

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146706.4 + phase: 0
(478 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
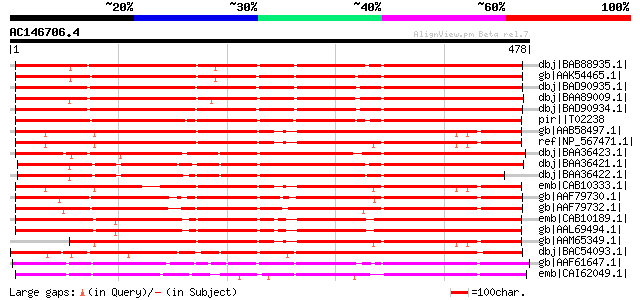
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB88935.1| glucosyltransferase NTGT2 [Nicotiana tabacum] 516 e-145
gb|AAK54465.1| cold-induced glucosyl transferase [Solanum sogara... 513 e-144
dbj|BAD90935.1| monoterpene glucosyltransferase [Eucalyptus perr... 498 e-139
dbj|BAA89009.1| anthocyanin 5-O-glucosyltransferase [Petunia x h... 490 e-137
dbj|BAD90934.1| monoterpene glucosyltransferase [Eucalyptus perr... 489 e-137
pir||T02238 glucosyl transferase, jasmonate-induced - common tob... 471 e-131
gb|AAB58497.1| UDP-glucose:indole-3-acetate beta-D-glucosyltrans... 447 e-124
ref|NP_567471.1| UDP-glucose:indole-3-acetate beta-D-glucosyltra... 446 e-123
dbj|BAA36423.1| UDP-glucose:anthocyanin 5-O-glucosyltransferase ... 435 e-120
dbj|BAA36421.1| UDP-glucose:anthocysnin 5-O-glucosyltransferase ... 434 e-120
dbj|BAA36422.1| UDP-glucose:anthocyanin 5-O-glucosyltransferase ... 412 e-114
emb|CAB10333.1| glucosyltransferase like protein [Arabidopsis th... 411 e-113
gb|AAF79730.1| T25N20.21 [Arabidopsis thaliana] gi|18700284|gb|A... 411 e-113
gb|AAF79732.1| T25N20.18 [Arabidopsis thaliana] gi|15220556|ref|... 405 e-111
emb|CAB10189.1| glucosyltransferase like protein [Arabidopsis th... 402 e-110
gb|AAL69494.1| putative glucosyltransferase [Arabidopsis thaliana] 402 e-110
gb|AAM65349.1| AT4g15550/dl3815c [Arabidopsis thaliana] gi|16604... 400 e-110
dbj|BAC54093.1| anthocyanin 5-glucosyltransferase [Torenia hybrida] 363 5e-99
gb|AAF61647.1| UDP-glucose:salicylic acid glucosyltransferase [N... 326 1e-87
emb|CAI62049.1| UDP-xylose phenolic glycosyltransferase [Lycoper... 319 1e-85
>dbj|BAB88935.1| glucosyltransferase NTGT2 [Nicotiana tabacum]
Length = 470
Score = 516 bits (1328), Expect = e-145
Identities = 266/473 (56%), Positives = 343/473 (72%), Gaps = 17/473 (3%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIP---GLSFATFS 62
H L++T+P QGHINP LQF KRLI +G +VTFAT++ + R+ T GL+FA FS
Sbjct: 5 HVLLVTFPAQGHINPCLQFAKRLIRMGIEVTFATSVFAHRRMAKTTTSTLSKGLNFAAFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DGYDDG K+ + D YMSE RGS+ L +IIL S E P T L+Y+L+L WA KVA
Sbjct: 65 DGYDDGFKA-DEHDSQHYMSEIKSRGSKTLKDIILKSSDEGRPVTSLVYSLLLPWAAKVA 123
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
E H+P LLWIQ ATV DI+YYYF+ + D I + D I LP L LKS+DLPSF
Sbjct: 124 REFHIPCALLWIQPATVLDIYYYYFNGYEDAIKGSTNDPNWCIQLPRLPL-LKSQDLPSF 182
Query: 183 LLASNT---YTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGP 239
LL+S+ Y+FALP+ KEQ+ L+ E NP+VLVNT + E L ++ K +I IGP
Sbjct: 183 LLSSSNEEKYSFALPTFKEQLDTLDVEENPKVLVNTFDALEPKELKAIE--KYNLIGIGP 240
Query: 240 LIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEE 299
LIPS FLDGKDP D+SFGGD+ + +DYI+WL+SK SVVY+SFG+L LSK Q EE
Sbjct: 241 LIPSTFLDGKDPLDSSFGGDLFQ--KSNDYIEWLNSKANSSVVYISFGSLLNLSKNQKEE 298
Query: 300 IARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSH 359
IA+ L++ FLWVIRD++ + E+E ++LSC ELE GKIV WCSQ+EVL+H
Sbjct: 299 IAKGLIEIKKPFLWVIRDQENGKGDEKE---EKLSCMMELEKQ--GKIVPWCSQLEVLTH 353
Query: 360 RSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVK 419
S+GCF++HCGWNSTLESL SGV +VAFP WTDQ TNAKLIEDVWKTG+R++ +E+G+V+
Sbjct: 354 PSIGCFVSHCGWNSTLESLSSGVSVVAFPHWTDQGTNAKLIEDVWKTGVRLKKNEDGVVE 413
Query: 420 VEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
EEI++C+E+VM GEKGEE+RRNA+KWK+LAR AVKEGGSS NL++++ ++
Sbjct: 414 SEEIKRCIEMVMDGGEKGEEMRRNAQKWKELAREAVKEGGSSEMNLKAFVQEV 466
>gb|AAK54465.1| cold-induced glucosyl transferase [Solanum sogarandinum]
Length = 473
Score = 513 bits (1321), Expect = e-144
Identities = 263/474 (55%), Positives = 347/474 (72%), Gaps = 16/474 (3%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIP---GLSFATFS 62
H L++T+P QGHINP+LQF K+LI +G +VTF T++ + R+ T GL+ A FS
Sbjct: 5 HVLLVTFPTQGHINPSLQFAKKLIKMGIEVTFTTSVFAHRRMAKTATSTAPKGLNLAAFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DG+DDG KS D D YMSE RGS+ L +IIL S E P T L+YTL+L WA +VA
Sbjct: 65 DGFDDGFKSNVD-DSKRYMSEIRSRGSQTLRDIILKSSDEGRPVTSLVYTLLLPWAAEVA 123
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
