

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146784.10 + phase: 0
(168 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
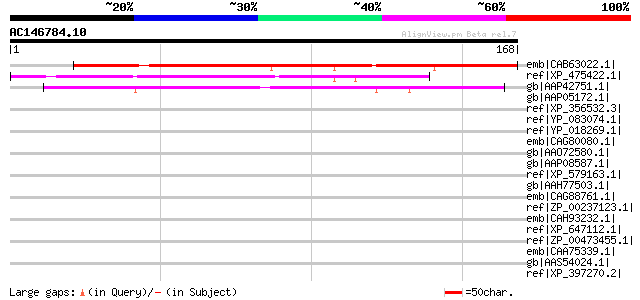
Sequences producing significant alignments: (bits) Value
emb|CAB63022.1| putative protein [Arabidopsis thaliana] gi|15230... 141 8e-33
ref|XP_475422.1| unknown protein [Oryza sativa (japonica cultiva... 88 8e-17
gb|AAP42751.1| At1g64385 [Arabidopsis thaliana] gi|27311781|gb|A... 65 6e-10
gb|AAP05172.1| conserved hypothetical protein [Chlamydophila cav... 39 0.042
ref|XP_356532.3| PREDICTED: similar to ribosomal protein S17 [Mu... 36 0.36
ref|YP_083074.1| hypothetical protein BCZK1479 [Bacillus cereus ... 36 0.47
ref|YP_018269.1| hypothetical protein GBAA1634 [Bacillus anthrac... 35 0.80
emb|CAG80080.1| unnamed protein product [Yarrowia lipolytica CLI... 35 0.80
gb|AAO72580.1| ADP ribosylation GTPase-like protein [Oryza sativ... 35 1.0
gb|AAP08587.1| hypothetical Membrane Spanning Protein [Bacillus ... 34 1.4
ref|XP_579163.1| PREDICTED: cell adhesion molecule JCAM [Rattus ... 34 1.4
gb|AAH77503.1| Snag1-prov protein [Xenopus laevis] 34 1.4
emb|CAG88761.1| unnamed protein product [Debaryomyces hansenii C... 33 3.0
ref|ZP_00237123.1| hypothetical protein membrane Spanning protei... 33 3.0
emb|CAH93232.1| hypothetical protein [Pongo pygmaeus] 33 3.0
ref|XP_647112.1| N-terminal kinase-like protein [Dictyostelium d... 33 3.0
ref|ZP_00473455.1| putative type-4 fimbrial pilin related signal... 33 4.0
emb|CAA75339.1| 70 kD tumor-specific antigen [Rattus norvegicus]... 32 5.2
gb|AAS54024.1| AFR652Wp [Ashbya gossypii ATCC 10895] gi|45199171... 32 5.2
ref|XP_397270.2| PREDICTED: similar to CG6757-PA, partial [Apis ... 32 5.2
>emb|CAB63022.1| putative protein [Arabidopsis thaliana]
gi|15230528|ref|NP_190726.1| expressed protein
[Arabidopsis thaliana] gi|11285599|pir||T45789
hypothetical protein F26O13.220 - Arabidopsis thaliana
Length = 390
Score = 141 bits (355), Expect = 8e-33
Identities = 70/153 (45%), Positives = 101/153 (65%), Gaps = 10/153 (6%)
Query: 22 IKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSFDKILTPVNGAYFLIFTVIVFA 81
I ++ ++ LD GKG C LH+ P E++ PS++K++TP+NGAYFLI +VI+F
Sbjct: 242 ISGDTNKIILDTGKGQCALHMY---PSEESTLPFHFPSYEKLVTPINGAYFLIVSVIIFG 298
Query: 82 VTWA-CCCIFKKKPRDEIPYQELEMA----LPESASATVVESAEGWDQGWDDDWDDNVAV 136
WA C C ++ +PY+ELE++ L + VE+A+ WD+GWDDDWD+N AV
Sbjct: 299 GIWAFCLCRKNRRAGSGVPYRELELSGGPGLENESGVHDVETAD-WDEGWDDDWDENNAV 357
Query: 137 KSP-VVRHAGSISANGLTSRSSNKDGWEDNWDD 168
KSP + SISANGLT+R+ N+DGW+ +WDD
Sbjct: 358 KSPGSAAKSVSISANGLTARAPNRDGWDHDWDD 390
>ref|XP_475422.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|46576005|gb|AAT01366.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 515
Score = 88.2 bits (217), Expect = 8e-17
Identities = 53/142 (37%), Positives = 71/142 (49%), Gaps = 8/142 (5%)
Query: 1 MSDLQLAYCFYNIVFQVTIKQIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSF 60
M LQLA F QV I E+T+ +GKG C LH T T F + ++
Sbjct: 307 MLPLQLAKGFSR---QVNITYSNPNGVEITVKSGKGQCSLH-TKQTVFDWQQQFQQFAAY 362
Query: 61 DKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMA--LPESASA-TVVE 117
P+ GA FL+FTV++ V ACC F ++ +PYQ+LEM P S+
