

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146855.3 - phase: 0
(186 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
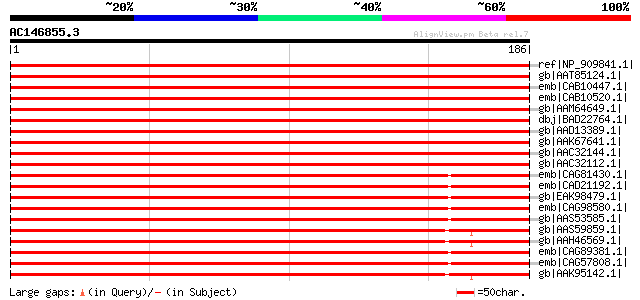
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_909841.1| ribosomal protein L15 [Oryza sativa (japonica c... 348 6e-95
gb|AAT85124.1| putative 60s ribosomal protein L15 [Oryza sativa ... 348 6e-95
emb|CAB10447.1| ribosomal protein [Arabidopsis thaliana] gi|7268... 344 6e-94
emb|CAB10520.1| ribosomal protein [Arabidopsis thaliana] gi|7268... 343 1e-93
gb|AAM64649.1| 60S ribosomal protein L15 homolog [Arabidopsis th... 343 1e-93
dbj|BAD22764.1| ribosomal protein [Bromus inermis] 341 5e-93
gb|AAD13389.1| ribosomal protein L15 [Petunia x hybrida] gi|6094... 341 7e-93
gb|AAK67641.1| ribosomal protein L15 [Homo sapiens] 340 2e-92
gb|AAC32144.1| probable 60S ribosomal protein L15 [Picea mariana... 336 2e-91
gb|AAC32112.1| probable 60S ribosomal protein L15 [Picea mariana... 335 4e-91
emb|CAG81430.1| unnamed protein product [Yarrowia lipolytica CLI... 288 5e-77
emb|CAD21192.1| probable ribosomal protein L15.e.B, cytosolic [N... 287 1e-76
gb|EAK98479.1| likely cytosolic ribosomal protein L15 [Candida a... 285 6e-76
emb|CAG98580.1| unnamed protein product [Kluyveromyces lactis NR... 282 4e-75
gb|AAS53585.1| AFR214Cp [Ashbya gossypii ATCC 10895] gi|45198732... 281 5e-75
gb|AAS59859.1| ribosomal protein L15 [Acipenser gueldenstaedtii]... 281 8e-75
gb|AAH46569.1| Rpl15-prov protein [Xenopus laevis] gi|49670582|g... 279 3e-74
emb|CAG89381.1| unnamed protein product [Debaryomyces hansenii C... 278 4e-74
emb|CAG57808.1| unnamed protein product [Candida glabrata CBS138... 278 7e-74
gb|AAK95142.1| ribosomal protein L15 [Ictalurus punctatus] gi|47... 278 7e-74
>ref|NP_909841.1| ribosomal protein L15 [Oryza sativa (japonica cultivar-group)]
gi|28875969|gb|AAO59978.1| ribosomal protein L15 [Oryza
sativa (japonica cultivar-group)]
gi|22795244|gb|AAN08216.1| ribosomal protein L15 [Oryza
sativa (japonica cultivar-group)]
Length = 204
Score = 348 bits (892), Expect = 6e-95
Identities = 166/186 (89%), Positives = 176/186 (94%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ +IVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPV KG
Sbjct: 19 MRFVQRVRCWEYRQQPAIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP +QG+TQLKFQR+KRSVAEERAGRKLGGLRVLNSYWVNEDST+KYFE+ILVDVA
Sbjct: 79 IVYGKPKHQGITQLKFQRNKRSVAEERAGRKLGGLRVLNSYWVNEDSTYKYFEIILVDVA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
HSAIRNDPRINWL PVHKHRELRGLTSAGK+ RGL GKGH +HKARPSRRA WKRN T+
Sbjct: 139 HSAIRNDPRINWLCKPVHKHRELRGLTSAGKKYRGLRGKGHTHHKARPSRRATWKRNQTV 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>gb|AAT85124.1| putative 60s ribosomal protein L15 [Oryza sativa (japonica
cultivar-group)]
Length = 186
Score = 348 bits (892), Expect = 6e-95