ELH+PS LLWIQ ATV DI+YYYF+ + D + S D I LP L LKS+DLPSF
Sbjct: 124 RELHIPSALLWIQPATVLDIYYYYFNGYEDEMKCSSNDPNWSIQLPRLPL-LKSQDLPSF 182
Query: 183 LLASNT----YTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIG 238
L++S++ Y+FALP+ KEQ+ L+ E NP+VLVNT + EL+ L + GK +I IG
Sbjct: 183 LVSSSSKDDKYSFALPTFKEQLDTLDGEENPKVLVNTFDALELEPLKAI--GKYNLIGIG 240
Query: 239 PLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQME 298
PLIPS+FL GKD ++ FGGD+ + S DDY++WL++K + S+VY+SFG+L LS+ Q E
Sbjct: 241 PLIPSSFLGGKDSLESRFGGDLFQ-KSNDDYMEWLNTKPKSSIVYISFGSLLNLSRNQKE 299
Query: 299 EIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLS 358
EIA+ L++ FLWVIRD+ + KE E ++++LSC ELE GKIV WCSQ+EVL+
Sbjct: 300 EIAKGLIEIKRPFLWVIRDQ--ENIKEVEKEEEKLSCMMELEKQ--GKIVPWCSQLEVLT 355
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMV 418
H SLGCF++HCGWNSTLESL SGVP+VAFP WTDQ TNAK IEDVWKTG+RM +E+G+V
Sbjct: 356 HPSLGCFVSHCGWNSTLESLSSGVPVVAFPHWTDQGTNAKWIEDVWKTGVRMRVNEDGVV 415
Query: 419 KVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+ EEI++C+E+VM GEKGEE+R+NA+KWK+LAR AVKEGGSS NL++++ ++
Sbjct: 416 ESEEIKRCIEIVMDGGEKGEEMRKNAQKWKELAREAVKEGGSSEVNLKAFVQEV 469
>dbj|BAD90935.1| monoterpene glucosyltransferase [Eucalyptus perriniana]
Length = 467
Score = 498 bits (1283), Expect = e-139
Identities = 253/468 (54%), Positives = 330/468 (70%), Gaps = 13/468 (2%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
++L++ +P QG INPALQF KRL+ GA VTFAT Y R+ GLSFA+FSDG
Sbjct: 5 NYLVVAFPAQGLINPALQFAKRLLHAGAHVTFATAASAYRRMAKSDPPQGLSFASFSDGS 64
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHEL 125
++G + D YM++ R GSE L +++++S E F C+ YT I+ WA +VAH L
Sbjct: 65 EEGLRP--GIDFEQYMADAERLGSETLRDLVVTSLNEGRKFECMFYTTIVPWAGQVAHSL 122
Query: 126 HLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDE-TCLISLPGLSFSLKSRDLPSFLL 184
+PSTL+W Q AT+ DI+YYYF+ +GD I N KD+ + + LPGL L SRD+PSF
Sbjct: 123 QIPSTLIWAQPATLLDIYYYYFNGYGDIIRNLGKDDPSASLHLPGLP-PLTSRDVPSFFT 181
Query: 185 ASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIPSA 244
N Y F L ++ Q ++ EE NPRVLVNT + E L + G + M+ IGPLIPSA
Sbjct: 182 PENQYAFTLSLMRVQFEVFKEEKNPRVLVNTFDALETGPLKAI--GNVTMLGIGPLIPSA 239
Query: 245 FLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARAL 304
FLDG+DP D SFGGD+ + DYI+WLD+K + SV+YVSFG+++VLSK Q EE+AR L
Sbjct: 240 FLDGQDPLDKSFGGDLFQ--GSKDYIRWLDTKPKGSVIYVSFGSISVLSKEQKEEMARGL 297
Query: 305 LDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGC 364
L +G FLWVIR K ++E E +DD+LSC EELE G IV WCSQVEVLSH S+GC
Sbjct: 298 LGTGRPFLWVIRKDK---REEGEGEDDQLSCVEELEQK--GMIVPWCSQVEVLSHASVGC 352
Query: 365 FMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIR 424
F+TH GWNST ESL GVPMVAFPQWTDQ TNA L+E+ WK G+R+ +E G+V+ +E++
Sbjct: 353 FVTHSGWNSTFESLACGVPMVAFPQWTDQQTNAMLVENEWKVGVRVSTNERGIVEGDELK 412
Query: 425 KCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+CLE+V+G GE+GEE+RRNA+KWK LAR A KEGGSS+RNL+ +L +I
Sbjct: 413 RCLELVVGDGEEGEEIRRNAEKWKGLAREAAKEGGSSDRNLKEFLEEI 460
>dbj|BAA89009.1| anthocyanin 5-O-glucosyltransferase [Petunia x hybrida]
Length = 468
Score = 490 bits (1261), Expect = e-137
Identities = 250/474 (52%), Positives = 337/474 (70%), Gaps = 19/474 (4%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTI---PGLSFATFS 62
H ++ T+P QGHINPALQF K L+ +G +VTF+T+I+ SR+ K + GL+F FS
Sbjct: 5 HVILTTFPAQGHINPALQFAKNLVKMGIEVTFSTSIYAQSRMDEKSILNAPKGLNFIPFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DG+D+G +D V YMS+ + GSE + IIL+ + P TCL+Y++ L WA +VA
Sbjct: 65 DGFDEGFDH--SKDPVFYMSQLRKCGSETVKKIILTCSENGQPITCLLYSIFLPWAAEVA 122
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
E+H+PS LLW Q AT+ DI+Y+ FH + + N+S D I LPGL L++RDLPSF
Sbjct: 123 REVHIPSALLWSQPATILDIYYFNFHGYEKAMANESNDPNWSIQLPGLPL-LETRDLPSF 181
Query: 183 LL---ASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGP 239
LL A + ALP KE I L+ E P++LVNT +E E +ALN ++ K IGP
Sbjct: 182 LLPYGAKGSLRVALPPFKELIDTLDAETTPKILVNTFDELEPEALNAIE--GYKFYGIGP 239
Query: 240 LIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEE 299
LIPSAFL G DP D SFGGD+ + + +DY++WL+SK SVVY+SFG+L S QMEE
Sbjct: 240 LIPSAFLGGNDPLDASFGGDLFQ--NSNDYMEWLNSKPNSSVVYISFGSLMNPSISQMEE 297
Query: 300 IARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSH 359
