

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146817.2 + phase: 0
(145 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
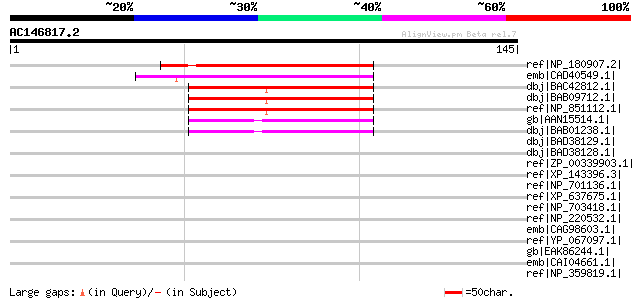
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_180907.2| hydroxyproline-rich glycoprotein family protein... 65 4e-10
emb|CAD40549.1| OSJNBa0072K14.9 [Oryza sativa (japonica cultivar... 48 6e-05
dbj|BAC42812.1| unknown protein [Arabidopsis thaliana] gi|306937... 47 7e-05
dbj|BAB09712.1| unnamed protein product [Arabidopsis thaliana] 47 7e-05
ref|NP_851112.1| expressed protein [Arabidopsis thaliana] 47 7e-05
gb|AAN15514.1| unknown protein [Arabidopsis thaliana] gi|2253102... 45 3e-04
dbj|BAB01238.1| unnamed protein product [Arabidopsis thaliana] 45 3e-04
dbj|BAD38129.1| hydroxyproline-rich glycoprotein-like [Oryza sat... 42 0.003
dbj|BAD38128.1| hydroxyproline-rich glycoprotein-like [Oryza sat... 42 0.003
ref|ZP_00339903.1| COG0086: DNA-directed RNA polymerase, beta' s... 37 0.13
ref|XP_143396.3| PREDICTED: similar to ifapsoriasin [Mus musculus] 36 0.22
ref|NP_701136.1| hypothetical protein PF11_0276 [Plasmodium falc... 35 0.29
ref|XP_637675.1| hypothetical protein DDB0218841 [Dictyostelium ... 35 0.29
ref|NP_703418.1| hypothetical protein [Plasmodium falciparum 3D7... 34 0.84
ref|NP_220532.1| DNA-DIRECTED RNA POLYMERASE BETA PRIME CHAIN (r... 34 0.84
emb|CAG98603.1| unnamed protein product [Kluyveromyces lactis NR... 34 0.84
ref|YP_067097.1| DNA-directed RNA polymerase beta prime subunit;... 34 0.84
gb|EAK86244.1| hypothetical protein UM04789.1 [Ustilago maydis 5... 34 0.84
emb|CAI04661.1| conserved hypothetical protein [Plasmodium berghei] 34 0.84
ref|NP_359819.1| DNA-directed RNA polymerase beta prime chain [E... 33 1.1
>ref|NP_180907.2| hydroxyproline-rich glycoprotein family protein [Arabidopsis
thaliana]
Length = 623
Score = 64.7 bits (156), Expect = 4e-10
Identities = 31/61 (50%), Positives = 42/61 (68%), Gaps = 2/61 (3%)
Query: 44 NSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEAT 103
N L++ E+ M+ CDEKR VYE M+ + +EKG+S GKGE + + LQ AHD+YE E T
Sbjct: 124 NELRIVEE--MQRLCDEKRNVYEGMLTRQREKGRSKGGKGETFSPQQLQEAHDDYENETT 181
Query: 104 L 104
L
Sbjct: 182 L 182
>emb|CAD40549.1| OSJNBa0072K14.9 [Oryza sativa (japonica cultivar-group)]
gi|50923911|ref|XP_472316.1| OSJNBa0072K14.9 [Oryza
sativa (japonica cultivar-group)]
Length = 624
Score = 47.8 bits (112), Expect = 6e-05
Identities = 28/70 (40%), Positives = 40/70 (57%), Gaps = 2/70 (2%)
Query: 37 NQIFEDSNSL--KVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAA 94
N I S SL +++ + MK CD KR+ YE M +EKG+S K E ++S LQA
