

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146819.13 + phase: 0
(97 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
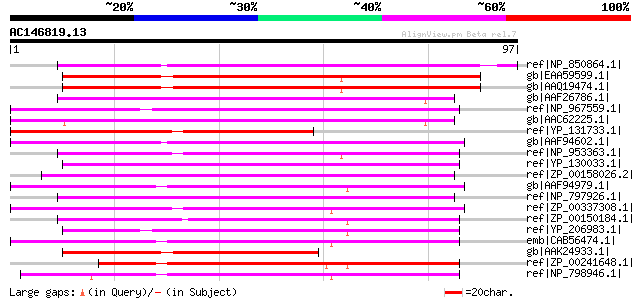
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_850864.1| two-component responsive regulator family prote... 63 2e-09
gb|EAA59599.1| hypothetical protein AN7945.2 [Aspergillus nidula... 58 5e-08
gb|AAQ19474.1| histidine kinase G2 [Emericella nidulans] 58 5e-08
gb|AAF26786.1| putative response regulator protein (receiver com... 55 4e-07
ref|NP_967559.1| two component sensor histidine kinase [Bdellovi... 55 6e-07
gb|AAC62225.1| response regulator protein [Brassica napus] 54 1e-06
ref|YP_131733.1| putative BaeS, Signal transduction histidine ki... 53 2e-06
gb|AAF94602.1| sensor histidine kinase/response regulator [Vibri... 52 4e-06
ref|NP_953363.1| sensory box histidine kinase/response regulator... 51 6e-06
ref|YP_130033.1| hypothetical protein PBPRA1827 [Photobacterium ... 51 8e-06
ref|ZP_00158026.2| COG0642: Signal transduction histidine kinase... 51 8e-06
gb|AAF94979.1| sensor histidine kinase [Vibrio cholerae O1 biova... 50 1e-05
ref|NP_797926.1| sensor histidine kinase/response regulator [Vib... 50 1e-05
ref|ZP_00337308.1| COG0642: Signal transduction histidine kinase... 50 1e-05
ref|ZP_00150184.1| COG0642: Signal transduction histidine kinase... 49 2e-05
ref|YP_206983.1| transcriptional regulator [Vibrio fischeri ES11... 49 3e-05
emb|CAB56474.1| putative histidine kinase [Pseudomonas stutzeri] 49 4e-05
gb|AAK24933.1| sensory box histidine kinase/response regulator [... 49 4e-05
ref|ZP_00241648.1| COG0642: Signal transduction histidine kinase... 49 4e-05
ref|NP_798946.1| sensor histidine kinase/response regulator [Vib... 48 5e-05
>ref|NP_850864.1| two-component responsive regulator family protein / response
regulator family protein [Arabidopsis thaliana]
Length = 139
Score = 62.8 bits (151), Expect = 2e-09
Identities = 35/88 (39%), Positives = 52/88 (58%), Gaps = 4/88 (4%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSR 69
+G++ V NGKE++D+ + +DLILM MP+M+GI ATK+LR++ +I GV+
Sbjct: 40 LGIKNDVVTNGKEAVDVYCS-GGNYDLILMDMDMPIMNGIQATKRLREMGIESKIAGVTT 98
Query: 70 HLTNDEYAEFCNAGLDDLHHEDGIPLAI 97
E EF AGL+D + PL I
Sbjct: 99 RANEGEKKEFMEAGLNDFQEK---PLTI 123
>gb|EAA59599.1| hypothetical protein AN7945.2 [Aspergillus nidulans FGSC A4]
gi|67901916|ref|XP_681214.1| hypothetical protein
AN7945_2 [Aspergillus nidulans FGSC A4]
gi|49115277|ref|XP_412082.1| hypothetical protein
AN7945.2 [Aspergillus nidulans FGSC A4]
Length = 676
Score = 58.2 bits (139), Expect = 5e-08
Identities = 34/84 (40%), Positives = 51/84 (60%), Gaps = 6/84 (7%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPG----QIVG 66
G E NG+E+L L + KFD ILM + MPVMDG TAT+++R +E I+G