I++ L+D G FLWVI++ +K +E ++ +L C EELE GKIV WCSQ+EVL H
Sbjct: 298 ISKGLIDIGRPFLWVIKEN----EKGKEEENKKLGCIEELEKI--GKIVPWCSQLEVLKH 351
Query: 360 RSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVK 419
SLGCF++HCGWNS LESL GVP+VAFPQWTDQ TNAK +EDVWK+G+R+ +E+G+V+
Sbjct: 352 PSLGCFVSHCGWNSALESLACGVPVVAFPQWTDQMTNAKQVEDVWKSGVRVRINEDGVVE 411
Query: 420 VEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIA 473
EEI++C+E+VM GEKGEELR+NAKKWK+LAR AVKEGGSS++NL+++++D+A
Sbjct: 412 SEEIKRCIELVMDGGEKGEELRKNAKKWKELAREAVKEGGSSHKNLKAFIDDVA 465
>dbj|BAD90934.1| monoterpene glucosyltransferase [Eucalyptus perriniana]
Length = 467
Score = 489 bits (1260), Expect = e-137
Identities = 253/468 (54%), Positives = 321/468 (68%), Gaps = 13/468 (2%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
++L++ +P QG INPALQ KRL+ GA VTFAT Y R+ GLSFA+FSDG
Sbjct: 5 NYLVVAFPAQGLINPALQIAKRLLHAGAHVTFATAGSAYRRMAKSDPPEGLSFASFSDGS 64
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHEL 125
D+G K D YM + R GSE L +++++S E F C+ YT I+ W +VAH L
Sbjct: 65 DEGLKP--GIDFNQYMVDVERLGSETLRDLVVTSLNEGRKFACIFYTTIIPWVAQVAHSL 122
Query: 126 HLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDE-TCLISLPGLSFSLKSRDLPSFLL 184
+PSTL+W Q AT+ DI+YYYF+ +GD I N KD+ + L+ LPGL L D+PSF
Sbjct: 123 QIPSTLIWTQPATLLDIYYYYFNGYGDIIRNLGKDDPSALLHLPGLP-PLTPPDIPSFFT 181
Query: 185 ASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIPSA 244
N Y F LP ++ Q +L EE PRVLVNT + E L + G + M IGPLIPSA
Sbjct: 182 PDNQYAFTLPLMQMQFELFKEEKYPRVLVNTFDALEPGPLKAI--GNVTMFGIGPLIPSA 239
Query: 245 FLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARAL 304
FLDG+DP D SFGGD+ + YIQWLD+K + SV+YVSFG+++VLSK Q EE+AR L
Sbjct: 240 FLDGQDPLDKSFGGDLFQ--GSKGYIQWLDTKPKGSVIYVSFGSISVLSKAQKEEMARGL 297
Query: 305 LDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGC 364
L +G FLWVIR K +E E + D LSC EELE G IV WCSQVEVLSH S+GC
Sbjct: 298 LGTGHPFLWVIRKDK---DEEGEGEQDHLSCMEELEQK--GMIVPWCSQVEVLSHASVGC 352
Query: 365 FMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIR 424
F+TH GWNST ESL GVPMVAFPQW DQ TNA L+E+ WK G+R+ +E G+V+ +EI+
Sbjct: 353 FVTHSGWNSTFESLACGVPMVAFPQWNDQLTNAMLVENEWKVGVRVNVNEGGVVEGDEIK 412
Query: 425 KCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+CLE+V+G GE+GEE+RRNAKKWK LAR A KEGGSS+RNL+++L +I
Sbjct: 413 RCLELVVGDGEQGEEIRRNAKKWKHLAREAAKEGGSSDRNLKAFLEEI 460
>pir||T02238 glucosyl transferase, jasmonate-induced - common tobacco
gi|1805359|dbj|BAA19155.1| glucosyl transferase
[Nicotiana tabacum]
Length = 467
Score = 471 bits (1213), Expect = e-131
Identities = 240/467 (51%), Positives = 332/467 (70%), Gaps = 15/467 (3%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H LI +P QGHINP+LQF+K+LI+LG KVT ++++ ++R+ N P I GL+FA FSDGY
Sbjct: 9 HVLIALFPGQGHINPSLQFSKKLINLGVKVTLSSSLSAFNRIKNLPKIEGLTFAPFSDGY 68
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHEL 125
D K D D + S GSEF+ N+I S + +PFT +IYT+++ WA VA +L
Sbjct: 69 DGNFKGSFD-DYHLFNSAIKSHGSEFIANLIKSKAKNGYPFTRVIYTILMDWAGSVAKKL 127
Query: 126 HLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFLLA 185
H+PSTL WIQ ATVFDI+YY F +Y N + +I LPGL SL S D PSF+
Sbjct: 128 HIPSTLFWIQPATVFDIYYYRFTNFANYFKNYDSQDQ-IIELPGLP-SLSSSDFPSFVFD 185
Query: 186 S-NTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIPSA 244
+ +A+ S+K QI++LN E NPR+LVNT + EL+AL + + M+ IGPLIPS+
Sbjct: 186 DVKSNDWAVESIKRQIEILNSEENPRILVNTFDALELNALRVLK--NVTMVGIGPLIPSS 243
Query: 245 FLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIARAL 304
FLD KD DN F D++ +S+++Y++WLD++ KSV+Y++FG+ A +S + MEEI++ L
Sbjct: 244 FLDEKDRKDNFFAADMI--ESENNYMEWLDARANKSVIYIAFGSYAEISSQWMEEISQGL 301
Query: 305 LDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSLGC 364
L G FLWVIR+ L +K EE +L+C++ELE G+IV+WCSQ+EVL H S+GC
Sbjct: 302 LKCGRPFLWVIRET-LNGEKPEE----KLTCKDELEKI--GRIVRWCSQMEVLKHSSVGC 354
Query: 365 FMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEEIR 424
F+THCGWNSTLESL SGVP+VA P W DQ NAKLI+DVWK G+R+ ++EG++K +E +
Sbjct: 355 FLTHCGWNSTLESLASGVPIVACPIWNDQICNAKLIQDVWKIGVRVNANKEGIIKRDEFQ 414
Query: 425 KCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
KC+E+VMG E+GEELR+NA+KWKDLA+ + KE SSN NL++Y+N+
Sbjct: 415 KCIEIVMGDAEEGEELRKNAQKWKDLAKESTKENSSSNVNLKAYVNE 461