Identities = 166/186 (89%), Positives = 176/186 (94%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ +IVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPV KG
Sbjct: 1 MRFVQRVRCWEYRQQPAIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVPKG 60
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP +QG+TQLKFQR+KRSVAEERAGRKLGGLRVLNSYWVNEDST+KYFE+ILVDVA
Sbjct: 61 IVYGKPKHQGITQLKFQRNKRSVAEERAGRKLGGLRVLNSYWVNEDSTYKYFEIILVDVA 120
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
HSAIRNDPRINWL PVHKHRELRGLTSAGK+ RGL GKGH +HKARPSRRA WKRN T+
Sbjct: 121 HSAIRNDPRINWLCKPVHKHRELRGLTSAGKKYRGLRGKGHTHHKARPSRRATWKRNQTV 180
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 181 SLRRYR 186
>emb|CAB10447.1| ribosomal protein [Arabidopsis thaliana] gi|7268422|emb|CAB78714.1|
ribosomal protein [Arabidopsis thaliana]
gi|23505795|gb|AAN28757.1| At4g16720/dl4385c
[Arabidopsis thaliana] gi|21592436|gb|AAM64387.1|
ribosomal protein [Arabidopsis thaliana]
gi|22136774|gb|AAM91731.1| putative ribosomal protein
[Arabidopsis thaliana] gi|13878179|gb|AAK44167.1|
putative ribosomal protein [Arabidopsis thaliana]
gi|19715591|gb|AAL91619.1| AT4g16720/dl4385c
[Arabidopsis thaliana] gi|16604446|gb|AAL24229.1|
AT4g16720/dl4385c [Arabidopsis thaliana]
gi|15235851|ref|NP_193405.1| 60S ribosomal protein L15
(RPL15A) [Arabidopsis thaliana] gi|7441105|pir||E71434
ribosomal protein L15.DL4385C, cytosolic - Arabidopsis
thaliana gi|3122673|sp|O23515|RL15_ARATH 60S ribosomal
protein L15
Length = 204
Score = 344 bits (883), Expect = 6e-94
Identities = 162/186 (87%), Positives = 175/186 (93%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ SIVRL RPTRPDKARRLGYKAKQG+VVYRVRVRRGGRKRPV KG
Sbjct: 19 MRFLQRVRCWEYRQQPSIVRLVRPTRPDKARRLGYKAKQGFVVYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRV+NSYW+NEDST+KY+E+ILVD A
Sbjct: 79 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVVNSYWLNEDSTYKYYEIILVDPA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+A+RNDPRINW+ NPVHKHRELRGLTS GK+NRGL GKGH HK RPSRRA WK+NN+L
Sbjct: 139 HNAVRNDPRINWICNPVHKHRELRGLTSEGKKNRGLRGKGHNNHKNRPSRRATWKKNNSL 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>emb|CAB10520.1| ribosomal protein [Arabidopsis thaliana] gi|7268491|emb|CAB78742.1|
ribosomal protein [Arabidopsis thaliana]
gi|7441107|pir||C71443 ribosomal protein L15.DL4730C,
cytosolic - Arabidopsis thaliana
Length = 216
Score = 343 bits (881), Expect = 1e-93
Identities = 161/186 (86%), Positives = 175/186 (93%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ SIVRL RPTRPDKARRLGYKAKQG+VVYRVRVRRGGRKRPV KG
Sbjct: 31 MRFLQRVRCWEYRQQPSIVRLVRPTRPDKARRLGYKAKQGFVVYRVRVRRGGRKRPVPKG 90
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRV+NSYW+NEDST+KY+E+ILVD A
Sbjct: 91 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVVNSYWLNEDSTYKYYEIILVDPA 150
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+A+RNDPRINW+ NPVHKHRELRGLTS GK+NRGL GKGH HK RPSRRA WK+NN++