Sbjct: 363 ATRANPIYGASFLVFTVVLVGVVCACCK-FARRRASGVPYQQLEMGDQAPNSSGVENTTS 421
Query: 118 SAEGWDQGWDDDWDDNVAVKSP 139
+ +GW+ GWDDDWDD A P
Sbjct: 422 TVDGWEDGWDDDWDDEEAAAKP 443
>gb|AAP42751.1| At1g64385 [Arabidopsis thaliana] gi|27311781|gb|AAO00856.1| Unknown
protein [Arabidopsis thaliana]
gi|22330420|ref|NP_683468.1| expressed protein
[Arabidopsis thaliana]
Length = 351
Score = 65.5 bits (158), Expect = 6e-10
Identities = 50/173 (28%), Positives = 77/173 (43%), Gaps = 23/173 (13%)
Query: 12 NIVFQVTIKQIKSESTELTLDAGKGDCVL---------HVTVVTPVPEASFFLRLPSFDK 62
+I +V+IK+ S + + L + KG C L H T S L +
Sbjct: 182 DIKVKVSIKKGGSNDSAIVLASSKGRCRLELKDLAAAAHETESDDTVSVSRPSILNISSR 241
Query: 63 ILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATVVESAE-- 120
L + FL+ ++++ V ++K K R YQ L+M LP S A V +S +
Sbjct: 242 TLIVIIMISFLVLSLVIIPVI---IHVYKNKSRGNNKYQRLDMELPVSNPALVTKSDQES 298
Query: 121 ---GWDQGWDDDWD------DNVAVKSPVVRHAGSISANGLTSRSSNKDGWED 164
GW+ W DDWD D +PV+ S+S+ GL R +K+GW+D
Sbjct: 299 GDDGWNNNWGDDWDDENGGGDEEQPNTPVLPLTPSLSSRGLAPRRLSKEGWKD 351
>gb|AAP05172.1| conserved hypothetical protein [Chlamydophila caviae GPIC]
gi|29840188|ref|NP_829294.1| hypothetical protein
CCA00426 [Chlamydophila caviae GPIC]
Length = 379
Score = 39.3 bits (90), Expect = 0.042
Identities = 19/46 (41%), Positives = 26/46 (56%), Gaps = 1/46 (2%)
Query: 74 IFTVIVFAVTWACCCIFKKKP-RDEIPYQELEMALPESASATVVES 118
+ TVI + ACC + KKP RD PY LE +PE+ T+ E+
Sbjct: 65 VATVIFSIIAVACCVLLNKKPQRDATPYPPLEEGMPENPRLTIPET 110
>ref|XP_356532.3| PREDICTED: similar to ribosomal protein S17 [Mus musculus]
Length = 528
Score = 36.2 bits (82), Expect = 0.36
Identities = 29/93 (31%), Positives = 41/93 (43%), Gaps = 10/93 (10%)
Query: 49 PEASFFLRLPSFDKILTP-VNG--AYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEM 105
PE S F R ++ + T NG A L+ TVI +CCC ++ + +P +
Sbjct: 233 PELSLFRRTYAWSTVYTSRTNGFPAPQLVATVITGFPNMSCCCCHRRCRHELLPLSCATL 292
Query: 106 ALPESASAT-------VVESAEGWDQGWDDDWD 131
PE AS+T V AEGW D+ D
Sbjct: 293 DTPEPASSTSPDPSPSSVPVAEGWKLSRVDETD 325
>ref|YP_083074.1| hypothetical protein BCZK1479 [Bacillus cereus E33L]
gi|51977222|gb|AAU18772.1| conserved hypothetical
protein [Bacillus cereus E33L]
Length = 113
Score = 35.8 bits (81), Expect = 0.47
Identities = 31/93 (33%), Positives = 45/93 (48%), Gaps = 13/93 (13%)
Query: 20 KQIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLP---SFDKIL-TPVNG--AYFL 73
K ++SE +L L+ K V + SFF+ LP S+ K+L TP G +
Sbjct: 19 KVVQSEEFQLLLNKKK-----KFIVPMSIFFLSFFIALPILTSYSKVLNTPAFGDVTWAW 73
Query: 74 IFTVIVFAVTWACCCIFKKKPR--DEIPYQELE 104
+F F +TWA C I+ KK DEI + L+
Sbjct: 74 VFAFAQFIMTWALCMIYSKKAESFDEISQKILQ 106
>ref|YP_018269.1| hypothetical protein GBAA1634 [Bacillus anthracis str. 'Ames
Ancestor'] gi|30261704|ref|NP_844081.1| hypothetical
protein BA1634 [Bacillus anthracis str. Ames]
gi|49477293|ref|YP_035821.1| hypothetical protein
BT9727_1489 [Bacillus thuringiensis serovar konkukian
str. 97-27] gi|49184532|ref|YP_027784.1| hypothetical
protein BAS1517 [Bacillus anthracis str. Sterne]
gi|30255932|gb|AAP25567.1| conserved hypothetical
protein [Bacillus anthracis str. Ames]
gi|49328849|gb|AAT59495.1| conserved hypothetical
protein [Bacillus thuringiensis serovar konkukian str.