>gb|AAB58497.1| UDP-glucose:indole-3-acetate beta-D-glucosyltransferase
[Arabidopsis thaliana]
Length = 474
Score = 447 bits (1150), Expect = e-124
Identities = 233/480 (48%), Positives = 326/480 (67%), Gaps = 34/480 (7%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISL--GAKVTFATTIHLYSR-LINKPTIPG-LSFATF 61
HFL +T+P QGHINP+L+ KRL GA+VTFA +I Y+R + + +P L FAT+
Sbjct: 13 HFLFVTFPAQGHINPSLELAKRLAGTISGARVTFAASISAYNRRMFSTENVPETLIFATY 72
Query: 62 SDGYDDGQKSFGDED------IVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
SDG+DDG KS D ++MSE RRG E LT +I ++++N PFTC++YT++L
Sbjct: 73 SDGHDDGFKSSAYSDKSRQDATGNFMSEMRRRGKETLTELIEDNRKQNRPFTCVVYTILL 132
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLK 175
+W ++A E HLPS LLW+Q TVF IFY+YF+ + D I+ + + I LP L L
Sbjct: 133 TWVAELAREFHLPSALLWVQPVTVFSIFYHYFNGYEDAISEMANTPSSSIKLPSLPL-LT 191
Query: 176 SRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMI 235
RD+PSF+++SN Y F LP+ +EQI L EEINP++L+NT +E E +A++ V K++
Sbjct: 192 VRDIPSFIVSSNVYAFLLPAFREQIDSLKEEINPKILINTFQELEPEAMSSVP-DNFKIV 250
Query: 236 PIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKR 295
P+GPL+ TD S S+ +YI+WLD+K + SV+YVSFGTLAVLSK+
Sbjct: 251 PVGPLLTLR-------TDFS---------SRGEYIEWLDTKADSSVLYVSFGTLAVLSKK 294
Query: 296 QMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVE 355
Q+ E+ +AL+ S FLWVI DK + +++E+ +++ E + G +V WC Q
Sbjct: 295 QLVELCKALIQSRRPFLWVITDKSYRNKEDEQEKEEDCISSSEKSFDEIGMVVSWCDQFR 354
Query: 356 VLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRM--EHD 413
VL+HRS+GCF+THCGWNSTLESL SGVP+VAFPQW DQ TNAKL+ED WKTG+R+ + +
Sbjct: 355 VLNHRSIGCFVTHCGWNSTLESLVSGVPVVAFPQWNDQMTNAKLLEDCWKTGVRVMEKKE 414
Query: 414 EEGMVKV--EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
EEG+V V EEIR+C+E VM +K EE R NA +WKDLA AV+EGGSS +L++++++
Sbjct: 415 EEGVVVVDSEEIRRCIEEVM--EDKAEEFRGNATRWKDLAAEAVREGGSSFNHLKAFVDE 472
>ref|NP_567471.1| UDP-glucose:indole-3-acetate beta-D-glucosyltransferase (IAGLU)
[Arabidopsis thaliana]
Length = 474
Score = 446 bits (1146), Expect = e-123
Identities = 236/482 (48%), Positives = 329/482 (67%), Gaps = 38/482 (7%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISL--GAKVTFATTIHLYSR-LINKPTIPG-LSFATF 61
HFL +T+P QGHINP+L+ KRL GA+VTFA +I Y+R + + +P L FAT+
Sbjct: 13 HFLFVTFPAQGHINPSLELAKRLAGTISGARVTFAASISAYNRRMFSTENVPETLIFATY 72
Query: 62 SDGYDDGQKSFGDED------IVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
SDG+DDG KS D ++MSE RRG E LT +I ++++N PFTC++YT++L
Sbjct: 73 SDGHDDGFKSSAYSDKSRQDATGNFMSEMRRRGKETLTELIEDNRKQNRPFTCVVYTILL 132
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLK 175
+W ++A E HLPS LLW+Q TVF IFY+YF+ + D I+ + + I LP L L
Sbjct: 133 TWVAELAREFHLPSALLWVQPVTVFSIFYHYFNGYEDAISEMANTPSSSIKLPSLPL-LT 191
Query: 176 SRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMI 235
RD+PSF+++SN Y F LP+ +EQI L EEINP++L+NT +E E +A++ V K++
Sbjct: 192 VRDIPSFIVSSNVYAFLLPAFREQIDSLKEEINPKILINTFQELEPEAMSSVP-DNFKIV 250
Query: 236 PIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKR 295
P+GPL+ TD S S+ +YI+WLD+K + SV+YVSFGTLAVLSK+
Sbjct: 251 PVGPLLTLR-------TDFS---------SRGEYIEWLDTKADSSVLYVSFGTLAVLSKK 294
Query: 296 QMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDEL--SCREELENNMNGKIVKWCSQ 353
Q+ E+ +AL+ S FLWVI DK + +++E+ +++ S REEL+ G +V WC Q
Sbjct: 295 QLVELCKALIQSRRPFLWVITDKSYRNKEDEQEKEEDCISSFREELDEI--GMVVSWCDQ 352
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRM--E 411
VL+HRS+GCF+THCGWNSTLESL SGVP+VAFPQW DQ NAKL+ED WKTG+R+ +
Sbjct: 353 FRVLNHRSIGCFVTHCGWNSTLESLVSGVPVVAFPQWNDQMMNAKLLEDCWKTGVRVMEK 412
Query: 412 HDEEGMVKV--EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYL 469
+EEG+V V EEIR+C+E VM +K EE R NA +WKDLA AV+EGGSS +L++++
Sbjct: 413 KEEEGVVVVDSEEIRRCIEEVM--EDKAEEFRGNATRWKDLAAEAVREGGSSFNHLKAFV 470
Query: 470 ND 471
++
Sbjct: 471 DE 472
>dbj|BAA36423.1| UDP-glucose:anthocyanin 5-O-glucosyltransferase [Verbena x hybrida]
Length = 461
Score = 435 bits (1118), Expect = e-120
Identities = 243/478 (50%), Positives = 318/478 (65%), Gaps = 29/478 (6%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPG----LSFATF 61
H L+ T+P QGHINPALQF KRL + +VTF T+++ + R+ T G ++F +F
Sbjct: 5 HVLLATFPAQGHINPALQFAKRLANADIQVTFFTSVYAWRRMSR--TAAGSNGLINFVSF 62
