

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.12 - phase: 0 /pseudo
(60 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
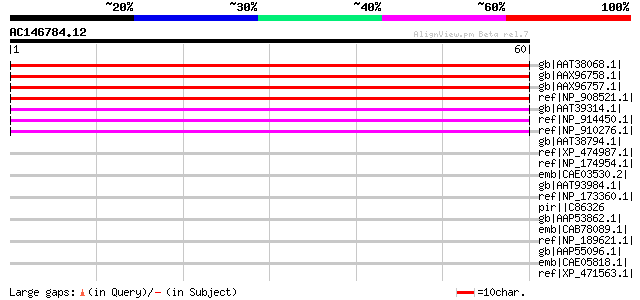
Score E
Sequences producing significant alignments: (bits) Value
gb|AAT38068.1| hypothetical protein [Oryza sativa (japonica cult... 62 3e-09
gb|AAX96758.1| hAT family dimerisation domain, putative [Oryza s... 62 3e-09
gb|AAX96757.1| hAT family dimerisation domain, putative [Oryza s... 62 3e-09
ref|NP_908521.1| putative transposable element Tip100 protein [O... 61 7e-09
gb|AAT39314.1| putative transposase [Solanum demissum] 45 4e-04
ref|NP_914450.1| putative transposable element [Oryza sativa (ja... 45 5e-04
ref|NP_910276.1| Similar to Arabidopsis thaliana DNA chromosome ... 44 9e-04
gb|AAT38794.1| putative hAT family dimerisation domain containin... 44 0.001
ref|XP_474987.1| OSJNBa0065B15.7 [Oryza sativa (japonica cultiva... 42 0.003
ref|NP_174954.1| hypothetical protein [Arabidopsis thaliana] 42 0.004
emb|CAE03530.2| OSJNBa0061C06.15 [Oryza sativa (japonica cultiva... 42 0.006
gb|AAT93984.1| hypothetical protein [Oryza sativa (japonica cult... 42 0.006
ref|NP_173360.1| hAT dimerisation domain-containing protein [Ara... 41 0.007
pir||C86326 hypothetical protein T29M8.13 [imported] - Arabidops... 41 0.007
gb|AAP53862.1| putative transposase [Oryza sativa (japonica cult... 41 0.010
emb|CAB78089.1| putative protein [Arabidopsis thaliana] gi|45388... 40 0.021
ref|NP_189621.1| hAT dimerisation domain-containing protein [Ara... 40 0.021
gb|AAP55096.1| putative transposase [Oryza sativa (japonica cult... 39 0.028
emb|CAE05818.1| OSJNBa0028M15.10 [Oryza sativa (japonica cultiva... 39 0.036
ref|XP_471563.1| OSJNBa0019J05.24 [Oryza sativa (japonica cultiv... 36 0.24
>gb|AAT38068.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 717
Score = 62.4 bits (150), Expect = 3e-09
Identities = 30/60 (50%), Positives = 40/60 (66%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
M++RF + +L CF+C D +SFS+FDVD L LADIY DFS DR +REQL ++
Sbjct: 551 MNNRFSKTFMDLLRCFACFDPDDSFSQFDVDMLLSLADIYSADFSMTDREILREQLHMFI 610
>gb|AAX96758.1| hAT family dimerisation domain, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1050
Score = 62.4 bits (150), Expect = 3e-09
Identities = 28/60 (46%), Positives = 42/60 (69%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+++RF ERS +L C +CLD +NSF+ FD DKL LA +Y DFS D +++QL+T++
Sbjct: 855 LNNRFAERSTQLLRCIACLDPRNSFANFDEDKLIELARMYAADFSPYDCIVLKDQLETFI 914
>gb|AAX96757.1| hAT family dimerisation domain, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1071
Score = 62.4 bits (150), Expect = 3e-09
Identities = 28/60 (46%), Positives = 42/60 (69%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+++RF ERS +L C +CLD +NSF+ FD DKL LA +Y DFS D +++QL+T++
Sbjct: 679 LNNRFAERSTQLLRCIACLDPRNSFANFDEDKLIELARMYAADFSPYDCIVLKDQLETFI 738
>ref|NP_908521.1| putative transposable element Tip100 protein [Oryza sativa
(japonica cultivar-group)]
Length = 802
Score = 61.2 bits (147), Expect = 7e-09
Identities = 31/60 (51%), Positives = 40/60 (66%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
M+HRF E S+ +L C S L+ +NSFS FDVDKL RLA+IY DF D +R QL ++
Sbjct: 619 MNHRFSEVSSELLVCMSSLNPRNSFSNFDVDKLVRLAEIYAEDFLVGDLMLLRTQLGNFI 678
>gb|AAT39314.1| putative transposase [Solanum demissum]
Length = 705
Score = 45.4 bits (106), Expect = 4e-04
Identities = 20/60 (33%), Positives = 36/60 (59%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
++ RF E + +L +CL+ SFS FD+ K+ R+A++Y DF + + + QL +Y+
