ++K+ QEIIEFP+V++S +G +E+ F +YV+P LS F
Sbjct: 30 QRFSFLIIIDFESTCWREKSSSGQEIIEFPAVLLSVCSGAVESEFHSYVQPQERPVLSAF 89
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q QV
Sbjct: 90 CTELTGITQDQV 101
>gb|AAC19158.1| unknown [Homo sapiens]
Length = 397
Score = 71.6 bits (174), Expect = 1e-11
Identities = 35/72 (48%), Positives = 49/72 (67%), Gaps = 1/72 (1%)
Query: 126 QEFQNFVVIDFEATCDKDKNPHPQEIIEFPSVIVSSVTGQLEACFQTYVRPTCNQHLSDF 185
Q + F+V+DFEATCDK + HPQEIIEFP + ++ T ++E+ F YV+P + L+ F
Sbjct: 201 QRYHYFLVLDFEATCDKPQI-HPQEIIEFPILKLNGRTMEIESTFHMYVQPVVHPQLTPF 259
Query: 186 CKDLTGIQQIQV 197
C +LTGI Q V
Sbjct: 260 CTELTGIIQAMV 271
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.135 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 393,621,274
Number of Sequences: 2540612
Number of extensions: 17650234
Number of successful extensions: 31977
Number of sequences better than 10.0: 135
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 55
Number of HSP's that attempted gapping in prelim test: 31781
Number of HSP's gapped (non-prelim): 170
length of query: 198
length of database: 863,360,394
effective HSP length: 121
effective length of query: 77
effective length of database: 555,946,342
effective search space: 42807868334
effective search space used: 42807868334
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146862.7