Sbjct: 577 GFEVTTANNGQEALGLWDS--DKFDCILMDQEMPVMDGNTATREIRAMEKEKGSHIPILG 634
Query: 67 VSRHLTNDEYAEFCNAGLDDLHHE 90
V+ ++ D+ A+ NAG+D + H+
Sbjct: 635 VTANVRQDQQADMLNAGMDGIIHK 658
>gb|AAQ19474.1| histidine kinase G2 [Emericella nidulans]
Length = 659
Score = 58.2 bits (139), Expect = 5e-08
Identities = 34/84 (40%), Positives = 51/84 (60%), Gaps = 6/84 (7%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPG----QIVG 66
G E NG+E+L L + KFD ILM + MPVMDG TAT+++R +E I+G
Sbjct: 560 GFEVTTANNGQEALGLWDS--DKFDCILMDQEMPVMDGNTATREIRAMEKEKGSHIPILG 617
Query: 67 VSRHLTNDEYAEFCNAGLDDLHHE 90
V+ ++ D+ A+ NAG+D + H+
Sbjct: 618 VTANVRQDQQADMLNAGMDGIIHK 641
>gb|AAF26786.1| putative response regulator protein (receiver component)
[Arabidopsis thaliana] gi|21553766|gb|AAM62859.1|
putative response regulator protein (receiver component)
[Arabidopsis thaliana] gi|51316101|sp|Q9M8Y4|ARR22_ARATH
Two-component response regulator ARR22
gi|30679083|ref|NP_850511.1| two-component responsive
regulator family protein / response regulator family
protein [Arabidopsis thaliana]
gi|15229263|ref|NP_187078.1| two-component responsive
regulator family protein / response regulator family
protein [Arabidopsis thaliana]
Length = 142
Score = 55.1 bits (131), Expect = 4e-07
Identities = 32/77 (41%), Positives = 46/77 (59%), Gaps = 1/77 (1%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSR 69
IG + KNG+E++ L E FDLILM + MP DG++ TKKLR+++ IVGV+
Sbjct: 44 IGGISQTAKNGEEAVILHRDGEASFDLILMDKEMPERDGVSTTKKLREMKVTSMIVGVTS 103
Query: 70 HLTNDEYAE-FCNAGLD 85
+E + F AGL+
Sbjct: 104 VADQEEERKAFMEAGLN 120
>ref|NP_967559.1| two component sensor histidine kinase [Bdellovibrio bacteriovorus
HD100] gi|39574710|emb|CAE78552.1| two component sensor
histidine kinase [Bdellovibrio bacteriovorus HD100]
Length = 666
Score = 54.7 bits (130), Expect = 6e-07
Identities = 31/86 (36%), Positives = 44/86 (51%), Gaps = 2/86 (2%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY 60
+++ L G NG+E L +E++FD+ILM MPVMDG TAT+KLR Y
Sbjct: 560 LLIQRMLAKRGANLTLATNGQEGLT--KALEKEFDIILMDIQMPVMDGYTATRKLRQAGY 617
Query: 61 PGQIVGVSRHLTNDEYAEFCNAGLDD 86
I+ ++ H D+ AG D
Sbjct: 618 SKPIIALTAHAMKDDRERCLEAGCTD 643
>gb|AAC62225.1| response regulator protein [Brassica napus]
Length = 136
Score = 53.9 bits (128), Expect = 1e-06
Identities = 34/87 (39%), Positives = 50/87 (57%), Gaps = 2/87 (2%)
Query: 1 MILFEWLID-IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
+I+ E +I IG + NG+E++ + FDLILM + MP DG++ TKKLR++E
Sbjct: 28 LIIHEKIIKAIGGISQTANNGEEAVIIHRDGGSSFDLILMDKEMPERDGVSTTKKLREME 87
Query: 60 YPGQIVGVSRHLTNDEYAE-FCNAGLD 85
IVGV+ N+E F AGL+
Sbjct: 88 VKSMIVGVTSLADNEEERRAFMEAGLN 114
>ref|YP_131733.1| putative BaeS, Signal transduction histidine kinase [Photobacterium
profundum SS9] gi|46915160|emb|CAG21933.1| putative
BaeS, Signal transduction histidine kinase