Sbjct: 151 HNAVRNDPRINWICNPVHKHRELRGLTSEGKKNRGLRGKGHNNHKNRPSRRATWKKNNSI 210
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 211 SLRRYR 216
>gb|AAM64649.1| 60S ribosomal protein L15 homolog [Arabidopsis thaliana]
gi|21689875|gb|AAM67498.1| putative 60S ribosomal
protein L15-like protein [Arabidopsis thaliana]
gi|18175858|gb|AAL59940.1| putative 60S ribosomal
protein L15-like protein [Arabidopsis thaliana]
gi|15236049|ref|NP_193470.1| 60S ribosomal protein L15
(RPL15B) [Arabidopsis thaliana]
Length = 204
Score = 343 bits (881), Expect = 1e-93
Identities = 161/186 (86%), Positives = 175/186 (93%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ SIVRL RPTRPDKARRLGYKAKQG+VVYRVRVRRGGRKRPV KG
Sbjct: 19 MRFLQRVRCWEYRQQPSIVRLVRPTRPDKARRLGYKAKQGFVVYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRV+NSYW+NEDST+KY+E+ILVD A
Sbjct: 79 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVVNSYWLNEDSTYKYYEIILVDPA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+A+RNDPRINW+ NPVHKHRELRGLTS GK+NRGL GKGH HK RPSRRA WK+NN++
Sbjct: 139 HNAVRNDPRINWICNPVHKHRELRGLTSEGKKNRGLRGKGHNNHKNRPSRRATWKKNNSI 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>dbj|BAD22764.1| ribosomal protein [Bromus inermis]
Length = 204
Score = 341 bits (875), Expect = 5e-93
Identities = 160/186 (86%), Positives = 176/186 (94%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ +IVR+TRPTRPD+ARRLG+KAKQGYVVYR+RVRRGGRKRPV KG
Sbjct: 19 MRFVQRVRCWEYRQQPAIVRITRPTRPDRARRLGFKAKQGYVVYRIRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP +QG+TQLKFQR+KRSVAEERAGR+LGGLRVLNSYWVNEDST+KYFEVILVDVA
Sbjct: 79 IVYGKPKHQGITQLKFQRNKRSVAEERAGRRLGGLRVLNSYWVNEDSTYKYFEVILVDVA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+A+RNDPRINWL PVHKHRELRGLTSAGK+ RGL GKGHR+HK RPSRRA WKRN T+
Sbjct: 139 HNAVRNDPRINWLCKPVHKHRELRGLTSAGKKFRGLRGKGHRHHKNRPSRRATWKRNQTV 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>gb|AAD13389.1| ribosomal protein L15 [Petunia x hybrida]
gi|6094014|sp|O82528|RL15_PETHY 60S ribosomal protein
L15
Length = 204
Score = 341 bits (874), Expect = 7e-93
Identities = 163/186 (87%), Positives = 175/186 (93%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQ SIVR+TRPTRPDKARRLGYKAKQGYVVYRVRV+RGGRKRPV KG
Sbjct: 19 MRFLQRVRCWEYRQLPSIVRVTRPTRPDKARRLGYKAKQGYVVYRVRVKRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKPTNQGVTQLKFQR+KRSVAEERAGRKLGGLRVLNSYW+NEDST+KYFEVILVD A
Sbjct: 79 IVYGKPTNQGVTQLKFQRTKRSVAEERAGRKLGGLRVLNSYWINEDSTYKYFEVILVDQA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+AIRNDPRINW+ NPVHKHRELRGLTSAGK+ RGL G+GH ++KA PSRRANWKRN TL
Sbjct: 139 HAAIRNDPRINWICNPVHKHRELRGLTSAGKKYRGLRGRGHLHNKAPPSRRANWKRNQTL 198
Query: 181 SLRRYR 186
SL RYR
Sbjct: 199 SLPRYR 204
>gb|AAK67641.1| ribosomal protein L15 [Homo sapiens]
Length = 204
Score = 340 bits (871), Expect = 2e-92
Identities = 159/186 (85%), Positives = 175/186 (93%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQQ +IVR+TRPTRPD+ARRLG+KAKQGYV+YR+RVRRGGRKRPV KG