Sbjct: 114 NTITNPSESLLKELQVVEEMKELCDHKRQEYEAMRAAYREKGRSRHSKTETLSSEQLQAY 173
Query: 95 HDEYEEEATL 104
+Y+E+A L
Sbjct: 174 FLDYQEDAAL 183
>dbj|BAC42812.1| unknown protein [Arabidopsis thaliana] gi|30693745|ref|NP_198926.2|
expressed protein [Arabidopsis thaliana]
Length = 582
Score = 47.4 bits (111), Expect = 7e-05
Identities = 22/54 (40%), Positives = 37/54 (67%), Gaps = 1/54 (1%)
Query: 52 DNMKHHCDEKREVYEYMIIQP-KEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK C+EKR+V ++M+++ K+K + KGE + R L+ A DE ++EATL
Sbjct: 131 EDMKQQCEEKRDVVKHMLMEHVKDKVQVKGTKGERLIRRQLETARDELQDEATL 184
>dbj|BAB09712.1| unnamed protein product [Arabidopsis thaliana]
Length = 534
Score = 47.4 bits (111), Expect = 7e-05
Identities = 22/54 (40%), Positives = 37/54 (67%), Gaps = 1/54 (1%)
Query: 52 DNMKHHCDEKREVYEYMIIQP-KEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK C+EKR+V ++M+++ K+K + KGE + R L+ A DE ++EATL
Sbjct: 60 EDMKQQCEEKRDVVKHMLMEHVKDKVQVKGTKGERLIRRQLETARDELQDEATL 113
>ref|NP_851112.1| expressed protein [Arabidopsis thaliana]
Length = 586
Score = 47.4 bits (111), Expect = 7e-05
Identities = 22/54 (40%), Positives = 37/54 (67%), Gaps = 1/54 (1%)
Query: 52 DNMKHHCDEKREVYEYMIIQP-KEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK C+EKR+V ++M+++ K+K + KGE + R L+ A DE ++EATL
Sbjct: 131 EDMKQQCEEKRDVVKHMLMEHVKDKVQVKGTKGERLIRRQLETARDELQDEATL 184
>gb|AAN15514.1| unknown protein [Arabidopsis thaliana] gi|22531026|gb|AAM97017.1|
unknown protein [Arabidopsis thaliana]
gi|30688552|ref|NP_189326.2| hydroxyproline-rich
glycoprotein family protein [Arabidopsis thaliana]
Length = 608
Score = 45.4 bits (106), Expect = 3e-04
Identities = 22/53 (41%), Positives = 32/53 (59%), Gaps = 2/53 (3%)
Query: 52 DNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK CD KR VYE ++ KEKG+ S KGE + A+ E+ +EAT+
Sbjct: 132 EDMKQQCDGKRNVYEMSLV--KEKGRPKSSKGERHIPPESRPAYSEFHDEATM 182
>dbj|BAB01238.1| unnamed protein product [Arabidopsis thaliana]
Length = 621
Score = 45.4 bits (106), Expect = 3e-04
Identities = 22/53 (41%), Positives = 32/53 (59%), Gaps = 2/53 (3%)
Query: 52 DNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
++MK CD KR VYE ++ KEKG+ S KGE + A+ E+ +EAT+
Sbjct: 132 EDMKQQCDGKRNVYEMSLV--KEKGRPKSSKGERHIPPESRPAYSEFHDEATM 182
>dbj|BAD38129.1| hydroxyproline-rich glycoprotein-like [Oryza sativa (japonica
cultivar-group)]
Length = 320
Score = 42.0 bits (97), Expect = 0.003
Identities = 20/53 (37%), Positives = 29/53 (53%)
Query: 52 DNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
+ MK CD KR+ YE M +KG S K E ++ L A+ EY+E++ L
Sbjct: 136 EEMKQQCDMKRDAYETMRASYSDKGGSRHSKTESFSTEQLDASFLEYQEDSAL 188
>dbj|BAD38128.1| hydroxyproline-rich glycoprotein-like [Oryza sativa (japonica
