

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146750.9 + phase: 0
(214 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
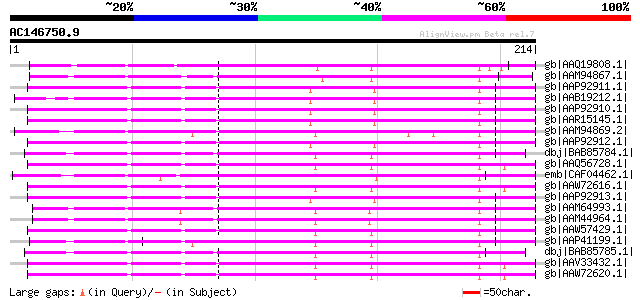
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ19808.1| polygalacturonase-inhibiting protein [Gossypium b... 135 6e-31
gb|AAM94867.1| polygalacturonase inhibitor protein [Brassica nap... 135 8e-31
gb|AAP92911.1| polygalacturonase-inhibiting protein [Pyrus pyrif... 135 8e-31
gb|AAB19212.1| polygalacturonase-inhibiting protein [Malus x dom... 135 8e-31
gb|AAP92910.1| polygalacturonase-inhibiting protein [Pyrus pyrif... 133 3e-30
gb|AAR15145.1| polygalacturonase-inhibiting protein [Eucalyptus ... 133 3e-30
gb|AAM94869.2| polygalacturonase inhibitor protein [Brassica nap... 133 4e-30
gb|AAP92912.1| polygalacturonase-inhibiting protein [Pyrus hybri... 132 9e-30
dbj|BAB85784.1| polygalacturonase-inhibiting protein [Citrus lat... 132 9e-30
gb|AAQ56728.1| polygalacturonase inhibiting protein [Prunus pers... 131 1e-29
emb|CAF04462.1| putative polygalacturonase-inhibiting protein [R... 130 2e-29
gb|AAW72616.1| polygalacturonase-inhibiting protein [Prunus pers... 130 2e-29
gb|AAP92913.1| polygalacturonase-inhibiting protein [Pyrus commu... 130 2e-29
gb|AAM64993.1| polygalacturonase inhibiting protein [Arabidopsis... 130 2e-29
gb|AAM44964.1| putative polygalacturonase inhibiting protein [Ar... 130 2e-29
gb|AAW57429.1| polygalacturonase-inhibiting protein [Prunus amer... 130 3e-29
gb|AAP41199.1| polygalacturonase-inhibiting protein [Cucumis melo] 129 4e-29
dbj|BAB85785.1| polygalacturonase-inhibiting protein [Citrus hys... 129 6e-29
gb|AAV33432.1| polygalacturonase inhibiting protein [Prunus mume] 128 9e-29
gb|AAW72620.1| polygalacturonase-inhibiting protein [Prunus mume... 128 9e-29
>gb|AAQ19808.1| polygalacturonase-inhibiting protein [Gossypium barbadense]
gi|33469564|gb|AAQ19807.1| polygalacturonase-inhibiting
protein [Gossypium barbadense]
Length = 330
Score = 135 bits (341), Expect = 6e-31
Identities = 82/200 (41%), Positives = 110/200 (55%), Gaps = 9/200 (4%)
Query: 9 LLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG 68
L FL F + SD CNA DK LLKI+ G P L WD T+CCDW + C
Sbjct: 8 LSFLFITIFISPSVSDH--CNAQDKKVLLKIKKALGNPY-LLASWDPKTDCCDWYCLEC- 63
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
P RV +T+ L+G +P E G+LPYL L +P + G I + +KL+ L+ L
Sbjct: 64 HPNTHRVVSLTLFSDDRLTGQIPPEVGDLPYLETLLFRHLPNLNGTIQPAIAKLKNLKML 123
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L +LSGP+P+FL +LK L +DLS N LSG+IP+SL L +L ++ N+L G IP
Sbjct: 124 RLSWTNLSGPVPNFLSQLKNLTYLDLSFNNLSGSIPSSLSTLPNLEDLHLDRNKLTGTIP 183
Query: 189 AGLKKF-NKNVF----EHNK 203
F KN++ HNK
Sbjct: 184 ESFGMFPRKNLYLFILSHNK 203
Score = 87.4 bits (215), Expect = 2e-16
Identities = 62/179 (34%), Positives = 84/179 (46%), Gaps = 52/179 (29%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR--LQNLDLGSNSLSGPIPSFL 143
LSG++P+ LP L L L + K+TG IP SF R L L N LSG IP+ L
Sbjct: 154 LSGSIPSSLSTLPNLEDLHL-DRNKLTGTIPESFGMFPRKNLYLFILSHNKLSGTIPASL 212
Query: 144 GKL----------------------------------------------KRLKEVDLSNN 157
+ K L +DL++N
Sbjct: 213 ANMDFNTIDLSRNLLEGDPSVLFGPNKTTFEIDLSRNMFQFGLSKVQFPKSLARLDLNHN 272
Query: 158 KLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
K++G+IPA L +L+ L NVS+N+LCG IP G L+ F+ + + HN+CLCGAPL CK
Sbjct: 273 KITGSIPAGLTDLE-LQFMNVSYNRLCGQIPVGGRLQSFDYSTYFHNRCLCGAPLDTCK 330
>gb|AAM94867.1| polygalacturonase inhibitor protein [Brassica napus]
gi|22256018|gb|AAM94868.1| polygalacturonase inhibitor
protein [Brassica napus]
Length = 342
Score = 135 bits (340), Expect = 8e-31
Identities = 79/191 (41%), Positives = 111/191 (57%), Gaps = 6/191 (3%)
Query: 9 LLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG 68
LL L ++ F+ S + D C+ DDK LLKI+ P + WD +CC W V CG
Sbjct: 8 LLLLFALLFTTSLSKD--LCHKDDKNTLLKIKKAMNNPY-TIISWDPKDDCCTWVSVECG 64