Sbjct: 19 MRFVQRVRCWEYRQQPAIVRITRPTRPDRARRLGFKAKQGYVIYRIRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP +QG+TQLKFQR+KRSVAEERAGR+LGGLRVLNSYWVNED T+KYFEVILVDVA
Sbjct: 79 IVYGKPKHQGITQLKFQRNKRSVAEERAGRRLGGLRVLNSYWVNEDFTYKYFEVILVDVA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
HSA+RNDPRINWL PVHKHRELRGLTSAGK+ RGL GKGHR+HK RPSRRA WKRN T+
Sbjct: 139 HSAVRNDPRINWLCKPVHKHRELRGLTSAGKKFRGLRGKGHRHHKNRPSRRATWKRNQTV 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>gb|AAC32144.1| probable 60S ribosomal protein L15 [Picea mariana]
gi|6093872|sp|O65082|RL15B_PICMA 60S ribosomal protein
L15-2
Length = 204
Score = 336 bits (862), Expect = 2e-91
Identities = 158/186 (84%), Positives = 171/186 (90%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWEYRQ IVR+T PTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPV KG
Sbjct: 19 MRFLQRVRCWEYRQLPGIVRITHPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP NQG+TQLKFQRS RSVAEERAGRKLGGLRVLNSYW+N+DST+KY+EV+LVD A
Sbjct: 79 IVYGKPKNQGITQLKFQRSLRSVAEERAGRKLGGLRVLNSYWINQDSTYKYYEVVLVDQA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H+ IRNDPRINW+ N VHKHRELRGLTSAGK+ RGL G+GH YHKARPS+RA WKRNNTL
Sbjct: 139 HTVIRNDPRINWICNAVHKHRELRGLTSAGKKYRGLRGRGHLYHKARPSKRATWKRNNTL 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>gb|AAC32112.1| probable 60S ribosomal protein L15 [Picea mariana]
gi|6093871|sp|O65050|RL15A_PICMA 60S ribosomal protein
L15-1
Length = 204
Score = 335 bits (859), Expect = 4e-91
Identities = 158/186 (84%), Positives = 172/186 (91%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+QRVRCWE+RQ IVR+TRP+RPDKARRLGYKAKQGYVVYRVRVRRGGRKRPV KG
Sbjct: 19 MRFLQRVRCWEFRQLPGIVRVTRPSRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
IVYGKP NQG+TQLKFQRS RSVAEERAGRKLGGLRVLNSYW+N+DST+KY+E+ILVD A
Sbjct: 79 IVYGKPKNQGITQLKFQRSLRSVAEERAGRKLGGLRVLNSYWINQDSTYKYYEIILVDQA 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
HSAIR DPRINW+ N VHKHRELRGLTSAGK+ RGL G+GH YHKARPS+RA WKRNNTL
Sbjct: 139 HSAIRKDPRINWICNAVHKHRELRGLTSAGKKYRGLRGRGHLYHKARPSKRATWKRNNTL 198
Query: 181 SLRRYR 186
SLRRYR
Sbjct: 199 SLRRYR 204
>emb|CAG81430.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50551511|ref|XP_503229.1| hypothetical protein
[Yarrowia lipolytica]
Length = 203
Score = 288 bits (737), Expect = 5e-77
Identities = 135/186 (72%), Positives = 155/186 (82%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+ RVRCWEYRQ++ I R +RP+RPDKARRLGYKAKQG+V+YRVRVRRG RKRP KG
Sbjct: 19 LRFLLRVRCWEYRQKTGIHRASRPSRPDKARRLGYKAKQGFVIYRVRVRRGNRKRPSKKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKPTNQG+ QLK QRS R+VAEER GR+ G LRVLNSYWVN+DST+KYFEVILVD +
Sbjct: 79 ATYGKPTNQGINQLKLQRSLRAVAEERVGRRAGNLRVLNSYWVNQDSTYKYFEVILVDPS 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H IRNDPRINW+ NPVHKHRE RGLT+AGK+NRG++ KGH+YH RR WK NTL