[Photobacterium profundum SS9]
Length = 971
Score = 53.1 bits (126), Expect = 2e-06
Identities = 29/58 (50%), Positives = 39/58 (67%), Gaps = 2/58 (3%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDL 58
+I+ +L G THAV+NG E+L ++ V FD+ILM MPVMDGI ATK++R L
Sbjct: 668 LIIESFLKQYGFNTHAVENGAEALKILEHVY--FDVILMDNHMPVMDGIEATKRIRAL 723
>gb|AAF94602.1| sensor histidine kinase/response regulator [Vibrio cholerae O1
biovar eltor str. N16961] gi|15641456|ref|NP_231088.1|
sensor histidine kinase/response regulator [Vibrio
cholerae O1 biovar eltor str. N16961]
gi|11356143|pir||E82198 sensor histidine kinase/response
regulator VC1445 [imported] - Vibrio cholerae (strain
N16961 serogroup O1)
Length = 572
Score = 52.0 bits (123), Expect = 4e-06
Identities = 28/87 (32%), Positives = 48/87 (54%), Gaps = 1/87 (1%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY 60
M++ L +G + +NG ++++L+ FDL+LM MP+MDGI ATK LRD
Sbjct: 467 MVIQLLLNKLGYDVTIAENGLQAVELLEK-NHVFDLVLMDISMPIMDGIAATKILRDKHI 525
Query: 61 PGQIVGVSRHLTNDEYAEFCNAGLDDL 87
I+ ++ H + +AG++D+
Sbjct: 526 EIPIIALTAHTAGSDKQNCIDAGMNDI 552
>ref|NP_953363.1| sensory box histidine kinase/response regulator [Geobacter
sulfurreducens PCA] gi|39984303|gb|AAR35690.1| sensory
box histidine kinase/response regulator [Geobacter
sulfurreducens PCA]
Length = 982
Score = 51.2 bits (121), Expect = 6e-06
Identities = 28/82 (34%), Positives = 47/82 (57%), Gaps = 7/82 (8%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPG-----QI 64
+G+ A NG E+LDL++T +DL+LM MPVMDG AT+ +RD+ P +
Sbjct: 747 LGLRGDAAANGAEALDLLATTP--YDLVLMDVQMPVMDGFEATRHIRDVRSPVLNHEIPV 804
Query: 65 VGVSRHLTNDEYAEFCNAGLDD 86
+ ++ H + + +AG++D
Sbjct: 805 IALTAHAMKGDRRKCLDAGMND 826
>ref|YP_130033.1| hypothetical protein PBPRA1827 [Photobacterium profundum SS9]
gi|46913445|emb|CAG20231.1| hypothetical protein
[Photobacterium profundum SS9]
Length = 693
Score = 50.8 bits (120), Expect = 8e-06
Identities = 25/76 (32%), Positives = 42/76 (54%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSRH 70
G NG+E++ + FD++LM MPV+DGI ATK+LR+ E I+ ++ +
Sbjct: 596 GHRVQIANNGEEAIKALENNTHYFDIVLMDISMPVLDGIQATKRLREEEITIPIIALTAN 655
Query: 71 LTNDEYAEFCNAGLDD 86
+ E+ AG++D
Sbjct: 656 AMQCDQVEYKKAGMND 671
>ref|ZP_00158026.2| COG0642: Signal transduction histidine kinase [Anabaena variabilis
ATCC 29413]
Length = 709
Score = 50.8 bits (120), Expect = 8e-06
Identities = 27/79 (34%), Positives = 47/79 (59%)
Query: 7 LIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVG 66
L D +T KNG E++ L + + K ++LM +MP MDG+TA + L+++ +I+
Sbjct: 606 LEDYNYKTLTAKNGIEAIALYAQHQDKISMVLMDIMMPSMDGLTAIRILQEMNPQLKIIA 665
Query: 67 VSRHLTNDEYAEFCNAGLD 85
+S N++ AE NAG++
Sbjct: 666 ISGITGNNQLAEAANAGVN 684
>gb|AAF94979.1| sensor histidine kinase [Vibrio cholerae O1 biovar eltor str.