Query: 62 SDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSK--QENHPFTCLIYTLILSWAP 119
SDGYDDG + +D +YMSE RG + L++ + ++ Q++ T ++Y+ + +WA
Sbjct: 63 SDGYDDGLQP--GDDGKNYMSEMKSRGIKALSDTLAANNVDQKSSKITFVVYSHLFAWAA 120
Query: 120 KVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDL 179
KVA E HL S LLWI+ ATV DIFY+YF+ + D I S I LPG L RDL
Sbjct: 121 KVAREFHLRSALLWIEPATVLDIFYFYFNGYSDEIDAGSD----AIHLPGGLPVLAQRDL 176
Query: 180 PSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGP 239
PSFLL S F +KE+++ L E P+VLVN+ + E DAL +D K +MI IGP
Sbjct: 177 PSFLLPSTHERFR-SLMKEKLETLEGEEKPKVLVNSFDALEPDALKAID--KYEMIAIGP 233
Query: 240 LIPSAFLDGKDPTDNSFGGDVVRVDSKDD-YIQWLDSKDEKSVVYVSFGTLAVLSKRQME 298
LIPSAFLDGKDP+D SFGGD+ S DD ++WL + SVVYVSFG+ +K QME
Sbjct: 234 LIPSAFLDGKDPSDRSFGGDLFEKGSNDDDCLEWLSTNPRSSVVYVSFGSFVNTTKSQME 293
Query: 299 EIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLS 358
EIAR LLD G FLWV+R E ++ +SC EEL+ GKIV WCSQ+EVL+
Sbjct: 294 EIARGLLDCGRPFLWVVR--------VNEGEEVLISCMEELKRV--GKIVSWCSQLEVLT 343
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEG-M 417
H SLGCF+THCGWNSTLES+ GVPMVAFPQW DQ TNAKL+EDVW+TG+R+ +EEG +
Sbjct: 344 HPSLGCFVTHCGWNSTLESISFGVPMVAFPQWFDQGTNAKLMEDVWRTGVRVRANEEGSV 403
Query: 418 VKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIACI 475
V +EIR+C+E VM GEK +LR +A KWKDLAR A++E GSS NL+ +L+++ I
Sbjct: 404 VDGDEIRRCIEEVMDGGEKSRKLRESAGKWKDLARKAMEEDGSSVNNLKVFLDEVVGI 461
>dbj|BAA36421.1| UDP-glucose:anthocysnin 5-O-glucosyltransferase [Perilla frutescens
var. crispa]
Length = 460
Score = 434 bits (1117), Expect = e-120
Identities = 234/471 (49%), Positives = 313/471 (65%), Gaps = 24/471 (5%)
Query: 8 LIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTI-----PGLSFATFS 62
L+ T+P QGHINPALQF KRL+ G VTF T+++ + R+ N + PGL F FS
Sbjct: 7 LLATFPAQGHINPALQFAKRLLKAGTDVTFFTSVYAWRRMANTASAAAGNPPGLDFVAFS 66
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DGYDDG K GD YMSE RGSE L N++L+ NH T ++Y+ + +WA +VA
Sbjct: 67 DGYDDGLKPCGDGK--RYMSEMKARGSEALRNLLLN----NHDVTFVVYSHLFAWAAEVA 120
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
E +PS LLW++ ATV I+Y+YF+ + D I + DE L LP L+ R LP+F
Sbjct: 121 RESQVPSALLWVEPATVLCIYYFYFNGYADEI-DAGSDEIQLPRLP----PLEQRSLPTF 175
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
LL F L +KE+++ L+ E +VLVNT + E DAL +D + ++I IGPLIP
Sbjct: 176 LLPETPERFRL-MMKEKLETLDGEEKAKVLVNTFDALEPDALTAID--RYELIGIGPLIP 232
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
SAFLDG DP++ S+GGD+ +++ ++WLD+K + SVVYVSFG++ K QMEEI +
Sbjct: 233 SAFLDGGDPSETSYGGDLFEKSEENNCVEWLDTKPKSSVVYVSFGSVLRFPKAQMEEIGK 292
Query: 303 ALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSL 362
LL G FLW+IR++K +EEE +ELSC EL+ GKIV WCSQ+EVL+H +L
Sbjct: 293 GLLACGRPFLWMIREQKNDDGEEEE---EELSCIGELKKM--GKIVSWCSQLEVLAHPAL 347
Query: 363 GCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEE 422
GCF+THCGWNS +ESL GVP+VA PQW DQTTNAKLIED W TG+R+ +E G V E
Sbjct: 348 GCFVTHCGWNSAVESLSCGVPVVAVPQWFDQTTNAKLIEDAWGTGVRVRMNEGGGVDGSE 407
Query: 423 IRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIA 473
I +C+E+VM GEK + +R NA KWK LAR A+ E GSS +NL ++L+ +A
Sbjct: 408 IERCVEMVMDGGEKSKLVRENAIKWKTLAREAMGEDGSSLKNLNAFLHQVA 458
>dbj|BAA36422.1| UDP-glucose:anthocyanin 5-O-glucosyltransferase homologue [Perilla
frutescens var. crispa]
Length = 443
Score = 412 bits (1060), Expect = e-114
Identities = 222/453 (49%), Positives = 304/453 (67%), Gaps = 22/453 (4%)
Query: 8 LIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTI-----PGLSFATFS 62
L+ T+P QGHINPALQF KRL+ G VTF T+++ + R+ N + PGL F FS
Sbjct: 7 LLATFPAQGHINPALQFAKRLLKAGTDVTFFTSVYAWRRMANTASAAAGNPPGLDFVAFS 66
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DGYDDG K GD YMSE RGSE L N++L+ N T ++Y+ + +WA +VA
Sbjct: 67 DGYDDGLKPGGDGK--RYMSEMKARGSEALRNLLLN----NDDVTFVVYSHLFAWAAEVA 120
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
H+P+ LLW++ ATV I+++YF+ + D I S + I LP L SL+ R LP+F
Sbjct: 121 RLSHVPTALLWVEPATVLCIYHFYFNGYADEIDAGSNE----IQLPRLP-SLEQRSLPTF 175
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
LL + F L +KE+++ L+ E +VLVNT + E DAL +D + ++I IGPLIP
Sbjct: 176 LLPATPERFRL-MMKEKLETLDGEEKAKVLVNTFDALEPDALTAID--RYELIGIGPLIP 232