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
RVT + IS +S +P E G+LPYL L+L ++P +TG IP + +KL+ L++L
Sbjct: 65 NA--NRVTSLDISDD-DVSAQIPPEVGDLPYLQYLTLRKLPNLTGEIPPTIAKLKYLKSL 121
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L NSL+GP+P FL +LK L+ ++LS NKLSG+IP SL L L +S N+L G IP
Sbjct: 122 WLSWNSLTGPVPEFLSQLKNLEYINLSFNKLSGSIPGSLSLLPKLDFLELSRNKLTGPIP 181
Query: 189 AGLKKFNKNVF 199
F + +
Sbjct: 182 ESFGSFKRAAY 192
Score = 84.0 bits (206), Expect = 3e-15
Identities = 60/177 (33%), Positives = 79/177 (43%), Gaps = 51/177 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQ-NLDLGSNSLSGPIPSFLG 144
LSG++P LP L L L+ K+TGPIP SF +R + L N LSG IP LG
Sbjct: 152 LSGSIPGSLSLLPKLDFLELSRN-KLTGPIPESFGSFKRAAYGIYLSHNQLSGSIPKSLG 210
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
+ K + +DL++N
Sbjct: 211 NVDFNTIDLSRNKLEGDASMLFGAKKTTWHIDLSRNMFQFDISKVKVAKTVNFLDLNHNS 270
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPC 213
L+G+IP L L FNVS+N+LCG IP G L++F+ + HNKCLC APL C
Sbjct: 271 LTGSIPDQWTQLD-LQTFNVSYNRLCGRIPQGGDLQRFDVYAYLHNKCLCDAPLPSC 326
>gb|AAP92911.1| polygalacturonase-inhibiting protein [Pyrus pyrifolia]
Length = 330
Score = 135 bits (340), Expect = 8e-31
Identities = 79/191 (41%), Positives = 111/191 (57%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
T L L+ ++ S N + CN DDK LL+I+ FG P L W ++T+CCDW V C
Sbjct: 7 TFLSLTLLFSSVLNPALSDLCNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA G+LPYL L + P +TGPI + +KL+ L++
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKS 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L +LSG +P FL +LK L +DLS N L+G IP+SL L +LS ++ N+L G I
Sbjct: 124 LRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLSALHLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F NV
Sbjct: 184 PKSFGQFIGNV 194
Score = 77.8 bits (190), Expect = 2e-13
Identities = 55/178 (30%), Positives = 81/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP LS L L + K+TG IP SF + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSELPNLSALHL-DRNKLTGHIPKSFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL CK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSCK 330
>gb|AAB19212.1| polygalacturonase-inhibiting protein [Malus x domestica]
Length = 330
Score = 135 bits (340), Expect = 8e-31
Identities = 84/196 (42%), Positives = 114/196 (57%), Gaps = 8/196 (4%)
Query: 3 LLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDW 62
+ L LTLLF S + K SD CN DDK LL+I+ FG P L W ++T+CCDW
Sbjct: 7 IFLSLTLLFSSVL---KPALSD--LCNPDDKKVLLQIKKAFGDPYV-LTSWKSDTDCCDW 60
Query: 63 SFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL 122
V C R+ +TI G +SG +PA G+LPYL L + P +TGPI + +KL
Sbjct: 61 YCVTCDST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKL 118
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
+ L+ L L +LSG +P FL +LK L +DLS N L+G IP+SL L +L+ ++ N+
Sbjct: 119 KGLKFLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSQLPNLNALHLDRNK 178
Query: 183 LCGAIPAGLKKFNKNV 198
L G IP L +F NV
Sbjct: 179 LTGHIPKSLGQFIGNV 194
Score = 74.3 bits (181), Expect = 2e-12
Identities = 53/178 (29%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L+ L L + K+TG IP S + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSQLPNLNALHL-DRNKLTGHIPKSLGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL CK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSCK 330
>gb|AAP92910.1| polygalacturonase-inhibiting protein [Pyrus pyrifolia]
gi|464367|sp|Q05091|PGIP_PYRCO Polygalacturonase
inhibitor precursor (Polygalacturonase-inhibiting
protein) gi|169684|gb|AAA33865.1| polygalacturonase
inhibitor
Length = 330
Score = 133 bits (335), Expect = 3e-30
Identities = 78/191 (40%), Positives = 109/191 (56%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
T L L+ ++ S N + CN DDK LL+I+ FG P L W ++T+CCDW V C