Sbjct: 139 HVVIRNDPRINWICNPVHKHRESRGLTAAGKKNRGIN-KGHKYHNTSAGRRHTWKLQNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>emb|CAD21192.1| probable ribosomal protein L15.e.B, cytosolic [Neurospora crassa]
gi|30316252|sp|Q8X034|RL15_NEUCR 60S ribosomal protein
L15 gi|32415471|ref|XP_328215.1| 60S RIBOSOMAL PROTEIN
L15 [Neurospora crassa] gi|28918286|gb|EAA27963.1| 60S
RIBOSOMAL PROTEIN L15 [Neurospora crassa]
Length = 203
Score = 287 bits (734), Expect = 1e-76
Identities = 136/186 (73%), Positives = 154/186 (82%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+ RVRCWE RQ + I R +RP+RPDKARRLGYKAKQGYV+YR RVRRGGRK+PV KG
Sbjct: 19 VRFLLRVRCWELRQLNVIHRASRPSRPDKARRLGYKAKQGYVIYRARVRRGGRKKPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKPTNQGV QLK+QRS +S AEER GR+ LRVLNSYW+N+DST+KYFEVILVD
Sbjct: 79 ATYGKPTNQGVNQLKYQRSLKSTAEERVGRRCANLRVLNSYWINQDSTYKYFEVILVDPQ 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H AIR DPRINW+ NPVHKHRE RGLTS GK +RGL+ KGHRY+K R RR WKR+NTL
Sbjct: 139 HKAIRRDPRINWIVNPVHKHRESRGLTSTGKRSRGLN-KGHRYNKTRAGRRKTWKRHNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>gb|EAK98479.1| likely cytosolic ribosomal protein L15 [Candida albicans SC5314]
gi|46439065|gb|EAK98387.1| likely cytosolic ribosomal
protein L15 [Candida albicans SC5314]
Length = 204
Score = 285 bits (728), Expect = 6e-76
Identities = 133/186 (71%), Positives = 157/186 (83%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+ RVRCWEYRQ++ I R +RP+RPDKARRLGYKAKQG+V+YR+RVRRGGRKRPV KG
Sbjct: 19 MRFLYRVRCWEYRQKNVIHRASRPSRPDKARRLGYKAKQGFVIYRIRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKPTNQGV QLK+Q+S RS AEER GR+ LRVLNSYWVN+DST+KYFEVILVD +
Sbjct: 79 ATYGKPTNQGVNQLKYQKSLRSTAEERVGRRASNLRVLNSYWVNQDSTYKYFEVILVDPS 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H AIR D R NW+ NPVHKHRE RGLTSAGK++RG++ KGH ++K + RR WK++NTL
Sbjct: 139 HKAIRRDARYNWIVNPVHKHREARGLTSAGKKSRGIN-KGHLFNKTKAGRRHTWKKHNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>emb|CAG98580.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50311691|ref|XP_455872.1| unnamed protein product
[Kluyveromyces lactis]
Length = 204
Score = 282 bits (721), Expect = 4e-75
Identities = 131/186 (70%), Positives = 155/186 (82%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+QRVR WEYRQ++ I R +RPTRPDKARRLGYKAKQG+V+YRVRVRRG RKRPVSKG
Sbjct: 19 LRFLQRVRVWEYRQKNVIHRASRPTRPDKARRLGYKAKQGFVIYRVRVRRGNRKRPVSKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKPTNQGV QLK+QRS R+ AEER GR+ LRVLNSYW+N+DST+KYFEVILVD +
Sbjct: 79 ATYGKPTNQGVNQLKYQRSLRATAEERVGRRAANLRVLNSYWINQDSTYKYFEVILVDPS 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H AIR D + NW+ NPVHKHRE RGLT+ GK++RG++ KGH++H + RR WKR NTL
Sbjct: 139 HKAIRRDAKYNWICNPVHKHREARGLTATGKKSRGIN-KGHKFHNTQAGRRKTWKRQNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>gb|AAS53585.1| AFR214Cp [Ashbya gossypii ATCC 10895] gi|45198732|ref|NP_985761.1|
AFR214Cp [Eremothecium gossypii]
Length = 204
