

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146723.12 + phase: 0
(172 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
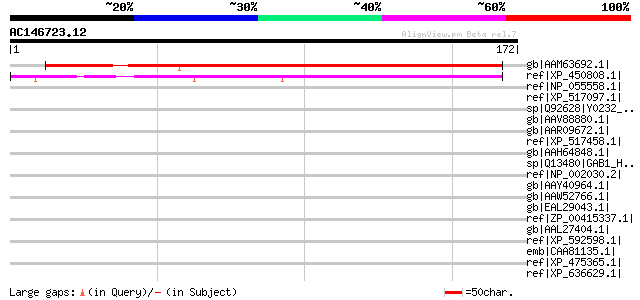
Sequences producing significant alignments: (bits) Value
gb|AAM63692.1| unknown [Arabidopsis thaliana] gi|5091545|gb|AAD3... 201 7e-51
ref|XP_450808.1| unknown protein [Oryza sativa (japonica cultiva... 148 7e-35
ref|NP_055558.1| hypothetical protein LOC9778 [Homo sapiens] 35 0.84
ref|XP_517097.1| PREDICTED: KIAA0232 gene product [Pan troglodytes] 35 0.84
sp|Q92628|Y0232_HUMAN Hypothetical protein KIAA0232 gi|20521850|... 35 0.84
gb|AAV88880.1| FAD/FMN-containing dehydrogenase [Zymomonas mobil... 35 1.1
gb|AAR09672.1| similar to Drosophila melanogaster Hsc70-4 [Droso... 33 2.5
ref|XP_517458.1| PREDICTED: similar to GRB2-associated binding p... 33 2.5
gb|AAH64848.1| GRB2-associated binding protein 1, isoform a [Hom... 33 2.5
sp|Q13480|GAB1_HUMAN GRB2-associated binding protein 1 (GRB2-ass... 33 2.5
ref|NP_002030.2| GRB2-associated binding protein 1 isoform b [Ho... 33 2.5
gb|AAY40964.1| unknown [Homo sapiens] 33 2.5
gb|AAW52766.1| HSP70 [Mytilus galloprovincialis] 33 2.5
gb|EAL29043.1| GA18066-PA [Drosophila pseudoobscura] 33 3.2
ref|ZP_00415337.1| thiol oxidoreductase [Azotobacter vinelandii ... 33 4.2
gb|AAL27404.1| 70 kDa heat shock protein [Artemia franciscana] 33 4.2
ref|XP_592598.1| PREDICTED: similar to Hypothetical protein KIAA... 32 5.5
emb|CAA81135.1| heat shock protein [Eimeria acervulina] gi|48067... 32 7.1
ref|XP_475365.1| putative hsp70 [Oryza sativa (japonica cultivar... 32 7.1
ref|XP_636629.1| hypothetical protein DDB0219366 [Dictyostelium ... 32 7.1
>gb|AAM63692.1| unknown [Arabidopsis thaliana] gi|5091545|gb|AAD39574.1| T10O24.14
[Arabidopsis thaliana] gi|18391169|ref|NP_563872.1|
expressed protein [Arabidopsis thaliana]
Length = 179
Score = 201 bits (511), Expect = 7e-51
Identities = 102/157 (64%), Positives = 118/157 (74%), Gaps = 7/157 (4%)
Query: 13 IPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRS--ITRAAEYKFPDPIP 70
+P SS C ++T QR KV SS+ R + RAAEYKFPDPIP
Sbjct: 26 LPIISSPAAVSCAIKSTQFFKQR-----CRTKVRDFSLSSLSRRGFVCRAAEYKFPDPIP 80
Query: 71 EFADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFID 130
EFA++ETEKF++H+LNKLSK+D+FE+SV+E+VGVCTEIF TFL SEYGGPGTLLV PFID
Sbjct: 81 EFAEAETEKFRDHMLNKLSKRDLFEDSVDEIVGVCTEIFETFLRSEYGGPGTLLVIPFID 140
Query: 131 MADIVNERGLPGGPQAARAAINWAQAHVDKDWNEWTG 167
MAD +NER LPGGPQAARAAI WAQ HVDKDW EWTG
Sbjct: 141 MADTLNERELPGGPQAARAAIKWAQDHVDKDWKEWTG 177
>ref|XP_450808.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|49387643|dbj|BAD25837.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 172
Score = 148 bits (373), Expect = 7e-35
Identities = 86/178 (48%), Positives = 106/178 (59%), Gaps = 19/178 (10%)
Query: 1 MASSTWC-------LVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSI 53
MA+ W L P L+ S+SS PS S+ + R R + P +
Sbjct: 1 MAARLWAAAVAPATLNPPLLTLSASSSPSS--SRLRRSVLGRL------RSRAPRPADFV 52
Query: 54 KRSITRAA--EYKFPDPIPEFADSETEKFQNHLLNKLSKK--DVFEESVEEVVGVCTEIF 109
R AA +YKFPDPIPEFA ET KF+ H++ +L +K D F E VEE+V VCTEI