cultivar-group)]
Length = 623
Score = 42.0 bits (97), Expect = 0.003
Identities = 20/53 (37%), Positives = 29/53 (53%)
Query: 52 DNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATL 104
+ MK CD KR+ YE M +KG S K E ++ L A+ EY+E++ L
Sbjct: 136 EEMKQQCDMKRDAYETMRASYSDKGGSRHSKTESFSTEQLDASFLEYQEDSAL 188
>ref|ZP_00339903.1| COG0086: DNA-directed RNA polymerase, beta' subunit/160 kD subunit
[Rickettsia akari str. Hartford]
Length = 1372
Score = 36.6 bits (83), Expect = 0.13
Identities = 24/65 (36%), Positives = 32/65 (48%), Gaps = 3/65 (4%)
Query: 67 YMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATLAKQGCPHIAKLGKGRRIKPLKH 126
Y++I P G S KGE +T LQ A D+Y E+A A G I ++ K LKH
Sbjct: 143 YVVIDP---GLSILQKGELLTEEELQKAKDKYGEDAFTASIGAEVIQQMLKELDFAKLKH 199
Query: 127 SNFNE 131
+ E
Sbjct: 200 ELYEE 204
>ref|XP_143396.3| PREDICTED: similar to ifapsoriasin [Mus musculus]
Length = 704
Score = 35.8 bits (81), Expect = 0.22
Identities = 26/89 (29%), Positives = 38/89 (42%), Gaps = 8/89 (8%)
Query: 24 GKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNS--- 80
G GP + RG N++ E + L E+ KHH ++ + KEK S S
Sbjct: 145 GSRGPAKHRRGSNSKRLERQDELSSSEESRKKHH----GSIFGHSWSSNKEKDGSRSEEL 200
Query: 81 -GKGEHITSRPLQAAHDEYEEEATLAKQG 108
KG+ P + + +EYE L QG
Sbjct: 201 GEKGDKSYDSPSRESEEEYESGYRLNHQG 229
>ref|NP_701136.1| hypothetical protein PF11_0276 [Plasmodium falciparum 3D7]
gi|23496201|gb|AAN35860.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 682
Score = 35.4 bits (80), Expect = 0.29
Identities = 18/57 (31%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Query: 11 ALTSKVDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEY 67
A + D NIK+ G G ++ +++ F D+ LK K N+ +CD K Y+Y
Sbjct: 358 ATQEREDANIKRGGTLGCDKIKEKESSEYFVDNKKLKNK-SSNLTDNCDNKINKYDY 413
>ref|XP_637675.1| hypothetical protein DDB0218841 [Dictyostelium discoideum]
gi|60466108|gb|EAL64174.1| hypothetical protein
DDB0218841 [Dictyostelium discoideum]
Length = 4592
Score = 35.4 bits (80), Expect = 0.29
Identities = 27/87 (31%), Positives = 38/87 (43%), Gaps = 2/87 (2%)
Query: 15 KVDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKE 74
K DK N I NN +D N K KEKD K EK++ E + ++ K+
Sbjct: 450 KDDKKDDINSSSSSIGSSNSSNNTPTKDKN--KEKEKDKEKEKEKEKKKEKEKLKLEEKK 507
Query: 75 KGKSNSGKGEHITSRPLQAAHDEYEEE 101
K K GK + S+ + E+E E
Sbjct: 508 KKKEEKGKSKSKDSKKNKIKGIEFEVE 534
>ref|NP_703418.1| hypothetical protein [Plasmodium falciparum 3D7]
gi|23504558|emb|CAD51438.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 893
Score = 33.9 bits (76), Expect = 0.84
Identities = 26/84 (30%), Positives = 38/84 (44%), Gaps = 7/84 (8%)