Score = 281 bits (720), Expect = 5e-75
Identities = 130/186 (69%), Positives = 155/186 (82%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+QR+R WEYRQ++ I R +RP+RPDKARRLGYKAKQG+V+YRVRVRRG RKRPV KG
Sbjct: 19 LRFLQRIRVWEYRQKTVIHRASRPSRPDKARRLGYKAKQGFVIYRVRVRRGNRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKPTNQGV QLK+QR+ R+ AEER GR+ LRVLNSYW+N+DST+KYFEVILVD +
Sbjct: 79 ATYGKPTNQGVNQLKYQRALRATAEERVGRRAANLRVLNSYWINQDSTYKYFEVILVDPS 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H AIR D RINW+ NPVHKHRE RGLT+ GK++RG++ KGHR+H + RR WKR NTL
Sbjct: 139 HKAIRRDARINWIANPVHKHREARGLTATGKKSRGIN-KGHRFHNTQAGRRKTWKRQNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>gb|AAS59859.1| ribosomal protein L15 [Acipenser gueldenstaedtii]
gi|45386081|gb|AAS59858.1| ribosomal protein L15
[Acipenser schrenckii] gi|45386079|gb|AAS59857.1|
ribosomal protein L15 [Acipenser sinensis]
Length = 204
Score = 281 bits (718), Expect = 8e-75
Identities = 135/187 (72%), Positives = 155/187 (82%), Gaps = 2/187 (1%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+ RVRCW+YRQ S++ R RPTRPDKARRLGYKAKQGYV+YRVRVRRGGRKRPV KG
Sbjct: 19 MRFLLRVRCWQYRQLSNLHRAPRPTRPDKARRLGYKAKQGYVIYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKP N G+ ++KF RS +SVAEERAGR GGLRVLNSYWV ED+T+K+FEVIL+D
Sbjct: 79 ATYGKPVNHGINEIKFARSLQSVAEERAGRHCGGLRVLNSYWVGEDATYKFFEVILIDPF 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYH-KARPSRRANWKRNNT 179
H AIR +P I WLT PVHKHRE+RGLTSAGK++RGL GKGH++H SRRA W+R NT
Sbjct: 139 HKAIRRNPDIQWLTKPVHKHREMRGLTSAGKKSRGL-GKGHKFHLTIGGSRRACWRRRNT 197
Query: 180 LSLRRYR 186
L LRRYR
Sbjct: 198 LQLRRYR 204
>gb|AAH46569.1| Rpl15-prov protein [Xenopus laevis] gi|49670582|gb|AAH75126.1|
Rpl15-prov protein [Xenopus laevis]
Length = 204
Score = 279 bits (713), Expect = 3e-74
Identities = 136/187 (72%), Positives = 152/187 (80%), Gaps = 2/187 (1%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+ RVRCW+YRQ SS+ R RPTRPDKARRLGYKAKQGYV+YR+RVRRGGRKRPV KG
Sbjct: 19 MRFLLRVRCWQYRQLSSLHRAPRPTRPDKARRLGYKAKQGYVIYRIRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKP N G+ QLKF RS +SVAEERAGR GGLRVL+SYWV EDST+K+FEVILVD
Sbjct: 79 ATYGKPVNHGINQLKFARSLQSVAEERAGRHCGGLRVLSSYWVGEDSTYKFFEVILVDTF 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYH-KARPSRRANWKRNNT 179
H AIR +P W+T VHKHRE+RGLTSAGK++RGL GKGH+YH SRRA WKR NT
Sbjct: 139 HKAIRRNPDTQWITKAVHKHREMRGLTSAGKKSRGL-GKGHKYHITIGGSRRACWKRRNT 197
Query: 180 LSLRRYR 186
L L RYR
Sbjct: 198 LQLHRYR 204
>emb|CAG89381.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50424843|ref|XP_461011.1| unnamed protein product
[Debaryomyces hansenii]
Length = 204
Score = 278 bits (712), Expect = 4e-74
Identities = 130/186 (69%), Positives = 152/186 (80%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+ RVRCWEYRQ++ I R +RP+RPDKARRLGYKAKQG+V+YRVRVRRGGRKRP KG
Sbjct: 19 MRFLHRVRCWEYRQKTVIHRASRPSRPDKARRLGYKAKQGFVIYRVRVRRGGRKRPAPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKP N GV QLK+QR+ R+ AEER GR+ LRVLNSYWVNEDST+KYFEVIL+D +
