

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146758.8 - phase: 0 /pseudo
(35 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
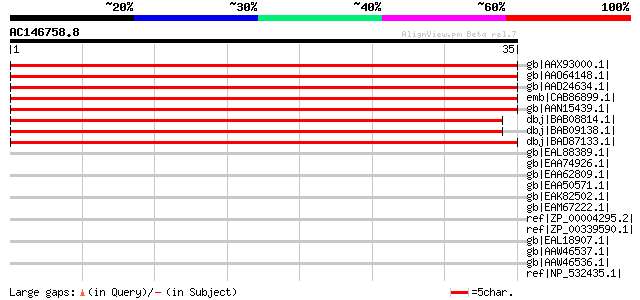
Score E
Sequences producing significant alignments: (bits) Value
gb|AAX93000.1| CBS domain, putative [Oryza sativa (japonica cult... 59 3e-08
gb|AAO64148.1| unknown protein [Arabidopsis thaliana] 56 2e-07
gb|AAD24634.1| hypothetical protein [Arabidopsis thaliana] gi|25... 56 2e-07
emb|CAB86899.1| putative protein [Arabidopsis thaliana] gi|42565... 55 7e-07
gb|AAN15439.1| putative protein [Arabidopsis thaliana] gi|221360... 55 7e-07
dbj|BAB08814.1| unnamed protein product [Arabidopsis thaliana] g... 52 3e-06
dbj|BAB09138.1| unnamed protein product [Arabidopsis thaliana] g... 51 8e-06
dbj|BAD87133.1| CBS domain-containing protein-like [Oryza sativa... 47 1e-04
gb|EAL88389.1| CBS and PB1 domain protein [Aspergillus fumigatus... 37 0.20
gb|EAA74926.1| hypothetical protein FG06309.1 [Gibberella zeae P... 36 0.25
gb|EAA62809.1| hypothetical protein AN5716.2 [Aspergillus nidula... 36 0.25
gb|EAA50571.1| hypothetical protein MG04330.4 [Magnaporthe grise... 36 0.25
gb|EAK82502.1| hypothetical protein UM01686.1 [Ustilago maydis 5... 35 0.57
gb|EAM67222.1| IMP dehydrogenase [Jannaschia sp. CCS1] 35 0.74
ref|ZP_00004295.2| COG0516: IMP dehydrogenase/GMP reductase [Rho... 34 1.3
ref|ZP_00339590.1| COG0516: IMP dehydrogenase/GMP reductase [Sil... 34 1.3
gb|EAL18907.1| hypothetical protein CNBI1680 [Cryptococcus neofo... 34 1.3
gb|AAW46537.1| conserved hypothetical protein [Cryptococcus neof... 34 1.3
gb|AAW46536.1| conserved hypothetical protein [Cryptococcus neof... 34 1.3
ref|NP_532435.1| hypothetical protein Atu1752 [Agrobacterium tum... 33 1.7
>gb|AAX93000.1| CBS domain, putative [Oryza sativa (japonica cultivar-group)]
Length = 575
Score = 59.3 bits (142), Expect = 3e-08
Identities = 30/35 (85%), Positives = 32/35 (90%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDAVLLTDANGLL GI+TDKD A RVI+EG
Sbjct: 77 MAARRVDAVLLTDANGLLSGIVTDKDIAKRVIAEG 111
>gb|AAO64148.1| unknown protein [Arabidopsis thaliana]
Length = 536
Score = 56.2 bits (134), Expect = 2e-07
Identities = 28/35 (80%), Positives = 32/35 (91%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDAVLLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 86 MAARRVDAVLLTDSSALLSGIVTDKDIATRVIAEG 120
>gb|AAD24634.1| hypothetical protein [Arabidopsis thaliana] gi|25408483|pir||E84781
hypothetical protein At2g36500 [imported] - Arabidopsis
thaliana gi|15227986|ref|NP_181191.1| CBS
domain-containing protein / octicosapeptide/Phox/Bemp1
(PB1) domain-containing protein [Arabidopsis thaliana]
Length = 536
Score = 56.2 bits (134), Expect = 2e-07
Identities = 28/35 (80%), Positives = 32/35 (91%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDAVLLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 86 MAARRVDAVLLTDSSALLSGIVTDKDIATRVIAEG 120
>emb|CAB86899.1| putative protein [Arabidopsis thaliana]
gi|42565877|ref|NP_190863.3| CBS domain-containing
protein / octicosapeptide/Phox/Bemp1 (PB1)
domain-containing protein [Arabidopsis thaliana]
gi|11357878|pir||T47552 hypothetical protein F8J2.120 -
Arabidopsis thaliana
Length = 556
Score = 54.7 bits (130), Expect = 7e-07
Identities = 27/35 (77%), Positives = 31/35 (88%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDA LLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 88 MAARRVDACLLTDSSALLSGIVTDKDVATRVIAEG 122
>gb|AAN15439.1| putative protein [Arabidopsis thaliana]
gi|22136010|gb|AAM91587.1| putative protein