Query: 1 MRPSKLHQIAALTSKV----DKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKH 56
M+ + H+I +TS D N N F + NN I+ N K K K+++K
Sbjct: 333 MKKLREHKIIIVTSSGKIYDDDNNNDNNNFYNDNI---YNNNIYNVHNDDKEKIKNHIKK 389
Query: 57 HCDEKREVYEYMIIQPKEKGKSNS 80
+ +YEY Q E+ KSNS
Sbjct: 390 KNKQNNYLYEYQRTQKDEEQKSNS 413
>ref|NP_220532.1| DNA-DIRECTED RNA POLYMERASE BETA PRIME CHAIN (rpoC) [Rickettsia
prowazekii str. Madrid E] gi|3860708|emb|CAA14609.1|
DNA-DIRECTED RNA POLYMERASE BETA PRIME CHAIN (rpoC)
[Rickettsia prowazekii] gi|7434730|pir||B71724
dna-directed RNA polymerase beta prime chain (rpoC)
RP141 - Rickettsia prowazekii
gi|6226044|sp|Q9ZE20|RPOC_RICPR DNA-directed RNA
polymerase beta' chain (RNAP beta' subunit)
(Transcriptase beta' chain) (RNA polymerase beta'
subunit)
Length = 1372
Score = 33.9 bits (76), Expect = 0.84
Identities = 22/65 (33%), Positives = 32/65 (48%), Gaps = 3/65 (4%)
Query: 67 YMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATLAKQGCPHIAKLGKGRRIKPLKH 126
Y+++ P G S KGE +T LQ A D+Y E+A A G I ++ K LK
Sbjct: 143 YVVVDP---GLSILQKGELLTEEELQKAKDKYGEDAFTASIGAEVIQQMLKELDFSKLKQ 199
Query: 127 SNFNE 131
++E
Sbjct: 200 ELYDE 204
>emb|CAG98603.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50312991|ref|XP_455895.1| unnamed protein product
[Kluyveromyces lactis]
Length = 326
Score = 33.9 bits (76), Expect = 0.84
Identities = 24/87 (27%), Positives = 37/87 (41%), Gaps = 4/87 (4%)
Query: 15 KVDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKE 74
K DK KK K + + + ED + K +K N KH D++ E + + K
Sbjct: 197 KEDKKTKKEKK----KAKKEEEKKQKEDKKNKKHSDKKNKKHDDDDEYSEAEPSVTEEKR 252
Query: 75 KGKSNSGKGEHITSRPLQAAHDEYEEE 101
S + K H TS L+ + E + E
Sbjct: 253 VETSFTKKPHHTTSAALKEGYKEVKNE 279
>ref|YP_067097.1| DNA-directed RNA polymerase beta prime subunit; RNA
nucleotidyltransferase (DNA-directed).; RNA polymerase
I.; RNA polymerase II.; RNA polymerase III. [Rickettsia
typhi str. Wilmington] gi|51459652|gb|AAU03615.1|
DNA-directed RNA polymerase beta prime subunit; RNA
nucleotidyltransferase (DNA-directed).; RNA polymerase
I.; RNA polymerase II.; RNA polymerase III. [Rickettsia
typhi str. Wilmington]
Length = 1372
Score = 33.9 bits (76), Expect = 0.84
Identities = 22/65 (33%), Positives = 32/65 (48%), Gaps = 3/65 (4%)
Query: 67 YMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATLAKQGCPHIAKLGKGRRIKPLKH 126
Y+++ P G S KGE +T LQ A D+Y E+A A G I ++ K LK
Sbjct: 143 YVVVDP---GLSILQKGELLTEEELQKAKDKYGEDAFTASIGAEVIQQMLKELDFSKLKQ 199
Query: 127 SNFNE 131
++E
Sbjct: 200 ELYDE 204
>gb|EAK86244.1| hypothetical protein UM04789.1 [Ustilago maydis 521]
gi|49076976|ref|XP_402404.1| hypothetical protein