97-27] gi|47502068|gb|AAT30744.1| conserved hypothetical
protein [Bacillus anthracis str. 'Ames Ancestor']
gi|65318973|ref|ZP_00391932.1| COG3162: Predicted
membrane protein [Bacillus anthracis str. A2012]
gi|49178459|gb|AAT53835.1| conserved hypothetical
protein [Bacillus anthracis str. Sterne]
Length = 113
Score = 35.0 bits (79), Expect = 0.80
Identities = 24/61 (39%), Positives = 33/61 (53%), Gaps = 8/61 (13%)
Query: 52 SFFLRLP---SFDKIL-TPVNG--AYFLIFTVIVFAVTWACCCIFKKKPR--DEIPYQEL 103
SFF+ LP S+ K+L TP G + +F F +TWA C I+ KK DEI + L
Sbjct: 46 SFFIALPILTSYSKVLNTPAFGDVTWAWVFAFAQFIMTWALCMIYSKKAESFDEISQKIL 105
Query: 104 E 104
+
Sbjct: 106 Q 106
>emb|CAG80080.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50554137|ref|XP_504477.1| hypothetical protein
[Yarrowia lipolytica]
Length = 749
Score = 35.0 bits (79), Expect = 0.80
Identities = 18/67 (26%), Positives = 31/67 (45%), Gaps = 3/67 (4%)
Query: 105 MALPESASATVVESAEGW--DQGWDDDWDDNVAVK-SPVVRHAGSISANGLTSRSSNKDG 161
+A + A+ ++ +GW D WDD D V +K +P + A ++DG
Sbjct: 682 LATKKPAAPPTLQDNDGWGLDDNWDDGSDGEVVIKEAPKTKPAPKTKKPAKLDLDDDEDG 741
Query: 162 WEDNWDD 168
+D WD+
Sbjct: 742 DDDGWDN 748
>gb|AAO72580.1| ADP ribosylation GTPase-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 308
Score = 34.7 bits (78), Expect = 1.0
Identities = 21/61 (34%), Positives = 27/61 (43%), Gaps = 3/61 (4%)
Query: 111 ASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRS---SNKDGWEDNWD 167
AS V E A+ GW DDW P R + NG S S S+K+ ++WD
Sbjct: 188 ASQKVEEYAKEGGNGWGDDWQRREQGSEPYHRFERETNGNGWNSSSHDGSSKNYNSNSWD 247
Query: 168 D 168
D
Sbjct: 248 D 248
>gb|AAP08587.1| hypothetical Membrane Spanning Protein [Bacillus cereus ATCC 14579]
gi|30019755|ref|NP_831386.1| hypothetical Membrane
Spanning Protein [Bacillus cereus ATCC 14579]
Length = 113
Score = 34.3 bits (77), Expect = 1.4
Identities = 24/61 (39%), Positives = 33/61 (53%), Gaps = 8/61 (13%)
Query: 52 SFFLRLP---SFDKIL-TPVNG--AYFLIFTVIVFAVTWACCCIFKKKPR--DEIPYQEL 103
SFF+ LP S+ K+L TP G + +F F +TWA C I+ KK DEI + L
Sbjct: 46 SFFIALPILTSYSKVLNTPAFGDVTWAWVFAFSQFIMTWALCMIYSKKAESFDEISRKIL 105
Query: 104 E 104
+
Sbjct: 106 Q 106
>ref|XP_579163.1| PREDICTED: cell adhesion molecule JCAM [Rattus norvegicus]
gi|41223398|gb|AAR99701.1| cell adhesion molecule JCAM
[Rattus norvegicus] gi|66912104|gb|AAH97947.1| Cell
adhesion molecule JCAM [Rattus norvegicus]
gi|55741625|ref|NP_001006991.1| cell adhesion molecule
JCAM [Rattus norvegicus]
Length = 369
Score = 34.3 bits (77), Expect = 1.4
Identities = 27/107 (25%), Positives = 48/107 (44%), Gaps = 10/107 (9%)
Query: 40 LHVTVVTPVPEA--SFFLRLPSFDKILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDE 97
