

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146719.12 + phase: 0 /pseudo
(758 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
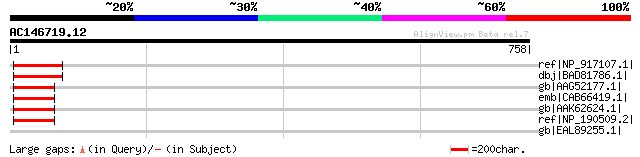
ref|NP_917107.1| P0421H07.9 [Oryza sativa (japonica cultivar-gro... 76 4e-12
dbj|BAD81786.1| transducin family protein / WD-40 repeat family ... 76 4e-12
gb|AAG52177.1| hypothetical protein; 45165-40041 [Arabidopsis th... 68 1e-09
emb|CAB66419.1| hypothetical protein [Arabidopsis thaliana] gi|1... 68 1e-09
gb|AAK62624.1| AT3g49400/F2K15_260 [Arabidopsis thaliana] 68 1e-09
ref|NP_190509.2| transducin family protein / WD-40 repeat family... 68 1e-09
gb|EAL89255.1| hypothetical protein Afu6g14190 [Aspergillus fumi... 36 4.1
>ref|NP_917107.1| P0421H07.9 [Oryza sativa (japonica cultivar-group)]
Length = 974
Score = 76.3 bits (186), Expect = 4e-12
Identities = 34/72 (47%), Positives = 49/72 (67%)
Query: 6 SPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNEPLILG 65
S Q A L+ SPS+PNA+AWS +NL+A ASGH +TIL P GPR L+ + P++P +G
Sbjct: 3 SHYQAATLIASPSYPNAIAWSTENLVAVASGHLITILNPSALEGPRELVVLRPSDPFPIG 62
Query: 66 FIDRKACLQLCI 77
++R+ + CI
Sbjct: 63 VVNREDLFEPCI 74
>dbj|BAD81786.1| transducin family protein / WD-40 repeat family protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 901
Score = 76.3 bits (186), Expect = 4e-12
Identities = 34/72 (47%), Positives = 49/72 (67%)
Query: 6 SPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNEPLILG 65
S Q A L+ SPS+PNA+AWS +NL+A ASGH +TIL P GPR L+ + P++P +G
Sbjct: 3 SHYQAATLIASPSYPNAIAWSTENLVAVASGHLITILNPSALEGPRELVVLRPSDPFPIG 62
Query: 66 FIDRKACLQLCI 77
++R+ + CI
Sbjct: 63 VVNREDLFEPCI 74
>gb|AAG52177.1| hypothetical protein; 45165-40041 [Arabidopsis thaliana]
Length = 798
Score = 68.2 bits (165), Expect = 1e-09
Identities = 33/60 (55%), Positives = 41/60 (68%)
Query: 6 SPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNEPLILG 65
S Q A L+ SPS+PNA+AWS +NLIA A+GH V I+ P LP GPRGLI + E +G
Sbjct: 3 SRFQEASLVTSPSYPNAVAWSSENLIAVAAGHLVIIINPALPTGPRGLITISDAELYQIG 62
>emb|CAB66419.1| hypothetical protein [Arabidopsis thaliana]
gi|11280702|pir||T45845 hypothetical protein F2K15.260
- Arabidopsis thaliana
Length = 719
Score = 68.2 bits (165), Expect = 1e-09
Identities = 33/60 (55%), Positives = 41/60 (68%)
Query: 6 SPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNEPLILG 65
S Q A L+ SPS+PNA+AWS +NLIA A+GH V I+ P LP GPRGLI + E +G
Sbjct: 3 SRFQEASLVTSPSYPNAVAWSSENLIAVAAGHLVIIINPALPTGPRGLITISDAELYQIG 62
>gb|AAK62624.1| AT3g49400/F2K15_260 [Arabidopsis thaliana]
Length = 793
Score = 68.2 bits (165), Expect = 1e-09
Identities = 33/60 (55%), Positives = 41/60 (68%)
Query: 6 SPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNEPLILG 65
S Q A L+ SPS+PNA+AWS +NLIA A+GH V I+ P LP GPRGLI + E +G
Sbjct: 3 SRFQEASLVTSPSYPNAVAWSSENLIAVAAGHLVIIINPALPTGPRGLITISDAELYQIG 62
>ref|NP_190509.2| transducin family protein / WD-40 repeat family protein
[Arabidopsis thaliana]
Length = 892
Score = 68.2 bits (165), Expect = 1e-09
Identities = 33/60 (55%), Positives = 41/60 (68%)
Query: 6 SPLQPAMLMGSPSFPNALAWSHDNLIAAASGHFVTILRPDLPNGPRGLIKVIPNEPLILG 65
S Q A L+ SPS+PNA+AWS +NLIA A+GH V I+ P LP GPRGLI + E +G
Sbjct: 3 SRFQEASLVTSPSYPNAVAWSSENLIAVAAGHLVIIINPALPTGPRGLITISDAELYQIG 62
>gb|EAL89255.1| hypothetical protein Afu6g14190 [Aspergillus fumigatus Af293]
Length = 700
Score = 36.2 bits (82), Expect = 4.1
Identities = 18/32 (56%), Positives = 20/32 (62%)
Query: 16 SPSFPNALAWSHDNLIAAASGHFVTILRPDLP 47
SPS N LAWS D IA A+G +V IL P P
Sbjct: 10 SPSCYNCLAWSADGEIALAAGEYVQILTPKKP 41
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.360 0.162 0.604
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,109,768,289
Number of Sequences: 2540612
Number of extensions: 40673950
Number of successful extensions: 139203
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 139196
Number of HSP's gapped (non-prelim): 7
length of query: 758
length of database: 863,360,394
effective HSP length: 136
effective length of query: 622
effective length of database: 517,837,162
effective search space: 322094714764
effective search space used: 322094714764
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC146719.12