Sbjct: 53 CRRAKNAAYDDYKFPDPIPEFAAQETSKFKEHMMWRLEQKKDDYFGEHVEEIVDVCTEIL 112
Query: 110 STFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQAARAAINWAQAHVDKDWNEWTG 167
TFL +Y GPGTLLV PF+DM + ERGLPG PQAARAAI WA+ ++DKDW WTG
Sbjct: 113 GTFLEHDYCGPGTLLVHPFLDMKGEIKERGLPGAPQAARAAIAWAEKNIDKDWKAWTG 170
>ref|NP_055558.1| hypothetical protein LOC9778 [Homo sapiens]
Length = 1395
Score = 35.0 bits (79), Expect = 0.84
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 13/87 (14%)
Query: 62 EYK-----FPDPIPEFAD--SETEKFQNHLLNKLSKKDVFEESVEE----VVGVCTEIFS 110
EYK + +PI E+ S K + N+ D+ E+VEE V G+C I +
Sbjct: 395 EYKEEPLWYTEPIAEYFVPLSRKSKLETTYRNRQDTSDLTSEAVEELSESVHGLC--ISN 452
Query: 111 TFLHSEYGGPGTLLVDPFIDMADIVNE 137
LH Y GT + F++M ++NE
Sbjct: 453 NNLHKTYLAAGTFIDGHFVEMPAVINE 479
>ref|XP_517097.1| PREDICTED: KIAA0232 gene product [Pan troglodytes]
Length = 1578
Score = 35.0 bits (79), Expect = 0.84
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 13/87 (14%)
Query: 62 EYK-----FPDPIPEFAD--SETEKFQNHLLNKLSKKDVFEESVEE----VVGVCTEIFS 110
EYK + +PI E+ S K + N+ D+ E+VEE V G+C I +
Sbjct: 443 EYKEEPLWYTEPIAEYFVPLSRKSKLETTYRNRQDTSDLTSEAVEELSESVHGLC--ISN 500
Query: 111 TFLHSEYGGPGTLLVDPFIDMADIVNE 137
LH Y GT + F++M ++NE
Sbjct: 501 NNLHKTYLAAGTFIDGHFVEMPAVINE 527
>sp|Q92628|Y0232_HUMAN Hypothetical protein KIAA0232 gi|20521850|dbj|BAA13221.3| KIAA0232
protein [Homo sapiens]
Length = 1402
Score = 35.0 bits (79), Expect = 0.84
Identities = 27/87 (31%), Positives = 41/87 (47%), Gaps = 13/87 (14%)
Query: 62 EYK-----FPDPIPEFAD--SETEKFQNHLLNKLSKKDVFEESVEE----VVGVCTEIFS 110
EYK + +PI E+ S K + N+ D+ E+VEE V G+C I +
Sbjct: 402 EYKEEPLWYTEPIAEYFVPLSRKSKLETTYRNRQDTSDLTSEAVEELSESVHGLC--ISN 459
Query: 111 TFLHSEYGGPGTLLVDPFIDMADIVNE 137
LH Y GT + F++M ++NE
Sbjct: 460 NNLHKTYLAAGTFIDGHFVEMPAVINE 486
>gb|AAV88880.1| FAD/FMN-containing dehydrogenase [Zymomonas mobilis subsp. mobilis
ZM4] gi|56551152|ref|YP_161991.1| FAD/FMN-containing
dehydrogenase [Zymomonas mobilis subsp. mobilis ZM4]
Length = 574
Score = 34.7 bits (78), Expect = 1.1
Identities = 30/123 (24%), Positives = 51/123 (41%), Gaps = 18/123 (14%)
Query: 38 FPSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEFADSETEKFQNHLLNKLSKKDVFEES 97
F +N +VS LP + + + A FP +P+ D +++++HL+ K+S EE
Sbjct: 353 FDGLNNRVSFLP-RYLSDRVMQFASTLFPHHLPKRMDDFRDRYEHHLILKVS-----EEG 406
Query: 98 VEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQAARAAINWAQAH 157
EE F GG + + +D ++ AA AAI + H
Sbjct: 407 TEE----ARSFLKDFFEKASGG--------YFECSDEEGKKAFLHRFAAAGAAIRYRDVH 454
Query: 158 VDK 160
D+
Sbjct: 455 QDE 457
>gb|AAR09672.1| similar to Drosophila melanogaster Hsc70-4 [Drosophila yakuba]
Length = 84
Score = 33.5 bits (75), Expect = 2.5
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 10/65 (15%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N+L+ K+ +E +E+ GVC I + S G PG + P G+PG
Sbjct: 17 NQLADKEEYEHRQKELEGVCNPIITKLYQSAGGAPGGMPGGP----------GGMPGAAG 66
Query: 146 AARAA 150
AA AA
Sbjct: 67 AAGAA 71
>ref|XP_517458.1| PREDICTED: similar to GRB2-associated binding protein 1 isoform a
[Pan troglodytes]
Length = 1089
Score = 33.5 bits (75), Expect = 2.5
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 762 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRTFPSD 821