Sbjct: 7 TFLSLTLLFSSVLNPALSDLCNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA G+LPYL L + P +TGPI + +KL+ L++
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKS 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L +LSG +P FL +LK L +DLS N L+G IP+SL L +L + N+L G I
Sbjct: 124 LRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F NV
Sbjct: 184 PISFGQFIGNV 194
Score = 75.1 bits (183), Expect = 1e-12
Identities = 54/178 (30%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L L L + K+TG IP SF + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSELPNLGALRL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL CK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSCK 330
>gb|AAR15145.1| polygalacturonase-inhibiting protein [Eucalyptus grandis]
Length = 331
Score = 133 bits (335), Expect = 3e-30
Identities = 78/191 (40%), Positives = 109/191 (56%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
T L L+ ++ S N + CN DDK LL+I+ FG P L W ++T+CCDW V C
Sbjct: 7 TFLSLTLLFSSVLNPALSDLCNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA G+LPYL L + P +TGPI + +KL+ L++
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKS 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L +LSG +P FL +LK L +DLS N L+G IP+SL L +L + N+L G I
Sbjct: 124 LRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F NV
Sbjct: 184 PISFGQFIGNV 194
Score = 78.6 bits (192), Expect = 1e-13
Identities = 55/178 (30%), Positives = 81/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L L L + K+TG IP SF + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSELPNLGALRL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL PCK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLGPCK 330
>gb|AAM94869.2| polygalacturonase inhibitor protein [Brassica napus]
gi|26094814|gb|AAM94870.2| polygalacturonase inhibitor
protein [Brassica napus]
Length = 331
Score = 133 bits (334), Expect = 4e-30
Identities = 81/197 (41%), Positives = 110/197 (55%), Gaps = 9/197 (4%)
Query: 3 LLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDW 62
LLL L + FSK+ CN DDK LLKI+ P L WD T+CC W
Sbjct: 7 LLLFFFLFTFLTTSFSKN------LCNQDDKTTLLKIKKALNNPY-HLASWDPRTDCCSW 59
Query: 63 SFVGCGRPYPG-RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK 121
+ CG RVT +TI G +SG +P E G+L YL L ++ +TG IP + +K
Sbjct: 60 YCLECGDATVNHRVTALTIFSGQ-ISGQIPPEVGDLSYLQTLVFRKLTNLTGQIPRTIAK 118
Query: 122 LQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN 181
L+ L++L L +L+GP+P FL +LK L+ VDLS N LSG+IP+SL L +L ++S N
Sbjct: 119 LKYLRSLRLSWTNLTGPVPGFLSELKNLQWVDLSFNDLSGSIPSSLSLLPNLVSLDLSRN 178
Query: 182 QLCGAIPAGLKKFNKNV 198
+L G+IP F V
Sbjct: 179 KLTGSIPESFGSFPAKV 195
Score = 84.0 bits (206), Expect = 3e-15
Identities = 64/178 (35%), Positives = 81/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ LP L L L+ K+TG IP SF ++ +L L N LSG IP LG
Sbjct: 156 LSGSIPSSLSLLPNLVSLDLSRN-KLTGSIPESFGSFPAKVPDLYLSHNQLSGYIPKTLG 214
Query: 145 KLKRLKEVDLSNNKLSG-----------------------------TIPASLGNLQ---- 171
L ++D S NKL G IP +LG L
Sbjct: 215 NLD-FSKIDFSRNKLGGDASMLFRANKTTWYIDLSRNMLQFDLSRVVIPKTLGILDLNHN 273
Query: 172 -------------SLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
L FNVS+N+LCG IP G L++F+ + HNKCLCGAPL CK
Sbjct: 274 GITGNIPVQWTEAPLQFFNVSYNRLCGHIPTGGTLQEFDTYSYFHNKCLCGAPLDSCK 331
>gb|AAP92912.1| polygalacturonase-inhibiting protein [Pyrus hybrid cultivar]
Length = 330
Score = 132 bits (331), Expect = 9e-30
Identities = 78/191 (40%), Positives = 108/191 (55%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
T L L+ ++ S N + CN DDK LL+I+ FG P L W ++T+CCDW V C
Sbjct: 7 TFLSLTLLFSSVLNPALSDLCNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA G+LPYL L + P +TGPI + +KL+ L+
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKF 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L +LSG +P FL +LK L +DLS N L+G IP+SL L +L + N+L G I