N16961] gi|15641833|ref|NP_231465.1| sensor histidine
kinase [Vibrio cholerae O1 biovar eltor str. N16961]
gi|11356137|pir||C82151 sensor histidine kinase VC1831
[imported] - Vibrio cholerae (strain N16961 serogroup
O1)
Length = 736
Score = 50.4 bits (119), Expect = 1e-05
Identities = 30/90 (33%), Positives = 50/90 (55%), Gaps = 5/90 (5%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY 60
MIL ++ + G E H+V +G +++ + E FDL+LM MP+ DGI AT+++R L
Sbjct: 615 MILEAFMRNKGFECHSVMDGVQAITALQ--ESSFDLVLMDNHMPLKDGIQATREIRQLPL 672
Query: 61 PGQ---IVGVSRHLTNDEYAEFCNAGLDDL 87
P + G + + D + +AG DD+
Sbjct: 673 PQAKILLFGCTADVFKDTRDKMLSAGADDI 702
>ref|NP_797926.1| sensor histidine kinase/response regulator [Vibrio parahaemolyticus
RIMD 2210633] gi|28806538|dbj|BAC59810.1| sensor
histidine kinase/response regulator [Vibrio
parahaemolyticus RIMD 2210633]
Length = 576
Score = 50.1 bits (118), Expect = 1e-05
Identities = 26/76 (34%), Positives = 41/76 (53%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQIVGVSR 69
+G H +G E+L + + + D+ILM MPVMDGITAT+ +R IV ++
Sbjct: 478 LGHNVHIASHGAEALTFLEENDTRIDMILMDVSMPVMDGITATRLIRKKGITIPIVALTA 537
Query: 70 HLTNDEYAEFCNAGLD 85
H + + +AG+D
Sbjct: 538 HALESDKDKCLDAGMD 553
>ref|ZP_00337308.1| COG0642: Signal transduction histidine kinase [Silicibacter sp.
TM1040]
Length = 699
Score = 50.1 bits (118), Expect = 1e-05
Identities = 29/92 (31%), Positives = 52/92 (56%), Gaps = 7/92 (7%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY 60
+IL ++L +V NG+E++ L+ + FD+ILM MP MDG+ T+K+R +E
Sbjct: 588 LILEKFLSSTPYRVKSVSNGQEAIGLVHSFS--FDVILMDIQMPRMDGVETTQKIRSIEQ 645
Query: 61 -----PGQIVGVSRHLTNDEYAEFCNAGLDDL 87
P IV V+ + ++ + + + G+DD+
Sbjct: 646 AKGREPSFIVAVTGNAMPEQLSHYLSVGMDDV 677
>ref|ZP_00150184.1| COG0642: Signal transduction histidine kinase [Dechloromonas
aromatica RCB]
Length = 804
Score = 49.3 bits (116), Expect = 2e-05
Identities = 30/82 (36%), Positives = 45/82 (54%), Gaps = 6/82 (7%)
Query: 10 IGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQ-----I 64
+G+E +A +NG E+ + ER DLILM MP+MDG+ AT+K+R E I
Sbjct: 698 LGIEANAAENGLEATQAATAAERP-DLILMDVQMPLMDGLEATRKIRAWEQANNQPRLVI 756
Query: 65 VGVSRHLTNDEYAEFCNAGLDD 86
V ++ ++Y AG+DD
Sbjct: 757 VALTAGAFAEDYEHCIAAGMDD 778
>ref|YP_206983.1| transcriptional regulator [Vibrio fischeri ES114]
gi|59482456|gb|AAW88095.1| transcriptional regulator
[Vibrio fischeri ES114]
Length = 766
Score = 48.9 bits (115), Expect = 3e-05
Identities = 28/79 (35%), Positives = 42/79 (52%), Gaps = 5/79 (6%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEYPGQ---IVGV 67
G +NGKE++ + ++DLI M MP MDG+TATKK+R L Q I+ +
Sbjct: 522 GYSIDIAQNGKEAIT--KAINTQYDLIFMDISMPEMDGMTATKKIRSLSDTHQNMPIIAL 579
Query: 68 SRHLTNDEYAEFCNAGLDD 86
+ H + + F AG+ D
Sbjct: 580 TAHALSGDKERFIEAGMTD 598
>emb|CAB56474.1| putative histidine kinase [Pseudomonas stutzeri]
Length = 417
Score = 48.5 bits (114), Expect = 4e-05