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 822 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 851
>gb|AAH64848.1| GRB2-associated binding protein 1, isoform a [Homo sapiens]
gi|46370071|ref|NP_997006.1| GRB2-associated binding
protein 1 isoform a [Homo sapiens]
gi|63021430|gb|AAY26398.1| GRB2-associated binding
protein 1 [Homo sapiens]
Length = 724
Score = 33.5 bits (75), Expect = 2.5
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 356 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRAFPSD 415
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 416 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 445
>sp|Q13480|GAB1_HUMAN GRB2-associated binding protein 1 (GRB2-associated binder-1)
gi|1199618|gb|AAC50380.1| Gab1
Length = 694
Score = 33.5 bits (75), Expect = 2.5
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 356 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRAFPSD 415
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 416 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 445
>ref|NP_002030.2| GRB2-associated binding protein 1 isoform b [Homo sapiens]
Length = 694
Score = 33.5 bits (75), Expect = 2.5
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 356 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRAFPSD 415
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 416 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 445
>gb|AAY40964.1| unknown [Homo sapiens]
Length = 670
Score = 33.5 bits (75), Expect = 2.5
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 3/90 (3%)
Query: 14 PSSSSSIPSLCFSQTTTKCIQRFSF-PSINRKVSTLPFSSIKRSITRAAEYKFPDPIPEF 72
P + SIP ++ CI PS + +ST+ + +++ + Y P P
Sbjct: 332 PVETCSIPRTASDTDSSYCIPTAGMSPSRSNTISTVDLNKLRKDASSQDCYDIPRAFPSD 391
Query: 73 ADSETEKFQNH--LLNKLSKKDVFEESVEE 100
S E F NH + N L+ V E ++E
Sbjct: 392 RSSSLEGFHNHFKVKNVLTVGSVSSEELDE 421
>gb|AAW52766.1| HSP70 [Mytilus galloprovincialis]
Length = 654
Score = 33.5 bits (75), Expect = 2.5
Identities = 20/62 (32%), Positives = 27/62 (43%), Gaps = 7/62 (11%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N L++K+ FE +E+ GVC I + S G PG M + G PGG
Sbjct: 585 NNLAEKEEFEHKQKELEGVCNPIITKLYQSAGGAPGG-------GMPNFGGAGGAPGGAP 637
Query: 146 AA 147
A
Sbjct: 638 GA 639
>gb|EAL29043.1| GA18066-PA [Drosophila pseudoobscura]
Length = 652
Score = 33.1 bits (74), Expect = 3.2
Identities = 21/71 (29%), Positives = 31/71 (43%), Gaps = 13/71 (18%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N+L+ K+ +E +E+ G+C I + S G PG + G+PG P
Sbjct: 584 NQLADKEEYEHRQKELEGICNPIVTKLYQSTGGAPGGM-------------PGGMPGAPG 630
Query: 146 AARAAINWAQA 156
AA A A A
Sbjct: 631 AAGAGAPGAGA 641
>ref|ZP_00415337.1| thiol oxidoreductase [Azotobacter vinelandii AvOP]
gi|67087725|gb|EAM07191.1| thiol oxidoreductase
[Azotobacter vinelandii AvOP]
Length = 452
Score = 32.7 bits (73), Expect = 4.2
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 5/41 (12%)
Query: 115 SEYGGPGTLLVDPFIDMADIVNERGLPGGPQAARAAINWAQ 155
+++GGP P D D +N+ G+PG P RAAI W +
Sbjct: 121 NDHGGPL-----PHPDYGDQLNDTGIPGVPPEGRAAIAWEE 156