UM04789.1 [Ustilago maydis 521]
Length = 426
Score = 33.9 bits (76), Expect = 0.84
Identities = 31/118 (26%), Positives = 47/118 (39%), Gaps = 14/118 (11%)
Query: 28 PIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKREVYEYMIIQPKEKGKSNSGKGEHIT 87
P + + N+ E + KEK K + E+ E + P + + EHI
Sbjct: 132 PTKKEKKEKNEKKEKKEKKEKKEKKEKKEQKTQSEEIAEQVPESPSDTEIDSDDAEEHIQ 191
Query: 88 SRPLQAAHDE---------YEEEATLAKQ---GCPHIAKLGKGRRIKPLKH--SNFNE 131
P+QAA E E EA KQ G +I+++ G ++H SNF E
Sbjct: 192 DDPMQAADQEEKIIKPLSTTELEAFKKKQRKLGIVYISRIPPGMTPAKVRHILSNFGE 249
>emb|CAI04661.1| conserved hypothetical protein [Plasmodium berghei]
Length = 469
Score = 33.9 bits (76), Expect = 0.84
Identities = 31/126 (24%), Positives = 54/126 (42%), Gaps = 9/126 (7%)
Query: 4 SKLHQIAALTSKVDKNIKKNGKFGPIRVPRGVNNQIFEDSNSLKVKEKDNMKHHCDEKRE 63
+K + ++ + K K I KN K+ + +P G ++I +L +K D K + E +
Sbjct: 118 NKNNILSCIDEKKKKKIDKNSKYKLVVLPNGKKHKI-----NLNIKLSDFQKLYISEDNK 172
Query: 64 VYEYMIIQPKEKGKSNSGKGEH-ITSRPLQAAHDEYEEEATLAKQGCPHIAKLGKGRRIK 122
+EY++ K K N K + I R +Y EE T C + +
Sbjct: 173 SFEYLLRNMKNK---NIEKNMYSIIKRNEYNMKMDYIEECTKNGIKCDMLHMNKSENELY 229
Query: 123 PLKHSN 128
P++ SN
Sbjct: 230 PMRISN 235
>ref|NP_359819.1| DNA-directed RNA polymerase beta prime chain [EC:2.7.7.6]
[Rickettsia conorii str. Malish 7]
gi|15619230|gb|AAL02720.1| DNA-directed RNA polymerase
beta prime chain [EC:2.7.7.6] [Rickettsia conorii str.
Malish 7] gi|25288418|pir||F97722 hypothetical protein
rpoC [imported] - Rickettsia conorii (strain Malish 7)
gi|14916697|sp|Q9RH40|RPOC_RICCN DNA-directed RNA
polymerase beta' chain (RNAP beta' subunit)
(Transcriptase beta' chain) (RNA polymerase beta'
subunit)
Length = 1372
Score = 33.5 bits (75), Expect = 1.1
Identities = 22/65 (33%), Positives = 31/65 (46%), Gaps = 3/65 (4%)
Query: 67 YMIIQPKEKGKSNSGKGEHITSRPLQAAHDEYEEEATLAKQGCPHIAKLGKGRRIKPLKH 126
Y+++ P G S KGE +T LQ A D+Y E+A A G I ++ K LK
Sbjct: 143 YVVVDP---GLSILQKGELLTEEELQKAKDKYGEDAFTASIGAEVIQQMLKELDFSKLKQ 199
Query: 127 SNFNE 131
+ E
Sbjct: 200 ELYEE 204
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 256,384,344
Number of Sequences: 2540612
Number of extensions: 10755808
Number of successful extensions: 20719
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 63
Number of HSP's that attempted gapping in prelim test: 20661
Number of HSP's gapped (non-prelim): 108
length of query: 145
length of database: 863,360,394
effective HSP length: 121
effective length of query: 24
effective length of database: 555,946,342
effective search space: 13342712208
effective search space used: 13342712208
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146817.2