Sbjct: 124 LRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLDALRLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F NV
Sbjct: 184 PISFGQFIGNV 194
Score = 75.1 bits (183), Expect = 1e-12
Identities = 54/178 (30%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L L L + K+TG IP SF + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSELPNLDALRL-DRNKLTGHIPISFGQFIGNVPDLCLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFGSIDLSRNKLEGDASVIFVLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL CK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSCK 330
>dbj|BAB85784.1| polygalacturonase-inhibiting protein [Citrus latipes]
Length = 327
Score = 132 bits (331), Expect = 9e-30
Identities = 73/192 (38%), Positives = 110/192 (57%), Gaps = 6/192 (3%)
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG 66
L+L+F+ S++ S S + CN +DK LLK + P L W+ T+CCDW V
Sbjct: 7 LSLIFILSLFISPSLSD---LCNPNDKKVLLKFKKALNNPYV-LASWNPKTDCCDWYCVT 62
Query: 67 CGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQ 126
C R+ +TI G L G +P+E G+LPYL L ++P +TGPI + +KL+ L+
Sbjct: 63 CDLS-TNRINSLTIFAG-DLPGQIPSEVGDLPYLETLMFHKLPSLTGPIQPAIAKLKNLK 120
Query: 127 NLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGA 186
L + ++SGP+P F+ +L L ++LS N LSGTIP+SL LQ L ++ N+L G+
Sbjct: 121 TLRISWTNISGPVPDFISQLTNLTFLELSFNNLSGTIPSSLSKLQKLGALHLDRNKLTGS 180
Query: 187 IPAGLKKFNKNV 198
IP F ++
Sbjct: 181 IPESFGTFTGSI 192
Score = 85.1 bits (209), Expect = 1e-15
Identities = 58/174 (33%), Positives = 81/174 (46%), Gaps = 50/174 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSGT+P+ L L L L + K+TG IP SF + +L L N LSG IP+ LG
Sbjct: 153 LSGTIPSSLSKLQKLGALHL-DRNKLTGSIPESFGTFTGSIPDLYLSHNQLSGKIPASLG 211
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
+ + L +DL++NK
Sbjct: 212 SMDFNTIDLSRNKLEGGASFLFGLNKTTQRIDVSRNLLEFNLSKVEFPQSLTNLDLNHNK 271
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPL 210
+ G+IPA + +L++L NVS+N+LCG IP G L+ F + HN+CLCGAPL
Sbjct: 272 IFGSIPAQITSLENLGFLNVSYNRLCGPIPVGGKLQSFGYTEYFHNRCLCGAPL 325
>gb|AAQ56728.1| polygalacturonase inhibiting protein [Prunus persica]
Length = 330
Score = 131 bits (330), Expect = 1e-29
Identities = 79/211 (37%), Positives = 118/211 (55%), Gaps = 7/211 (3%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
TLL L+ ++ + N + CN +DK LL+I+ F P L W T+CCDW V C
Sbjct: 7 TLLCLTLLFSTILNQALSELCNPEDKKVLLQIKKAFNDPYV-LTSWKPETDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +P + G+LPYL L + P +TGPI S +KL+RL+
Sbjct: 66 DST-TNRINSLTIFSGQ-VSGQIPTQVGDLPYLETLEFHKQPNLTGPIQPSIAKLKRLKE 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L ++SG +P FL +LK L +DLS + L+G+IP+SL L +L+ ++ N+L G I
Sbjct: 124 LRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALHLDRNKLTGHI 183
Query: 188 PAGLKKFNKNVFE----HNKCLCGAPLAPCK 214
P +F+ +V E HN+ P + K
Sbjct: 184 PKSFGEFHGSVPELYLSHNQLSGNIPTSLAK 214
Score = 85.5 bits (210), Expect = 9e-16
Identities = 60/178 (33%), Positives = 83/178 (45%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
L+G++P+ LP L+ L L + K+TG IP SF + + L L N LSG IP+ L
Sbjct: 155 LTGSIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGEFHGSVPELYLSHNQLSGNIPTSLA 213
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
KL K L +DL++NK
Sbjct: 214 KLDFNRIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLISLDLNHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G IP L + L NVS+N+LCG IP G L+ F+ + + HN+CLCGAPL CK
Sbjct: 274 ITGGIPVGLTQVD-LQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSCK 330
>emb|CAF04462.1| putative polygalacturonase-inhibiting protein [Rubus idaeus]
Length = 331
Score = 130 bits (328), Expect = 2e-29
Identities = 77/194 (39%), Positives = 110/194 (56%), Gaps = 8/194 (4%)
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECC- 60
F L LTLLF + + + S CN DK L +I+ F P L W ++ +CC
Sbjct: 4 FKLFSLTLLFSTILTPALSE-----LCNPKDKKVLFEIKTAFNNPY-ILSSWKSDADCCT 57