Sbjct: 536 LNDRFDEVTTDLLHGIACLNPIKSFSSFDIRKIMRMAELYPDDFDESNMNILENQLASYI 595
>ref|NP_914450.1| putative transposable element [Oryza sativa (japonica
cultivar-group)]
Length = 796
Score = 45.1 bits (105), Expect = 5e-04
Identities = 20/60 (33%), Positives = 35/60 (58%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+D+RF E+++ +L C + L + SF F+++ L LA +Y DF D + +R L Y+
Sbjct: 625 LDNRFNEKTSQLLICAAALSPRQSFHDFNLEHLMSLAKLYPKDFDDGELMDLRHHLSLYI 684
>ref|NP_910276.1| Similar to Arabidopsis thaliana DNA chromosome 4, BAC clone F17A8;
putative protein. (AL049482) [Oryza sativa (japonica
cultivar-group)]
Length = 757
Score = 44.3 bits (103), Expect = 9e-04
Identities = 22/60 (36%), Positives = 33/60 (54%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+++RF E S +L C S + +SF+ FD K+ RLA Y D + + QLDTY+
Sbjct: 607 LENRFDEVSMELLLCMSAFNPTDSFASFDAQKILRLASFYPKDIEGSNLMKLELQLDTYI 666
>gb|AAT38794.1| putative hAT family dimerisation domain containing protein [Solanum
demissum]
Length = 805
Score = 43.9 bits (102), Expect = 0.001
Identities = 20/60 (33%), Positives = 35/60 (58%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
++ RF E + +L +CL+ SFS FD+ K R+A++Y DF + + + QL +Y+
Sbjct: 636 LNDRFDEVTTDLLHGIACLNPIKSFSSFDIRKKMRMAELYPDDFDESNMNILENQLASYI 695
>ref|XP_474987.1| OSJNBa0065B15.7 [Oryza sativa (japonica cultivar-group)]
gi|38344561|emb|CAD39903.2| OSJNBa0065B15.7 [Oryza
sativa (japonica cultivar-group)]
Length = 639
Score = 42.4 bits (98), Expect = 0.003
Identities = 20/60 (33%), Positives = 34/60 (56%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+D+RF E S+ +L C S ++SF F+++ L LA +Y DF+ + + QL Y+
Sbjct: 478 LDNRFSETSSQLLICSSAFSPRDSFHDFNLENLMSLAKLYPSDFNSGNLRDLSHQLGLYI 537
>ref|NP_174954.1| hypothetical protein [Arabidopsis thaliana]
Length = 496
Score = 42.0 bits (97), Expect = 0.004
Identities = 21/57 (36%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E + +L C + L +SF +FD K+ RL++ Y DF+ DR ++ QL Y+
Sbjct: 429 RFDEVNTELLSCVASLSPIDSFHEFDQLKVLRLSEFYPQDFTHVDRRSLEHQLGLYI 485
>emb|CAE03530.2| OSJNBa0061C06.15 [Oryza sativa (japonica cultivar-group)]
Length = 587
Score = 41.6 bits (96), Expect = 0.006
Identities = 20/60 (33%), Positives = 33/60 (54%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+++RF E + +L C + SFS F+VD L +LA Y DF ++ + QL+ Y+
Sbjct: 516 LNNRFDEVNTELLRCMASFSPAKSFSAFNVDNLVKLAKFYPSDFDVEEMNQLPFQLNRYI 575
>gb|AAT93984.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 774
Score = 41.6 bits (96), Expect = 0.006
Identities = 20/60 (33%), Positives = 33/60 (54%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+++RF E + +L C + SFS F+VD L +LA Y DF ++ + QL+ Y+
Sbjct: 650 LNNRFDEVNTELLRCMASFSPAKSFSAFNVDNLVKLAKFYPSDFDVEEMNQLPFQLNRYI 709
>ref|NP_173360.1| hAT dimerisation domain-containing protein [Arabidopsis thaliana]
Length = 769
Score = 41.2 bits (95), Expect = 0.007
Identities = 22/57 (38%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E ++ +L C S L +SF +FD L RL + Y DFS +R ++ QL+ Y+
Sbjct: 604 RFDEVNSELLICMSSLSPIDSFRQFDKSMLVRLTEFYPDDFSFVERRSLDHQLEIYL 660
>pir||C86326 hypothetical protein T29M8.13 [imported] - Arabidopsis thaliana
gi|8954063|gb|AAF82236.1| Contains similarity to a
transposable element Tip100 protein for transposase from
Ipomoea purpurea gb|4063769 and is a member of the
transmembrane 4 family PF|00335. [Arabidopsis thaliana]
Length = 811
Score = 41.2 bits (95), Expect = 0.007
Identities = 22/57 (38%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E ++ +L C S L +SF +FD L RL + Y DFS +R ++ QL+ Y+