>gb|AAL27404.1| 70 kDa heat shock protein [Artemia franciscana]
Length = 644
Score = 32.7 bits (73), Expect = 4.2
Identities = 23/72 (31%), Positives = 36/72 (49%), Gaps = 11/72 (15%)
Query: 64 KFPDPIPEFADSET------EKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFLHSEY 117
KF D +PE AD T E + +N+L++K+ +EE +E+ VC I + Y
Sbjct: 557 KFKDKLPE-ADKNTILDKCNETIKWLDVNQLAEKEEYEEKQKEIEKVCNPIITKL----Y 611
Query: 118 GGPGTLLVDPFI 129
G G +L D +
Sbjct: 612 GQAGGMLADSLV 623
>ref|XP_592598.1| PREDICTED: similar to Hypothetical protein KIAA0232, partial [Bos
taurus]
Length = 1139
Score = 32.3 bits (72), Expect = 5.5
Identities = 26/87 (29%), Positives = 40/87 (45%), Gaps = 13/87 (14%)
Query: 62 EYK-----FPDPIPEFAD--SETEKFQNHLLNKLSKKDVFEESVEE----VVGVCTEIFS 110
EYK + +PI E+ S K + N+ DV +VEE V G+C I +
Sbjct: 230 EYKEEPLWYTEPIAEYFVPLSRKSKLETTYRNRQDAGDVTSAAVEELSESVHGLC--ISN 287
Query: 111 TFLHSEYGGPGTLLVDPFIDMADIVNE 137
+H Y GT + F++M ++NE
Sbjct: 288 NNIHKTYLAAGTFIDGHFVEMPAVINE 314
>emb|CAA81135.1| heat shock protein [Eimeria acervulina] gi|480678|pir||S37165
dnaK-type molecular chaperone - Eimeria acervulina
Length = 646
Score = 32.0 bits (71), Expect = 7.1
Identities = 20/59 (33%), Positives = 28/59 (46%), Gaps = 4/59 (6%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGP 144
N+L++K+ FE +EV VCT I + S G G + M D+ G GGP
Sbjct: 586 NQLAEKEEFEAKQKEVEAVCTPIVTKMYQSAAGAQGGMPG----GMPDMSAAAGAAGGP 640
>ref|XP_475365.1| putative hsp70 [Oryza sativa (japonica cultivar-group)]
gi|47900318|gb|AAT39165.1| putative hsp70 [Oryza sativa
(japonica cultivar-group)]
Length = 646
Score = 32.0 bits (71), Expect = 7.1
Identities = 20/61 (32%), Positives = 31/61 (50%), Gaps = 11/61 (18%)
Query: 86 NKLSKKDVFEESVEEVVGVCTEIFSTFLHSEYGGPGTLLVDPFIDMADIVNERGLPGGPQ 145
N+L++ D FE+ ++E+ G+C I + Y GPG DMA ++E GG
Sbjct: 589 NQLAEADEFEDKMKELEGICNPIIAKM----YQGPGA-------DMAGGMDEDAPAGGSG 637
Query: 146 A 146
A
Sbjct: 638 A 638
>ref|XP_636629.1| hypothetical protein DDB0219366 [Dictyostelium discoideum]
gi|60465017|gb|EAL63126.1| hypothetical protein
DDB0219366 [Dictyostelium discoideum]
Length = 499
Score = 32.0 bits (71), Expect = 7.1
Identities = 29/111 (26%), Positives = 47/111 (42%), Gaps = 2/111 (1%)
Query: 3 SSTWCLVPFLIPSSSSSIPSLCFSQTTTKCIQRFSFPSINRKVSTLPFSSIKRSITRAAE 62
SST + S++SSIPS S + T + S N ST SS + ++
Sbjct: 170 SSTASYTSTAVQSTASSIPSSPRSSSQTTSLSSSSSSISN--FSTSSNSSFSSFSSLSSS 227
Query: 63 YKFPDPIPEFADSETEKFQNHLLNKLSKKDVFEESVEEVVGVCTEIFSTFL 113
+ P P F K +++L L +K++ ES +++ E F L
Sbjct: 228 LEVPIPSNSFPTPTNVKLTSNILKHLPQKELKMESKSKLIRRAKEAFQFIL 278
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.132 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 317,232,501
Number of Sequences: 2540612
Number of extensions: 12851375
Number of successful extensions: 32589
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 32583
Number of HSP's gapped (non-prelim): 27
length of query: 172
length of database: 863,360,394
effective HSP length: 119
effective length of query: 53
effective length of database: 561,027,566
effective search space: 29734460998
effective search space used: 29734460998
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146723.12