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
SAFLDG+DP++ S+GGD+ +++ ++WL+SK + SVVYVSFG++ K QMEEI +
Sbjct: 233 SAFLDGEDPSETSYGGDLFEKSEENNCVEWLNSKPKSSVVYVSFGSVLRFPKAQMEEIGK 292
Query: 303 ALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRSL 362
LL G FLW+IR++K +EEE +++ELSC EL+ GKIV WCSQ+EVL+H +L
Sbjct: 293 GLLACGRPFLWMIREQKNDDGEEEE-EEEELSCIGELKKM--GKIVSWCSQLEVLAHPAL 349
Query: 363 GCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVEE 422
GCF+THCGWNS +ESL G+P+VA PQW DQTTNAKLIED W TG+R+ +E G V E
Sbjct: 350 GCFVTHCGWNSAVESLSCGIPVVAVPQWFDQTTNAKLIEDAWGTGVRVRMNEGGGVDGCE 409
Query: 423 IRKCLEVVMGKGEKGEELRRNAKKWKDLARAAV 455
I +C+E+VM G+K + +R NA KWK LAR A+
Sbjct: 410 IERCVEMVMDGGDKTKLVRENAIKWKTLARQAM 442
>emb|CAB10333.1| glucosyltransferase like protein [Arabidopsis thaliana]
gi|7268302|emb|CAB78597.1| glucosyltransferase like
protein [Arabidopsis thaliana] gi|7433892|pir||C71420
hypothetical protein - Arabidopsis thaliana
Length = 458
Score = 411 bits (1057), Expect = e-113
Identities = 225/482 (46%), Positives = 318/482 (65%), Gaps = 54/482 (11%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISL--GAKVTFATTIHLYSR-LINKPTIPG-LSFATF 61
HFL +T+P QGHINP+L+ KRL GA+VTFA +I Y+R + + +P L FAT+
Sbjct: 13 HFLFVTFPAQGHINPSLELAKRLAGTISGARVTFAASISAYNRRMFSTENVPETLIFATY 72
Query: 62 SDGYDDGQKSFGDED------IVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLIL 115
SDG+DDG KS D ++MSE RRG E LT +I ++++N PFTC++YT++L
Sbjct: 73 SDGHDDGFKSSAYSDKSRQDATGNFMSEMRRRGKETLTELIEDNRKQNRPFTCVVYTILL 132
Query: 116 SWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLK 175
+W ++A +F IFY+YF+ + D I+ + + I LP L L
Sbjct: 133 TWVAELA----------------LFSIFYHYFNGYEDAISEMANTPSSSIKLPSLPL-LT 175
Query: 176 SRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMI 235
RD+PSF+++SN Y F LP+ +EQI L EEINP++L+NT +E E +A++ V K++
Sbjct: 176 VRDIPSFIVSSNVYAFLLPAFREQIDSLKEEINPKILINTFQELEPEAMSSVP-DNFKIV 234
Query: 236 PIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKR 295
P+GPL+ TD S S+ +YI+WLD+K + SV+YVSFGTLAVLSK+
Sbjct: 235 PVGPLLTLR-------TDFS---------SRGEYIEWLDTKADSSVLYVSFGTLAVLSKK 278
Query: 296 QMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDEL--SCREELENNMNGKIVKWCSQ 353
Q+ E+ +AL+ S FLWVI DK + +++E+ +++ S REEL+ G +V WC Q
Sbjct: 279 QLVELCKALIQSRRPFLWVITDKSYRNKEDEQEKEEDCISSFREELDEI--GMVVSWCDQ 336
Query: 354 VEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRM--E 411
VL+HRS+GCF+THCGWNSTLESL SGVP+VAFPQW DQ NAKL+ED WKTG+R+ +
Sbjct: 337 FRVLNHRSIGCFVTHCGWNSTLESLVSGVPVVAFPQWNDQMMNAKLLEDCWKTGVRVMEK 396
Query: 412 HDEEGMVKV--EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYL 469
+EEG+V V EEIR+C+E VM +K EE R NA +WKDLA AV+EGGSS +L++++
Sbjct: 397 KEEEGVVVVDSEEIRRCIEEVM--EDKAEEFRGNATRWKDLAAEAVREGGSSFNHLKAFV 454
Query: 470 ND 471
++
Sbjct: 455 DE 456
>gb|AAF79730.1| T25N20.21 [Arabidopsis thaliana] gi|18700284|gb|AAL77752.1|
At1g05560/T25N20_20 [Arabidopsis thaliana]
gi|13605918|gb|AAK32944.1| At1g05560/T25N20_20
[Arabidopsis thaliana] gi|18390540|ref|NP_563742.1|
UDP-glucose transferase (UGT75B2) [Arabidopsis thaliana]
gi|13661275|gb|AAK37839.1| UDP-glucosyltransferase
[Arabidopsis thaliana]
Length = 469
Score = 411 bits (1057), Expect = e-113
Identities = 228/474 (48%), Positives = 308/474 (64%), Gaps = 34/474 (7%)
Query: 6 HFLIITYPLQGHINPALQFTKRLIS-LGAKVTFATTIHLY--SRLINKPTIPGLSFATFS 62
HFL++T+P QGH+NP+L+F +RLI GA+VTF T + ++ S + N + LSF TFS
Sbjct: 5 HFLLVTFPAQGHVNPSLRFARRLIKRTGARVTFVTCVSVFHNSMIANHNKVENLSFLTFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DG+DDG S ED G + L++ I ++K + P TCLIYT++L+WAPKVA
Sbjct: 65 DGFDDGGISTY-EDRQKRSVNLKVNGDKALSDFIEATKNGDSPVTCLIYTILLNWAPKVA 123
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
LPS LLWIQ A VF+I+Y +F + NKS + LP LS SL+ RDLPSF
Sbjct: 124 RRFQLPSALLWIQPALVFNIYYTHF------MGNKS-----VFELPNLS-SLEIRDLPSF 171
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L SNT A + +E ++ L +E P++L+NT + E +AL I M+ +GPL+P
Sbjct: 172 LTPSNTNKGAYDAFQEMMEFLIKETKPKILINTFDSLEPEALTAFP--NIDMVAVGPLLP 229
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
+ G T+ S D Y WLDSK E SV+YVSFGT+ LSK+Q+EE+AR
Sbjct: 230 TEIFSGS--TNKSVK------DQSSSYTLWLDSKTESSVIYVSFGTMVELSKKQIEELAR 281