[Arabidopsis thaliana]
Length = 469
Score = 54.7 bits (130), Expect = 7e-07
Identities = 27/35 (77%), Positives = 31/35 (88%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAARRVDA LLTD++ LL GI+TDKD ATRVI+EG
Sbjct: 1 MAARRVDACLLTDSSALLSGIVTDKDVATRVIAEG 35
>dbj|BAB08814.1| unnamed protein product [Arabidopsis thaliana]
gi|15242788|ref|NP_201154.1| CBS domain-containing
protein / octicosapeptide/Phox/Bemp1 (PB1)
domain-containing protein [Arabidopsis thaliana]
Length = 543
Score = 52.4 bits (124), Expect = 3e-06
Identities = 25/34 (73%), Positives = 31/34 (90%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISE 34
MA+RRVDA+LLTD+N +L GI+TDKD ATRVIS+
Sbjct: 79 MASRRVDALLLTDSNEMLCGILTDKDIATRVISQ 112
>dbj|BAB09138.1| unnamed protein product [Arabidopsis thaliana]
gi|8777387|dbj|BAA96977.1| unnamed protein product
[Arabidopsis thaliana] gi|22327688|ref|NP_680412.1| CBS
domain-containing protein / octicosapeptide/Phox/Bemp1
(PB1) domain-containing protein [Arabidopsis thaliana]
gi|18423173|ref|NP_568736.1| CBS domain-containing
protein / octicosapeptide/Phox/Bemp1 (PB1)
domain-containing protein [Arabidopsis thaliana]
Length = 548
Score = 51.2 bits (121), Expect = 8e-06
Identities = 24/34 (70%), Positives = 31/34 (90%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISE 34
MAARRVDA+LLTD+N LL GI+TD+D AT+VI++
Sbjct: 87 MAARRVDALLLTDSNALLCGILTDRDIATKVIAK 120
>dbj|BAD87133.1| CBS domain-containing protein-like [Oryza sativa (japonica
cultivar-group)] gi|57899170|dbj|BAD87222.1| CBS
domain-containing protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 533
Score = 47.0 bits (110), Expect = 1e-04
Identities = 22/35 (62%), Positives = 29/35 (82%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MA +RVDA LLTD+NG+L GI+T +D + RVI+EG
Sbjct: 78 MALKRVDAALLTDSNGMLSGILTAEDISGRVIAEG 112
>gb|EAL88389.1| CBS and PB1 domain protein [Aspergillus fumigatus Af293]
Length = 661
Score = 36.6 bits (83), Expect = 0.20
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 126 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGNG 160
>gb|EAA74926.1| hypothetical protein FG06309.1 [Gibberella zeae PH-1]
gi|46123863|ref|XP_386485.1| hypothetical protein
FG06309.1 [Gibberella zeae PH-1]
Length = 680
Score = 36.2 bits (82), Expect = 0.25
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 121 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGAG 155
>gb|EAA62809.1| hypothetical protein AN5716.2 [Aspergillus nidulans FGSC A4]
gi|67539092|ref|XP_663320.1| hypothetical protein
AN5716_2 [Aspergillus nidulans FGSC A4]
gi|49096786|ref|XP_409853.1| hypothetical protein
AN5716.2 [Aspergillus nidulans FGSC A4]
Length = 666
Score = 36.2 bits (82), Expect = 0.25
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 132 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGAG 166
>gb|EAA50571.1| hypothetical protein MG04330.4 [Magnaporthe grisea 70-15]
gi|39944638|ref|XP_361856.1| hypothetical protein
MG04330.4 [Magnaporthe grisea 70-15]
Length = 733
Score = 36.2 bits (82), Expect = 0.25
Identities = 18/35 (51%), Positives = 23/35 (65%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
MAA+R D VL+TD + + GI T KD A RV+ G
Sbjct: 161 MAAKREDCVLVTDDDDRIAGIFTAKDLAFRVVGAG 195
>gb|EAK82502.1| hypothetical protein UM01686.1 [Ustilago maydis 521]
gi|49070024|ref|XP_399301.1| hypothetical protein
UM01686.1 [Ustilago maydis 521]
Length = 708
Score = 35.0 bits (79), Expect = 0.57
Identities = 18/34 (52%), Positives = 22/34 (63%)
Query: 2 AARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