Query: 61 DWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFS 120
DW V C P R+ +TI L+G +PA+ G+LPYL L L ++P +TGPI S +
Sbjct: 58 DWYCVECD-PTTHRINSLTIFTDNNLTGQIPAQVGDLPYLETLELRKLPHLTGPIQPSIA 116
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
KL+ L+ L L N LSG +P F+ +LK L ++L+ NK +G+IP+SL L +L ++
Sbjct: 117 KLKHLKMLRLSWNGLSGSVPDFISQLKNLTFLELNFNKFTGSIPSSLSQLPNLGALHLDR 176
Query: 181 NQLCGAIPAGLKKF 194
NQL G IP+ KF
Sbjct: 177 NQLTGQIPSSFGKF 190
Score = 87.4 bits (215), Expect = 2e-16
Identities = 53/154 (34%), Positives = 73/154 (46%), Gaps = 26/154 (16%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P+ FG ++TG IP SF+ + +DL N L G G
Sbjct: 179 LTGQIPSSFGKFVGTVPALFLSHNQLTGKIPTSFANMN-FDQIDLSRNKLEGDASVIFGL 237
Query: 146 LKR-----------------------LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
K L+ VDL++N ++G+IPA L L L FNVS+N+
Sbjct: 238 NKTTQIVDLSRNMLEFDLSKVVFSTSLRAVDLNHNSITGSIPAQLTQLDDLVLFNVSYNR 297
Query: 183 LCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
LCG IP G L+ + + HN+CLCGAPL CK
Sbjct: 298 LCGKIPVGGKLQSLDTTSYFHNRCLCGAPLPSCK 331
>gb|AAW72616.1| polygalacturonase-inhibiting protein [Prunus persica]
Length = 330
Score = 130 bits (328), Expect = 2e-29
Identities = 79/211 (37%), Positives = 118/211 (55%), Gaps = 7/211 (3%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
TLL L+ ++ S N + CN +DK LL+I+ F P L W T+CCDW V C
Sbjct: 7 TLLCLTLLFSSILNQALSELCNPEDKKVLLQIKKAFNDPYV-LASWKPETDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +P + G+LPYL L + P +TGPI S +KL+RL+
Sbjct: 66 DST-TNRINSLTIFSGQ-VSGQIPTQVGDLPYLETLEFHKQPNLTGPIQPSIAKLKRLKE 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L ++SG +P FL +LK L ++LS + L+G+IP+SL L +L+ ++ N+L G I
Sbjct: 124 LRLSWTNISGSVPDFLSQLKNLTFLELSFSNLTGSIPSSLSQLPNLNALHLDRNKLTGHI 183
Query: 188 PAGLKKFNKNVFE----HNKCLCGAPLAPCK 214
P +F+ +V E HN+ P + K
Sbjct: 184 PKSFGEFHGSVPELYLSHNQLSGNIPTSLAK 214
Score = 86.7 bits (213), Expect = 4e-16
Identities = 61/178 (34%), Positives = 83/178 (46%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
L+G++P+ LP L+ L L + K+TG IP SF + + L L N LSG IP+ L
Sbjct: 155 LTGSIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGEFHGSVPELYLSHNQLSGNIPTSLA 213
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
KL K L +DL++NK
Sbjct: 214 KLDFNRIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLISLDLNHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G IP L L L NVS+N+LCG IP G L+ F+ + + HN+CLCGAPL CK
Sbjct: 274 ITGGIPVGLTQLD-LQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSCK 330
>gb|AAP92913.1| polygalacturonase-inhibiting protein [Pyrus communis]
Length = 330
Score = 130 bits (327), Expect = 2e-29
Identities = 77/191 (40%), Positives = 108/191 (56%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
T L L+ ++ S N + CN DDK LL+I+ FG P L W ++T+CCDW V C
Sbjct: 7 TFLSLTLLFSSVLNPALSDLCNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA G+LPYL L + P +TGPI + + L+ L+
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIANLKGLKF 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L +LSG +P FL +LK L +DLS N L+G IP+SL L +L ++ N+L G I
Sbjct: 124 LRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALHLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F NV
Sbjct: 184 PISFGQFIGNV 194
Score = 75.1 bits (183), Expect = 1e-12
Identities = 54/178 (30%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L L L + K+TG IP SF + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSELPNLGALHL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFGSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL CK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSCK 330
>gb|AAM64993.1| polygalacturonase inhibiting protein [Arabidopsis thaliana]
gi|7800201|gb|AAF69828.1| polygalacturonase inhibiting
protein 2; PGIP2 [Arabidopsis thaliana]