Query: 303 ALLDSGFSFLWVIRDKKLQQQKEEEVDDDELS----CREELENNMNGKIVKWCSQVEVLS 358
AL++ FLWVI DK ++ K E ++ E+ R ELE G IV WCSQ+EVLS
Sbjct: 282 ALIEGKRPFLWVITDKSNRETKTEGEEETEIEKIAGFRHELEEV--GMIVSWCSQIEVLS 339
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMV 418
HR++GCF+THCGW+STLESL GVP+VAFP W+DQ TNAKL+E+ WKTG+R+ +++G+V
Sbjct: 340 HRAVGCFVTHCGWSSTLESLVLGVPVVAFPMWSDQPTNAKLLEESWKTGVRVRENKDGLV 399
Query: 419 KVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+ EIR+CLE VM EK ELR NAKKWK LA A +EGGSS++N+ +++ DI
Sbjct: 400 ERGEIRRCLEAVM--EEKSVELRENAKKWKRLAMEAGREGGSSDKNMEAFVEDI 451
>gb|AAF79732.1| T25N20.18 [Arabidopsis thaliana] gi|15220556|ref|NP_172044.1|
UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana]
Length = 455
Score = 405 bits (1042), Expect = e-111
Identities = 220/474 (46%), Positives = 305/474 (63%), Gaps = 31/474 (6%)
Query: 6 HFLIITYPLQGHINPALQFTKRLI-SLGAKVTFATTIHLYSRLI--NKPTIPGLSFATFS 62
HFL++T+P QGH+NP+L+F +RLI + GA+VTFAT + + R + N + LSF TFS
Sbjct: 5 HFLLVTFPAQGHVNPSLRFARRLIKTTGARVTFATCLSVIHRSMIPNHNNVENLSFLTFS 64
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DG+DDG S D D+ + + F R G + L++ I +++ + P +CLIYT++ +W PKVA
Sbjct: 65 DGFDDGVISNTD-DVQNRLVHFERNGDKALSDFIEANQNGDSPVSCLIYTILPNWVPKVA 123
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
HLPS LWIQ A FDI+Y Y S + P L SL+ RDLPSF
Sbjct: 124 RRFHLPSVHLWIQPAFAFDIYYNY-----------STGNNSVFEFPNLP-SLEIRDLPSF 171
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L SNT A +E + L EE NP++LVNT + E + L + I+M+ +GPL+P
Sbjct: 172 LSPSNTNKAAQAVYQELMDFLKEESNPKILVNTFDSLEPEFLTAIP--NIEMVAVGPLLP 229
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIAR 302
+ G + G D+ R Y WLDSK E SV+YVSFGT+ LSK+Q+EE+AR
Sbjct: 230 AEIFTGSES-----GKDLSRDHQSSSYTLWLDSKTESSVIYVSFGTMVELSKKQIEELAR 284
Query: 303 ALLDSGFSFLWVIRDKKLQQQK---EEEVDDDELS-CREELENNMNGKIVKWCSQVEVLS 358
AL++ G FLWVI DK ++ K EEE + ++++ R ELE G IV WCSQ+EVL
Sbjct: 285 ALIEGGRPFLWVITDKLNREAKIEGEEETEIEKIAGFRHELEEV--GMIVSWCSQIEVLR 342
Query: 359 HRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMV 418
HR++GCF+THCGW+S+LESL GVP+VAFP W+DQ NAKL+E++WKTG+R+ + EG+V
Sbjct: 343 HRAIGCFLTHCGWSSSLESLVLGVPVVAFPMWSDQPANAKLLEEIWKTGVRVRENSEGLV 402
Query: 419 KVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDI 472
+ EI +CLE VM K ELR NA+KWK LA A +EGGSS++N+ +++ +
Sbjct: 403 ERGEIMRCLEAVM--EAKSVELRENAEKWKRLATEAGREGGSSDKNVEAFVKSL 454
>emb|CAB10189.1| glucosyltransferase like protein [Arabidopsis thaliana]
gi|7268115|emb|CAB78452.1| glucosyltransferase like
protein [Arabidopsis thaliana]
gi|22136734|gb|AAM91686.1| putative glucosyltransferase
[Arabidopsis thaliana] gi|21387139|gb|AAM47973.1|
glucosyltransferase [Arabidopsis thaliana]
gi|17065026|gb|AAL32667.1| glucosyltransferase
[Arabidopsis thaliana] gi|15236407|ref|NP_193146.1|
UDP-glucoronosyl/UDP-glucosyl transferase family protein
[Arabidopsis thaliana] gi|7433897|pir||C71402 probable
glucosyltransferase - Arabidopsis thaliana
Length = 456
Score = 402 bits (1033), Expect = e-110
Identities = 219/470 (46%), Positives = 301/470 (63%), Gaps = 32/470 (6%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H+L++T+P QGHINPALQ RLI GA VT++T + + R+ P+ GLSFA F+DG+
Sbjct: 13 HYLLVTFPAQGHINPALQLANRLIHHGATVTYSTAVSAHRRMGEPPSTKGLSFAWFTDGF 72
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNII---LSSKQENHPFTCLIYTLILSWAPKVA 122
DDG KSF D+ I YMSE R GS L +II L + E P T +IY++++ W VA
Sbjct: 73 DDGLKSFEDQKI--YMSELKRCGSNALRDIIKANLDATTETEPITGVIYSVLVPWVSTVA 130
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
E HLP+TLLWI+ ATV DI+YYYF+ ++ + I LP L + + DLPSF
Sbjct: 131 REFHLPTTLLWIEPATVLDIYYYYFNTSYKHLFDVEP-----IKLPKLPL-ITTGDLPSF 184
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L S AL +L+E I+ L E NP++LVNT E DAL V+ K+KMIPIGPL+
Sbjct: 185 LQPSKALPSALVTLREHIEALETESNPKILVNTFSALEHDALTSVE--KLKMIPIGPLVS 242
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAV-LSKRQMEEIA 301
S+ +GK S S +DY +WLDSK E+SV+Y+S GT A L ++ ME +
Sbjct: 243 SS--EGKTDLFKS---------SDEDYTKWLDSKLERSVIYISLGTHADDLPEKHMEALT 291
Query: 302 RALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRS 361
+L + FLW++R+K +++K+ E + + G +V WCSQ VL+H +
Sbjct: 292 HGVLATNRPFLWIVREKNPEEKKKNRF-------LELIRGSDRGLVVGWCSQTAVLAHCA 344