+++TVV P P++ LP++ IL V A+ L+ +I+ + CCC ++ ++E
Sbjct: 216 VNLTVVQPPPDSIGGEGQALPTWAIILLAV--AFSLLLILIIALIIIFCCCCVSRREKEE 273
Query: 98 IPYQELEMALPESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHA 144
YQ E + V + + + W +N KS R A
Sbjct: 274 STYQN------EIRKSANVRTGKADPETWLKSGKENYGYKSDEARAA 314
>gb|AAH77503.1| Snag1-prov protein [Xenopus laevis]
Length = 591
Score = 34.3 bits (77), Expect = 1.4
Identities = 18/59 (30%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Query: 93 KPRDEIPYQELEMALPESASATVVESAEGWDQGWDDDWDD--NVAVKSPVVRHAGSISA 149
+P + L +A A + ++++ D WDD+WDD + A P + AGS SA
Sbjct: 91 QPFQSVAAPHLGLAFQPRAPSLGYQTSQPSDDDWDDEWDDSSSTAADEPAGQPAGSYSA 149
>emb|CAG88761.1| unnamed protein product [Debaryomyces hansenii CBS767]
gi|50423739|ref|XP_460454.1| unnamed protein product
[Debaryomyces hansenii]
Length = 413
Score = 33.1 bits (74), Expect = 3.0
Identities = 27/104 (25%), Positives = 46/104 (43%), Gaps = 8/104 (7%)
Query: 63 ILTPVNGAYFLIFTVIVFAVTWACCCIFKKKPRDEIPYQELEMALPESASATV--VESAE 120
I+ V G FL+ V++ W CC I+ K P+DE +L + + T+ + A
Sbjct: 11 IIGAVIGTCFLLSFVMILYCYW-CCVIYVKSPQDEYNVSDLHFRDKQKTTHTLRHMFKAM 69
Query: 121 GWDQGW--DDDWDDNVAVKSP---VVRHAGSISANGLTSRSSNK 159
W G D +D + + P V ++++N SR + K
Sbjct: 70 SWIYGGLRSKDLEDQIDIDKPPSSVTNCDTNLNSNSTFSRLALK 113
>ref|ZP_00237123.1| hypothetical protein membrane Spanning protein [Bacillus cereus
G9241] gi|47557002|gb|EAL15332.1| hypothetical protein
membrane Spanning protein [Bacillus cereus G9241]
gi|42780799|ref|NP_978046.1| hypothetical protein
BCE1725 [Bacillus cereus ATCC 10987]
gi|42736719|gb|AAS40654.1| conserved hypothetical
protein [Bacillus cereus ATCC 10987]
Length = 113
Score = 33.1 bits (74), Expect = 3.0
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 6/48 (12%)
Query: 52 SFFLRLP---SFDKIL-TPVNG--AYFLIFTVIVFAVTWACCCIFKKK 93
SFF+ LP S+ K+L TP G + IF F +TW+ C I+ KK
Sbjct: 46 SFFIALPILTSYSKVLNTPAFGDVTWAWIFAFAQFIMTWSLCMIYSKK 93
>emb|CAH93232.1| hypothetical protein [Pongo pygmaeus]
Length = 549
Score = 33.1 bits (74), Expect = 3.0
Identities = 23/71 (32%), Positives = 36/71 (50%), Gaps = 5/71 (7%)
Query: 93 KPRDEIPYQELEMALPESASATVV-ESAEGWDQGWDDDWDDNVAV----KSPVVRHAGSI 147
K +D+I +++L+ + E+ + E EG+D DDDWD + V K V +
Sbjct: 33 KEKDDILFEDLQDNVNENGEGEIEDEEEEGYDDDDDDDWDWDEGVGKLTKGYVWNGGSNP 92
Query: 148 SANGLTSRSSN 158
AN TS SS+
Sbjct: 93 QANRQTSDSSS 103
>ref|XP_647112.1| N-terminal kinase-like protein [Dictyostelium discoideum]
gi|60475288|gb|EAL73223.1| N-terminal kinase-like (NTKL)
protein [Dictyostelium discoideum]
Length = 813
Score = 33.1 bits (74), Expect = 3.0
Identities = 16/51 (31%), Positives = 24/51 (46%), Gaps = 3/51 (5%)
Query: 118 SAEGWDQGWDDDW-DDNVAVKSPVVRHAGSISANGLTSRSSNKDGWEDNWD 167