Length = 326
Score = 130 bits (327), Expect = 2e-29
Identities = 77/190 (40%), Positives = 108/190 (56%), Gaps = 5/190 (2%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG- 68
L LS++ + S A D C+ DDK LLKI+ P L WD T+CC W + CG
Sbjct: 5 LLLSTLLLTTSLAKD--LCHKDDKTTLLKIKKSLNNPY-HLASWDPKTDCCSWYCLECGD 61
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
RVT + I G +SG +P E G+LPYL+ L ++ +TG I + +KL+ L L
Sbjct: 62 ATVNHRVTSLIIQDG-EISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLTFL 120
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L +L+GP+P FL +LK L+ +DLS N LSG+IP+SL +L+ L +S N+L G IP
Sbjct: 121 RLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGPIP 180
Query: 189 AGLKKFNKNV 198
F+ V
Sbjct: 181 ESFGTFSGKV 190
Score = 82.8 bits (203), Expect = 6e-15
Identities = 58/178 (32%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ +L L L L+ K+TGPIP SF ++ +L L N LSG IP LG
Sbjct: 151 LSGSIPSSLSSLRKLEYLELSRN-KLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLG 209
Query: 145 K----------------------------------------------LKRLKEVDLSNNK 158
K L +D+++N
Sbjct: 210 NPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLNNLDMNHNG 269
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G+IPA NVS+N+LCG IP G +++F+ F HNKCLCGAPL CK
Sbjct: 270 ITGSIPAEWSKAY-FQLLNVSYNRLCGRIPKGEYIQRFDSYSFFHNKCLCGAPLPSCK 326
>gb|AAM44964.1| putative polygalacturonase inhibiting protein [Arabidopsis
thaliana] gi|14334896|gb|AAK59626.1| putative
polygalacturonase inhibiting protein [Arabidopsis
thaliana] gi|9759543|dbj|BAB11145.1| polygalacturonase
inhibiting protein [Arabidopsis thaliana]
gi|15240183|ref|NP_196305.1| polygalacturonase
inhibiting protein 2 (PGIP2) [Arabidopsis thaliana]
gi|21263837|sp|Q9M5J8|PGIP2_ARATH Polygalacturonase
inhibitor 2 precursor (Polygalacturonase-inhibiting
protein) (PGIP-2)
Length = 330
Score = 130 bits (327), Expect = 2e-29
Identities = 77/190 (40%), Positives = 108/190 (56%), Gaps = 5/190 (2%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG- 68
L LS++ + S A D C+ DDK LLKI+ P L WD T+CC W + CG
Sbjct: 9 LLLSTLLLTTSLAKD--LCHKDDKTTLLKIKKSLNNPY-HLASWDPKTDCCSWYCLECGD 65
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
RVT + I G +SG +P E G+LPYL+ L ++ +TG I + +KL+ L L
Sbjct: 66 ATVNHRVTSLIIQDG-EISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLTFL 124
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L +L+GP+P FL +LK L+ +DLS N LSG+IP+SL +L+ L +S N+L G IP
Sbjct: 125 RLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGPIP 184
Query: 189 AGLKKFNKNV 198
F+ V
Sbjct: 185 ESFGTFSGKV 194
Score = 82.8 bits (203), Expect = 6e-15
Identities = 58/178 (32%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ +L L L L+ K+TGPIP SF ++ +L L N LSG IP LG
Sbjct: 155 LSGSIPSSLSSLRKLEYLELSRN-KLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLG 213
Query: 145 K----------------------------------------------LKRLKEVDLSNNK 158
K L +D+++N
Sbjct: 214 NPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLNNLDMNHNG 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G+IPA NVS+N+LCG IP G +++F+ F HNKCLCGAPL CK
Sbjct: 274 ITGSIPAEWSKAY-FQLLNVSYNRLCGRIPKGEYIQRFDSYSFFHNKCLCGAPLPSCK 330
>gb|AAW57429.1| polygalacturonase-inhibiting protein [Prunus americana]
gi|57868643|gb|AAW57430.1| polygalacturonase-inhibiting
protein [Prunus americana]
Length = 330
Score = 130 bits (326), Expect = 3e-29
Identities = 74/191 (38%), Positives = 110/191 (56%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
TLL L+ ++ + N + CN +DK LL+I+ F P L W T+CCDW V C
Sbjct: 7 TLLCLTLLFSTILNPALSELCNPEDKKVLLQIKKAFNDPYV-LTSWKPETDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +P + G+LPYL L + P +TGPI S +KL+RL+
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPTQVGDLPYLETLEFHKQPNLTGPIQPSIAKLKRLKE 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L ++SG +P FL +LK L +DLS + L+G+IP+SL L +L+ + N+L G I