Identities = 30/91 (32%), Positives = 50/91 (53%), Gaps = 7/91 (7%)
Query: 1 MILFEWLIDIGVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY 60
M+L L D+G E AV +G+++L+++ ++ FD++ M MP MDG T+ +R E
Sbjct: 181 MLLETLLTDMGGEVVAVSSGQQALEVVQ--QQSFDMVFMDVQMPGMDGRQTTEAIRRWEL 238
Query: 61 -----PGQIVGVSRHLTNDEYAEFCNAGLDD 86
P IV ++ H ++E +GLDD
Sbjct: 239 ESGQPPLPIVALTAHALSNERRSLLQSGLDD 269
>gb|AAK24933.1| sensory box histidine kinase/response regulator [Caulobacter
crescentus CB15] gi|16127201|ref|NP_421765.1| sensory
box histidine kinase/response regulator [Caulobacter
crescentus CB15] gi|25397492|pir||A87617 sensory box
histidine kinase/response regulator [imported] -
Caulobacter crescentus
Length = 713
Score = 48.5 bits (114), Expect = 4e-05
Identities = 25/49 (51%), Positives = 36/49 (73%), Gaps = 2/49 (4%)
Query: 11 GVETHAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE 59
G E +NG E+L +++T ++FDLILM MPVMDG+TAT+++R LE
Sbjct: 610 GAELVMAENGAEALTILAT--QRFDLILMDMQMPVMDGLTATREVRCLE 656
>ref|ZP_00241648.1| COG0642: Signal transduction histidine kinase [Rubrivivax
gelatinosus PM1]
Length = 527
Score = 48.5 bits (114), Expect = 4e-05
Identities = 29/75 (38%), Positives = 48/75 (63%), Gaps = 8/75 (10%)
Query: 18 KNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLE--YPGQ----IVGVSRHL 71
+NG+E+LD ++ E KFD++LM MPV+DGI+A ++LR E PG +V ++ +
Sbjct: 415 RNGQEALDALA--EHKFDVVLMDVQMPVLDGISAVRELRARESSNPGSGRTTVVAMTANT 472
Query: 72 TNDEYAEFCNAGLDD 86
+ A+ +AG+DD
Sbjct: 473 EPQDIAQCRSAGMDD 487
>ref|NP_798946.1| sensor histidine kinase/response regulator [Vibrio parahaemolyticus
RIMD 2210633] gi|28807577|dbj|BAC60830.1| sensor
histidine kinase/response regulator [Vibrio
parahaemolyticus RIMD 2210633]
Length = 932
Score = 48.1 bits (113), Expect = 5e-05
Identities = 31/87 (35%), Positives = 50/87 (56%), Gaps = 5/87 (5%)
Query: 3 LFEWLIDIGVET-HAVKNGKESLDLISTVERKFDLILMARIMPVMDGITATKKLRDLEY- 60
L L+ VET A NG+++++L + E+K+DLI M MP+MDG+TA + +++LE
Sbjct: 682 LISALLKERVETVTACSNGQQAVNLAT--EKKYDLIFMDIQMPLMDGVTACQNIKELELN 739
Query: 61 -PGQIVGVSRHLTNDEYAEFCNAGLDD 86
++ V+ H E AG+DD
Sbjct: 740 KDTPVIAVTAHAMAGERDRLLAAGMDD 766
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.142 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 168,719,613
Number of Sequences: 2540612
Number of extensions: 5878194
Number of successful extensions: 15319
Number of sequences better than 10.0: 1535
Number of HSP's better than 10.0 without gapping: 607
Number of HSP's successfully gapped in prelim test: 928
Number of HSP's that attempted gapping in prelim test: 14562
Number of HSP's gapped (non-prelim): 1608
length of query: 97
length of database: 863,360,394
effective HSP length: 73
effective length of query: 24
effective length of database: 677,895,718
effective search space: 16269497232
effective search space used: 16269497232
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146819.13


