

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.13 + phase: 0 /pseudo
(36 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
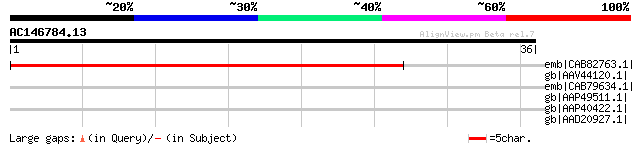
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB82763.1| (1-4)-beta-mannan endohydrolase-like protein [Ar... 46 3e-04
gb|AAV44120.1| hypothetical protein [Oryza sativa (japonica cult... 33 1.6
emb|CAB79634.1| putative (1-4)-beta-mannan endohydrolase [Arabid... 32 3.7
gb|AAP49511.1| At5g66460 [Arabidopsis thaliana] gi|23306404|gb|A... 32 4.8
gb|AAP40422.1| putative glycosyl hydrolase family 5 protein/cell... 32 4.8
gb|AAD20927.1| (1-4)-beta-mannan endohydrolase [Arabidopsis thal... 32 4.8
>emb|CAB82763.1| (1-4)-beta-mannan endohydrolase-like protein [Arabidopsis thaliana]
gi|29028774|gb|AAO64766.1| At5g01930 [Arabidopsis
thaliana] gi|15241093|ref|NP_195813.1| (1-4)-beta-mannan
endohydrolase, putative [Arabidopsis thaliana]
gi|11282585|pir||T48214 endo-1,4-beta-mannosidase-like
protein - Arabidopsis thaliana
Length = 448
Score = 45.8 bits (107), Expect = 3e-04
Identities = 19/27 (70%), Positives = 23/27 (84%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSKD 27
M AH+ED E YLGMPV+F+EFGVS+ D
Sbjct: 320 MQAHVEDAEMYLGMPVLFTEFGVSAHD 346
>gb|AAV44120.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|55168214|gb|AAV44080.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 520
Score = 33.5 bits (75), Expect = 1.6
Identities = 16/27 (59%), Positives = 22/27 (81%), Gaps = 1/27 (3%)
Query: 1 MDAHIEDTEKYLG-MPVVFSEFGVSSK 26
++AHI D E LG MPV+F+EFGVS++
Sbjct: 381 VEAHIADAEGALGGMPVLFAEFGVSTR 407
>emb|CAB79634.1| putative (1-4)-beta-mannan endohydrolase [Arabidopsis thaliana]
gi|15235255|ref|NP_194561.1| glycosyl hydrolase family 5
protein / cellulase family protein [Arabidopsis
thaliana] gi|7488130|pir||T09048 probable mannan
endo-1,4-beta-mannosidase (EC 3.2.1.78) - Arabidopsis
thaliana
Length = 431
Score = 32.3 bits (72), Expect = 3.7
Identities = 14/27 (51%), Positives = 21/27 (76%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSKD 27
M +HIED K L PV+F+EFG+S+++
Sbjct: 315 MQSHIEDGLKELKKPVLFTEFGLSNQN 341
>gb|AAP49511.1| At5g66460 [Arabidopsis thaliana] gi|23306404|gb|AAN17429.1| mannan
endo-1,4-beta-mannosidase [Arabidopsis thaliana]
gi|10177527|dbj|BAB10922.1| mannan
endo-1,4-beta-mannosidase [Arabidopsis thaliana]
gi|15239973|ref|NP_201447.1| (1-4)-beta-mannan
endohydrolase, putative [Arabidopsis thaliana]
Length = 431
Score = 32.0 bits (71), Expect = 4.8
Identities = 12/26 (46%), Positives = 18/26 (69%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSSK 26
+DAHI+D + L P++ +EFG S K
Sbjct: 301 LDAHIQDAQNVLHKPIILAEFGKSMK 326
>gb|AAP40422.1| putative glycosyl hydrolase family 5 protein/cellulase
((1-4)-beta-mannan endohydrolase) [Arabidopsis thaliana]
gi|30681096|ref|NP_179660.2| glycosyl hydrolase family 5
protein / cellulase family protein [Arabidopsis
thaliana]
Length = 433
Score = 32.0 bits (71), Expect = 4.8
Identities = 14/25 (56%), Positives = 20/25 (80%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSS 25
M +HIED +K L PV+F+EFG+S+
Sbjct: 316 MLSHIEDGDKELKKPVLFTEFGLSN 340
>gb|AAD20927.1| (1-4)-beta-mannan endohydrolase [Arabidopsis thaliana]
gi|25354515|pir||A84592 (1-4)-beta-mannan endohydrolase
[imported] - Arabidopsis thaliana
Length = 403
Score = 32.0 bits (71), Expect = 4.8
Identities = 14/25 (56%), Positives = 20/25 (80%)
Query: 1 MDAHIEDTEKYLGMPVVFSEFGVSS 25
M +HIED +K L PV+F+EFG+S+
Sbjct: 286 MLSHIEDGDKELKKPVLFTEFGLSN 310
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.130 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,972,932
Number of Sequences: 2540612
Number of extensions: 1069949
Number of successful extensions: 1273
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 1268
Number of HSP's gapped (non-prelim): 6
length of query: 36
length of database: 863,360,394
effective HSP length: 12
effective length of query: 24
effective length of database: 832,873,050
effective search space: 19988953200
effective search space used: 19988953200
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146784.13