Sbjct: 124 LRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALRLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F+ +V
Sbjct: 184 PKSFGEFHGSV 194
Score = 88.2 bits (217), Expect = 1e-16
Identities = 61/178 (34%), Positives = 84/178 (46%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
L+G++P+ LP L+ L L + K+TG IP SF + + +L L N LSG IP+ L
Sbjct: 155 LTGSIPSSLSQLPNLNALRL-DRNKLTGHIPKSFGEFHGSVPDLYLSHNQLSGTIPTSLA 213
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
KL K L +DL++NK
Sbjct: 214 KLNFTTIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSNVEFSKSLTSLDLNHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G IP L L L NVS+N+LCG IP G L+ F+ + + HN+CLCGAPL CK
Sbjct: 274 ITGGIPVGLTQLD-LQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSCK 330
>gb|AAP41199.1| polygalacturonase-inhibiting protein [Cucumis melo]
Length = 326
Score = 129 bits (325), Expect = 4e-29
Identities = 77/190 (40%), Positives = 103/190 (53%), Gaps = 5/190 (2%)
Query: 9 LLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG 68
LL L +F+ S A C+ +DK LL I+ F P L W +CC W V C
Sbjct: 6 LLLLLFFFFTVSFAE---LCHPNDKEVLLNIKKAFNNPY-ILTSWKPEEDCCTWYCVECD 61
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
R+ +TI LSG +P G+LP+L L ++P +TGPIP + +KL L+ L
Sbjct: 62 LK-SHRIIALTIFADDELSGPIPPFVGDLPFLENLMFHKLPNLTGPIPPTIAKLHNLKYL 120
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
DL N LSGPIPSFLG L L +DLS N+ +G+IP+SL NL+ L ++ N+L G IP
Sbjct: 121 DLSWNGLSGPIPSFLGSLSNLDILDLSFNRFTGSIPSSLANLRRLGTLHLDRNKLTGPIP 180
Query: 189 AGLKKFNKNV 198
F V
Sbjct: 181 DSFGNFKGKV 190
Score = 82.8 bits (203), Expect = 6e-15
Identities = 65/215 (30%), Positives = 96/215 (44%), Gaps = 58/215 (26%)
Query: 55 NNTECCDWSFVGCGRPYPG------RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEM 108
+N + D S+ G P P + ++ +S +G++P+ NL L L L +
Sbjct: 115 HNLKYLDLSWNGLSGPIPSFLGSLSNLDILDLSFN-RFTGSIPSSLANLRRLGTLHL-DR 172
Query: 109 PKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLGKL--------------------- 146
K+TGPIP+SF + ++ L L N LSG IP+ +GK+
Sbjct: 173 NKLTGPIPDSFGNFKGKVPYLYLSHNQLSGKIPTSMGKVDFNYIDLSRNKLVGDGSLIFG 232
Query: 147 -------------------------KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN 181
+ L +DL++NK+ G IP + L+ L NVS+N
Sbjct: 233 SKKTTEIVDLSRNLLEFNISKVVFPRTLTWLDLNHNKIFGEIPTGVVKLE-LQMLNVSYN 291
Query: 182 QLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
LCG IP G L+ F+ + HNKCLCG PL CK
Sbjct: 292 ALCGRIPMGGKLQSFDVYSYFHNKCLCGKPLGSCK 326
>dbj|BAB85785.1| polygalacturonase-inhibiting protein [Citrus hystrix]
Length = 327
Score = 129 bits (324), Expect = 6e-29
Identities = 72/188 (38%), Positives = 107/188 (56%), Gaps = 6/188 (3%)
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG 66
L+L+F+ S++ S + CN +DK LLK + P L W+ T+CCDW V
Sbjct: 7 LSLIFILSLFIPPSLSD---LCNPNDKKVLLKFKKALNNPYV-LASWNPKTDCCDWYCVT 62
Query: 67 CGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQ 126
C R+ +TI G L G +P+E G+LPYL L ++P +TGPI + +KL+ L+
Sbjct: 63 CDLS-TNRINSLTIFAG-DLPGQIPSEVGDLPYLETLMFHKLPSLTGPIQPAIAKLKNLK 120
Query: 127 NLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGA 186
L + ++SGP+P F+ +L L ++LS N LSGTIP+SL LQ L ++ N+L G+
Sbjct: 121 TLRISWTNISGPVPDFISQLTNLTFLELSFNNLSGTIPSSLSKLQKLGALHLDRNKLTGS 180
Query: 187 IPAGLKKF 194
IP F
Sbjct: 181 IPESFGTF 188
Score = 84.0 bits (206), Expect = 3e-15
Identities = 58/174 (33%), Positives = 80/174 (45%), Gaps = 50/174 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSGT+P+ L L L L + K+TG IP SF +L L N LSG IP+ LG
Sbjct: 153 LSGTIPSSLSKLQKLGALHL-DRNKLTGSIPESFGTFTGSTPDLYLSHNQLSGKIPASLG 211
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
+ + L +DL++NK
Sbjct: 212 SMDFNTIDLSRNKLEGDASFLIGLNKTTQRIDVSRNLLEFNLSKVEFPESLTNLDLNHNK 271
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPL 210