Sbjct: 646 RFDEVNSELLICMSSLSPIDSFRQFDKSMLVRLTEFYPDDFSFVERRSLDHQLEIYL 702
>gb|AAP53862.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|37534546|ref|NP_921575.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
Length = 413
Score = 40.8 bits (94), Expect = 0.010
Identities = 20/60 (33%), Positives = 33/60 (54%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
++++F E + +L C S SFS F+VD L +LA Y DF ++ + QL+ Y+
Sbjct: 131 LNNKFDEVNPELLRCMSSFSPAKSFSDFNVDNLVKLAKFYPNDFDVEEMNQLTFQLNRYI 190
>emb|CAB78089.1| putative protein [Arabidopsis thaliana] gi|4538896|emb|CAB39633.1|
putative protein [Arabidopsis thaliana]
gi|15233987|ref|NP_192704.1| hypothetical protein
[Arabidopsis thaliana] gi|7485592|pir||T04013
hypothetical protein F17A8.10 - Arabidopsis thaliana
Length = 664
Score = 39.7 bits (91), Expect = 0.021
Identities = 21/57 (36%), Positives = 33/57 (57%)
Query: 4 RFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
RF E ++ +L C S L +SF +FD L RL + Y +FS +R ++ QL+ Y+
Sbjct: 548 RFDEVNSELLICMSSLSPIDSFCQFDKSMLVRLTEFYPDEFSFVERRSLDHQLEIYL 604
>ref|NP_189621.1| hAT dimerisation domain-containing protein [Arabidopsis thaliana]
Length = 536
Score = 39.7 bits (91), Expect = 0.021
Identities = 21/56 (37%), Positives = 32/56 (56%)
Query: 5 FRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
F E ++ +L C S L +SF +FD L RL + Y DFS +R ++ QL+ Y+
Sbjct: 372 FDELNSELLICMSSLSPIDSFRQFDKSMLVRLTEFYPDDFSFVERRSLDHQLEIYL 427
>gb|AAP55096.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|37537014|ref|NP_922809.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|19225003|gb|AAL86479.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
Length = 811
Score = 39.3 bits (90), Expect = 0.028
Identities = 20/60 (33%), Positives = 34/60 (56%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
++ RF E + +L C S L+ NSF+ +D ++ +LA Y +FS D + QL T++
Sbjct: 642 LNSRFDEVNMELLICMSALNPFNSFASYDAQQVLKLAKFYPKEFSTMDLIRLEMQLGTFI 701
>emb|CAE05818.1| OSJNBa0028M15.10 [Oryza sativa (japonica cultivar-group)]
gi|50930943|ref|XP_474999.1| OSJNBa0028M15.10 [Oryza
sativa (japonica cultivar-group)]
Length = 437
Score = 38.9 bits (89), Expect = 0.036
Identities = 19/60 (31%), Positives = 32/60 (52%)
Query: 1 MDHRFRERSNIVLDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
+D+ F E S +L C S ++SF F+++ L LA +Y DF+ + + QL Y+
Sbjct: 259 LDNHFSETSFQLLICSSAFSPRDSFHDFNLENLMSLAKLYPSDFNSGNLRDLSHQLGLYI 318
>ref|XP_471563.1| OSJNBa0019J05.24 [Oryza sativa (japonica cultivar-group)]
gi|32487628|emb|CAE04074.1| OSJNBb0032D24.4 [Oryza
sativa (japonica cultivar-group)]
gi|38346439|emb|CAD40226.2| OSJNBa0019J05.24 [Oryza
sativa (japonica cultivar-group)]
Length = 629
Score = 36.2 bits (82), Expect = 0.24
Identities = 17/48 (35%), Positives = 25/48 (51%)
Query: 13 LDCFSCLDSKNSFSKFDVDKLARLADIYHVDFSDDDRGTIREQLDTYV 60
L C + SFS F+VD L +LA Y DF ++ + QL+ Y+
Sbjct: 552 LRCMASFSPAKSFSAFNVDNLVKLAKFYPSDFDVEEMNQLPFQLNRYI 599
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.141 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 92,703,583
Number of Sequences: 2540612
Number of extensions: 2735984
Number of successful extensions: 7771
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 7745
Number of HSP's gapped (non-prelim): 26
length of query: 60
length of database: 863,360,394
effective HSP length: 36
effective length of query: 24
effective length of database: 771,898,362
effective search space: 18525560688
effective search space used: 18525560688
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146784.12