Sbjct: 79 ATYGKPVNHGVNQLKYQRALRATAEERVGRRASNLRVLNSYWVNEDSTYKYFEVILLDPS 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H AIR D R NW+ NPVHKHRE RGLTS GK++RG++ KGH ++ + RR WKRNNTL
Sbjct: 139 HKAIRRDARYNWIANPVHKHREARGLTSTGKKSRGIN-KGHLFNNTKAGRRHTWKRNNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>emb|CAG57808.1| unnamed protein product [Candida glabrata CBS138]
gi|50284973|ref|XP_444915.1| unnamed protein product
[Candida glabrata]
Length = 204
Score = 278 bits (710), Expect = 7e-74
Identities = 130/186 (69%), Positives = 154/186 (81%), Gaps = 1/186 (0%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
+RF+QRVR WEYRQ++ I R +RP+RPDKARR+GYKAKQG+V+YRVRVRRG RKRPV KG
Sbjct: 19 LRFLQRVRVWEYRQKNVIHRASRPSRPDKARRMGYKAKQGFVIYRVRVRRGNRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKPTNQGV +LK+QRS R+ AEER GR+ LRVLNSYWVN+DST+KYFEVILVD +
Sbjct: 79 ATYGKPTNQGVNELKYQRSLRATAEERVGRRASNLRVLNSYWVNQDSTYKYFEVILVDPS 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYHKARPSRRANWKRNNTL 180
H AIR D R NW+ NPVHKHRE RGLT+ GK++RG++ KGHRY + RR WKR+NTL
Sbjct: 139 HKAIRRDARYNWICNPVHKHREARGLTATGKKSRGIN-KGHRYTNTKAGRRKTWKRHNTL 197
Query: 181 SLRRYR 186
SL RYR
Sbjct: 198 SLWRYR 203
>gb|AAK95142.1| ribosomal protein L15 [Ictalurus punctatus]
gi|47606044|sp|Q90YV2|RL15_ICTPU 60S ribosomal protein
L15
Length = 204
Score = 278 bits (710), Expect = 7e-74
Identities = 135/187 (72%), Positives = 153/187 (81%), Gaps = 2/187 (1%)
Query: 1 MRFMQRVRCWEYRQQSSIVRLTRPTRPDKARRLGYKAKQGYVVYRVRVRRGGRKRPVSKG 60
MRF+ RVRCW+YRQ S++ R RPTRPDKARRLGYKAKQGYV+YRVRVRRGGRKRPV KG
Sbjct: 19 MRFLLRVRCWQYRQLSALHRAPRPTRPDKARRLGYKAKQGYVIYRVRVRRGGRKRPVPKG 78
Query: 61 IVYGKPTNQGVTQLKFQRSKRSVAEERAGRKLGGLRVLNSYWVNEDSTFKYFEVILVDVA 120
YGKP + GV QLKF RS +SVAEERAGR GGLRVLNSYWV EDST+K+FEVIL+D
Sbjct: 79 ATYGKPVHHGVNQLKFARSLQSVAEERAGRHCGGLRVLNSYWVGEDSTYKFFEVILIDTF 138
Query: 121 HSAIRNDPRINWLTNPVHKHRELRGLTSAGKENRGLSGKGHRYH-KARPSRRANWKRNNT 179
H AIR +P W+T VHKHRE+RGLTSAGK++RGL GKGH++H SRRA W+R NT
Sbjct: 139 HKAIRRNPDTQWITKAVHKHREMRGLTSAGKKSRGL-GKGHKFHLTIGGSRRAPWRRRNT 197
Query: 180 LSLRRYR 186
L LRRYR
Sbjct: 198 LQLRRYR 204
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 325,222,073
Number of Sequences: 2540612
Number of extensions: 13026354
Number of successful extensions: 26311
Number of sequences better than 10.0: 198
Number of HSP's better than 10.0 without gapping: 187
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 26003
Number of HSP's gapped (non-prelim): 203
length of query: 186
length of database: 863,360,394
effective HSP length: 120
effective length of query: 66
effective length of database: 558,486,954
effective search space: 36860138964
effective search space used: 36860138964
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 71 (32.0 bits)
Medicago: description of AC146855.3