AA+R D VL+ D + L GI T KD A RV+S G
Sbjct: 87 AAKRTDCVLVVDEDEHLAGIFTAKDLAFRVVSAG 120
>gb|EAM67222.1| IMP dehydrogenase [Jannaschia sp. CCS1]
Length = 482
Score = 34.7 bits (78), Expect = 0.74
Identities = 15/33 (45%), Positives = 23/33 (69%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVIS 33
M ARR++ +L+TD G L G++T KDT V++
Sbjct: 172 MKARRIEKLLITDGQGALTGLLTLKDTEQAVLN 204
>ref|ZP_00004295.2| COG0516: IMP dehydrogenase/GMP reductase [Rhodobacter sphaeroides
2.4.1]
Length = 482
Score = 33.9 bits (76), Expect = 1.3
Identities = 15/33 (45%), Positives = 23/33 (69%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVIS 33
M ARR++ +L+TD G L G++T KDT V++
Sbjct: 172 MKARRIEKLLVTDGQGKLTGLLTLKDTEKSVLN 204
>ref|ZP_00339590.1| COG0516: IMP dehydrogenase/GMP reductase [Silicibacter sp. TM1040]
Length = 482
Score = 33.9 bits (76), Expect = 1.3
Identities = 15/33 (45%), Positives = 24/33 (72%)
Query: 1 MAARRVDAVLLTDANGLLFGIMTDKDTATRVIS 33
M ARR++ +L+TD +G L G++T KDT V++
Sbjct: 172 MKARRIEKLLVTDKDGKLTGLLTLKDTEQAVLN 204
>gb|EAL18907.1| hypothetical protein CNBI1680 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 831
Score = 33.9 bits (76), Expect = 1.3
Identities = 18/34 (52%), Positives = 21/34 (60%)
Query: 2 AARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
AA+R D VL+ D L GI T KD A RV +EG
Sbjct: 235 AAKRADCVLVVDEEEGLSGIFTAKDLAFRVTAEG 268
>gb|AAW46537.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58261288|ref|XP_568054.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 831
Score = 33.9 bits (76), Expect = 1.3
Identities = 18/34 (52%), Positives = 21/34 (60%)
Query: 2 AARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
AA+R D VL+ D L GI T KD A RV +EG
Sbjct: 235 AAKRADCVLVVDEEEGLSGIFTAKDLAFRVTAEG 268
>gb|AAW46536.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58261286|ref|XP_568053.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 704
Score = 33.9 bits (76), Expect = 1.3
Identities = 18/34 (52%), Positives = 21/34 (60%)
Query: 2 AARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
AA+R D VL+ D L GI T KD A RV +EG
Sbjct: 108 AAKRADCVLVVDEEEGLSGIFTAKDLAFRVTAEG 141
>ref|NP_532435.1| hypothetical protein Atu1752 [Agrobacterium tumefaciens str. C58]
gi|17740192|gb|AAL42751.1| conserved hypothetical
protein [Agrobacterium tumefaciens str. C58]
gi|15156853|gb|AAK87522.1| AGR_C_3216p [Agrobacterium
tumefaciens str. C58] gi|25523676|pir||AI2791 conserved
hypothetical protein Atu1752 [imported] - Agrobacterium
tumefaciens (strain C58, Dupont)
gi|25518976|pir||A97571 hypothetical protein AGR_C_3216
[imported] - Agrobacterium tumefaciens (strain C58,
Cereon) gi|15889056|ref|NP_354737.1| hypothetical
protein AGR_C_3216 [Agrobacterium tumefaciens str. C58]
Length = 144
Score = 33.5 bits (75), Expect = 1.7
Identities = 13/33 (39%), Positives = 23/33 (69%)
Query: 3 ARRVDAVLLTDANGLLFGIMTDKDTATRVISEG 35
A ++ AV++TDA+G++ GI T++D V +G
Sbjct: 34 AHKIGAVVVTDADGVVLGIFTERDLVKAVAGQG 66
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.136 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,394,699
Number of Sequences: 2540612
Number of extensions: 665094
Number of successful extensions: 1723
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 1679
Number of HSP's gapped (non-prelim): 46
length of query: 35
length of database: 863,360,394
effective HSP length: 11
effective length of query: 24
effective length of database: 835,413,662
effective search space: 20049927888
effective search space used: 20049927888
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146758.8