+ G+IPA + +L++L NVS+N+LCG IP G L+ F + HN+CLCGAPL
Sbjct: 272 IFGSIPAQITSLENLGFLNVSYNRLCGPIPVGGKLQSFGYTEYFHNRCLCGAPL 325
>gb|AAV33432.1| polygalacturonase inhibiting protein [Prunus mume]
Length = 330
Score = 128 bits (322), Expect = 9e-29
Identities = 79/211 (37%), Positives = 117/211 (55%), Gaps = 7/211 (3%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
TLL L+ ++ + N + CN +DK LL+I+ F P L W T+CCDW V C
Sbjct: 7 TLLCLTLLFSAILNPALSELCNQEDKKVLLQIKKAFNDPYV-LTSWKPETDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA+ G+LPYL L + P +TGPI S KL+ L+
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPAQVGDLPYLETLEFHKQPNLTGPIQPSIVKLKSLKF 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L ++SG +P FL +LK L +DLS + L+G+IP+SL L +L+ ++ N+L G I
Sbjct: 124 LRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALHLDRNKLTGHI 183
Query: 188 PAGLKKFNKNVFE----HNKCLCGAPLAPCK 214
P +F+ +V E HN+ P + K
Sbjct: 184 PKSFGEFHGSVPELYLSHNQLSGNIPTSLAK 214
Score = 85.5 bits (210), Expect = 9e-16
Identities = 60/178 (33%), Positives = 83/178 (45%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
L+G++P+ LP L+ L L + K+TG IP SF + + L L N LSG IP+ L
Sbjct: 155 LTGSIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGEFHGSVPELYLSHNQLSGNIPTSLA 213
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
KL K L +DL++NK
Sbjct: 214 KLDFNRIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLISLDLNHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G IP L + L NVS+N+LCG IP G L+ F+ + + HN+CLCGAPL CK
Sbjct: 274 ITGGIPVGLTQVD-LQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSCK 330
>gb|AAW72620.1| polygalacturonase-inhibiting protein [Prunus mume]
gi|58379370|gb|AAW72619.1| polygalacturonase-inhibiting
protein [Prunus mume]
Length = 330
Score = 128 bits (322), Expect = 9e-29
Identities = 79/211 (37%), Positives = 117/211 (55%), Gaps = 7/211 (3%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
TLL L+ ++ + N + CN +DK LL+I+ F P L W T+CCDW V C
Sbjct: 7 TLLCLTLLFTAILNPALSELCNQEDKKVLLQIKKAFNDPYV-LTSWKPETDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA+ G+LPYL L + P +TGPI S KL+ L+
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPAQVGDLPYLETLEFHKQPNLTGPIQPSIVKLKSLKF 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L ++SG +P FL +LK L +DLS + L+G+IP+SL L +L+ ++ N+L G I
Sbjct: 124 LRLSWTNISGSVPDFLSQLKNLTFLDLSFSNLTGSIPSSLSQLPNLNALHLDRNKLTGHI 183
Query: 188 PAGLKKFNKNVFE----HNKCLCGAPLAPCK 214
P +F+ +V E HN+ P + K
Sbjct: 184 PKSFGEFHGSVPELYLSHNQLSGNIPTSLAK 214
Score = 86.7 bits (213), Expect = 4e-16
Identities = 61/178 (34%), Positives = 83/178 (46%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
L+G++P+ LP L+ L L + K+TG IP SF + + L L N LSG IP+ L
Sbjct: 155 LTGSIPSSLSQLPNLNALHL-DRNKLTGHIPKSFGEFHGSVPELYLSHNQLSGNIPTSLA 213
Query: 145 KL----------------------------------------------KRLKEVDLSNNK 158
KL K L +DL++NK
Sbjct: 214 KLDFNRIDFSRNKLEGDASMIFGLNKTTQIVDLSRNLLEFNLSKVEFSKSLISLDLNHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G IP L L L NVS+N+LCG IP G L+ F+ + + HN+CLCGAPL CK
Sbjct: 274 ITGGIPVGLTQLD-LQFLNVSYNRLCGQIPVGGKLQSFDSSTYFHNRCLCGAPLPSCK 330
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 391,336,361
Number of Sequences: 2540612
Number of extensions: 17229043
Number of successful extensions: 75997
Number of sequences better than 10.0: 4106
Number of HSP's better than 10.0 without gapping: 2446
Number of HSP's successfully gapped in prelim test: 1698
Number of HSP's that attempted gapping in prelim test: 41387
Number of HSP's gapped (non-prelim): 20032
length of query: 214
length of database: 863,360,394
effective HSP length: 123
effective length of query: 91
effective length of database: 550,865,118
effective search space: 50128725738
effective search space used: 50128725738
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146750.9