+++GW G DD W +D + V S + +S DGW+D WD
Sbjct: 765 NSDGW--GDDDGWGNDEIQVPSTTTSNIKKTVEKKTIVTASKSDGWDDGWD 813
>ref|ZP_00473455.1| putative type-4 fimbrial pilin related signal peptide protein
[Chromohalobacter salexigens DSM 3043]
gi|67519231|gb|EAM23187.1| putative type-4 fimbrial
pilin related signal peptide protein [Chromohalobacter
salexigens DSM 3043]
Length = 175
Score = 32.7 bits (73), Expect = 4.0
Identities = 19/62 (30%), Positives = 27/62 (42%), Gaps = 5/62 (8%)
Query: 100 YQELEMALPESASATVVESAEGWDQGW----DDDWDDNVAVKSPVVRHAGSISANGLTSR 155
Y A+ + +V GW+QGW D D D P +R G + ANGL R
Sbjct: 52 YYTRSEAIRQGVPVMLVSEENGWEQGWRVFADRDDDGRQGPSEPTLRAGGGL-ANGLRLR 110
Query: 156 SS 157
++
Sbjct: 111 AN 112
>emb|CAA75339.1| 70 kD tumor-specific antigen [Rattus norvegicus]
gi|3123027|sp|O35828|W70T_RAT 70 kDa WD-repeat tumor
rejection antigen
Length = 443
Score = 32.3 bits (72), Expect = 5.2
Identities = 21/55 (38%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Query: 6 LAYCFYNIVFQVTIKQIKSESTELTLDAGKGDCVLHVTVVTPVPEASFFLRLPSF 60
L +C+ +IV T+ T L L GKGD V + +PEA FFL SF
Sbjct: 240 LQHCWASIVAPSTLLPSYDPDTGLVLLTGKGD--TRVFLYEVIPEAPFFLECNSF 292
>gb|AAS54024.1| AFR652Wp [Ashbya gossypii ATCC 10895] gi|45199171|ref|NP_986200.1|
AFR652Wp [Eremothecium gossypii]
Length = 762
Score = 32.3 bits (72), Expect = 5.2
Identities = 20/61 (32%), Positives = 28/61 (45%), Gaps = 3/61 (4%)
Query: 109 ESASATVVESAEGWDQGWDDDWDDNVAVKSPVVRHAGSISANGLTSRSSNK-DGWEDNWD 167
+S S E E D+GW+ ++D + S R + S L N DGW+D WD
Sbjct: 428 KSMSTNEWEWNENDDEGWNAEFDTEI--NSEEHRLSDSKDKGVLVEEDQNNADGWDDVWD 485
Query: 168 D 168
D
Sbjct: 486 D 486
>ref|XP_397270.2| PREDICTED: similar to CG6757-PA, partial [Apis mellifera]
Length = 532
Score = 32.3 bits (72), Expect = 5.2
Identities = 17/69 (24%), Positives = 33/69 (47%), Gaps = 2/69 (2%)
Query: 102 ELEMALPESASATVVESAEGWDQGWDDDWDDNV--AVKSPVVRHAGSISANGLTSRSSNK 159
EL + E S +GW +G++ + + A ++ S+ ++S+ S+
Sbjct: 15 ELSITAGEVLSVMRDNVGDGWCEGFNQNGKSGLFPAAYVQIIETPTSVPNAMISSQQSSG 74
Query: 160 DGWEDNWDD 168
D W+D+WDD
Sbjct: 75 DYWDDDWDD 83
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.135 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 282,482,197
Number of Sequences: 2540612
Number of extensions: 11078945
Number of successful extensions: 26296
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 26255
Number of HSP's gapped (non-prelim): 47
length of query: 168
length of database: 863,360,394
effective HSP length: 118
effective length of query: 50
effective length of database: 563,568,178
effective search space: 28178408900
effective search space used: 28178408900
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146784.10