Query: 362 LGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVE 421
+GCF+THCGWNSTLESL SGVP+VAFPQ+ DQ T AKL+ED W+ G++++ EEG V E
Sbjct: 345 VGCFVTHCGWNSTLESLESGVPVVAFPQFADQCTTAKLVEDTWRIGVKVKVGEEGDVDGE 404
Query: 422 EIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
EIR+CLE VM GE+ EE+R NA+KWK +A A EGG S+ NL+ ++++
Sbjct: 405 EIRRCLEKVMSGGEEAEEMRENAEKWKAMAVDAAAEGGPSDLNLKGFVDE 454
>gb|AAL69494.1| putative glucosyltransferase [Arabidopsis thaliana]
Length = 466
Score = 402 bits (1033), Expect = e-110
Identities = 219/470 (46%), Positives = 301/470 (63%), Gaps = 32/470 (6%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H+L++T+P QGHINPALQ RLI GA VT++T + + R+ P+ GLSFA F+DG+
Sbjct: 23 HYLLVTFPAQGHINPALQLANRLIHHGATVTYSTAVSAHRRMGEPPSTKGLSFAWFTDGF 82
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNII---LSSKQENHPFTCLIYTLILSWAPKVA 122
DDG KSF D+ I YMSE R GS L +II L + E P T +IY++++ W VA
Sbjct: 83 DDGLKSFEDQKI--YMSELKRCGSNALRDIIKANLDATTETEPITGVIYSVLVPWVSTVA 140
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
E HLP+TLLWI+ ATV DI+YYYF+ ++ + I LP L + + DLPSF
Sbjct: 141 REFHLPTTLLWIEPATVLDIYYYYFNTSYKHLFDVEP-----IKLPKLPL-ITTGDLPSF 194
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIPIGPLIP 242
L S AL +L+E I+ L E NP++LVNT E DAL V+ K+KMIPIGPL+
Sbjct: 195 LQPSKALPSALVTLREHIEALETESNPKILVNTFSALEHDALTSVE--KLKMIPIGPLVS 252
Query: 243 SAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAV-LSKRQMEEIA 301
S+ +GK S S +DY +WLDSK E+SV+Y+S GT A L ++ ME +
Sbjct: 253 SS--EGKTDLFKS---------SDEDYTKWLDSKLERSVIYISLGTHADDLPEKHMEALT 301
Query: 302 RALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRS 361
+L + FLW++R+K +++K+ E + + G +V WCSQ VL+H +
Sbjct: 302 HGVLATNRPFLWIVREKNPEEKKKNRF-------LELIRGSDRGLVVGWCSQTAVLAHCA 354
Query: 362 LGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVE 421
+GCF+THCGWNSTLESL SGVP+VAFPQ+ DQ T AKL+ED W+ G++++ EEG V E
Sbjct: 355 VGCFVTHCGWNSTLESLESGVPVVAFPQFADQCTTAKLVEDTWRIGVKVKVGEEGDVDGE 414
Query: 422 EIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLND 471
EIR+CLE VM GE+ EE+R NA+KWK +A A EGG S+ NL+ ++++
Sbjct: 415 EIRRCLEKVMSGGEEAEEMRENAEKWKAMAVDAAAEGGPSDLNLKGFVDE 464
>gb|AAM65349.1| AT4g15550/dl3815c [Arabidopsis thaliana] gi|16604440|gb|AAL24226.1|
AT4g15550/dl3815c [Arabidopsis thaliana]
Length = 418
Score = 400 bits (1028), Expect = e-110
Identities = 210/428 (49%), Positives = 293/428 (68%), Gaps = 34/428 (7%)
Query: 56 LSFATFSDGYDDGQKSFGDED------IVSYMSEFTRRGSEFLTNIILSSKQENHPFTCL 109
L FAT+SDG+DD KS D ++MSE RRG E LT +I ++++N PFTC+
Sbjct: 11 LIFATYSDGHDDVFKSSAYSDKSRQDATGNFMSEMRRRGKETLTELIEDNRKQNRPFTCV 70
Query: 110 IYTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPG 169
+YT++L+W ++A E HLPS LLW+Q TVF IFY+YF+ + D I+ + + I LP
Sbjct: 71 VYTILLTWVAELAREFHLPSALLWVQPVTVFSIFYHYFNGYEDAISEMANTPSSSIKLPS 130
Query: 170 LSFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDV 229
L L RD+PSF+++SN Y F LP+ +EQI L EEINP++L+NT +E E +A++ V
Sbjct: 131 LPL-LTVRDIPSFIVSSNVYAFLLPAFREQIDSLKEEINPKILINTFQELEPEAMSSVP- 188
Query: 230 GKIKMIPIGPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTL 289
K++P+GPL+ TD S S+ +YI+WLD+K + SV+YVSFGTL
Sbjct: 189 DNFKIVPVGPLLTLR-------TDFS---------SRGEYIEWLDTKADSSVLYVSFGTL 232
Query: 290 AVLSKRQMEEIARALLDSGFSFLWVIRDKKLQQQKEEEVDDDEL--SCREELENNMNGKI 347
AVLSK+Q+ E+ +AL+ S FLWVI DK + +++E+ +++ S REEL+ G +
Sbjct: 233 AVLSKKQLVELCKALIQSRRPFLWVITDKSYRNKEDEQEKEEDCISSFREELDEI--GMV 290
Query: 348 VKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTG 407
V WC Q VL+HRS+GCF+THCGWNSTLESL SGVP+VAFPQW DQ NAKL+ED WKTG
Sbjct: 291 VSWCDQFRVLNHRSIGCFVTHCGWNSTLESLVSGVPVVAFPQWNDQMMNAKLLEDCWKTG 350
Query: 408 LRM--EHDEEGMVKV--EEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNR 463
+R+ + +EEG+V V EEIR+C+E VM +K EE R NA +WKDLA AV+EGGSS
Sbjct: 351 VRVMEKKEEEGVVVVDSEEIRRCIEEVM--EDKAEEFRGNATRWKDLAAEAVREGGSSFN 408
Query: 464 NLRSYLND 471
+L++++++
Sbjct: 409 HLKAFVDE 416
>dbj|BAC54093.1| anthocyanin 5-glucosyltransferase [Torenia hybrida]
Length = 478
Score = 363 bits (933), Expect = 5e-99
Identities = 212/488 (43%), Positives = 309/488 (62%), Gaps = 32/488 (6%)
Query: 1 MAQNHHFLIITYPLQGHINPALQFTKRLISLGA--KVTFATTIHLYSRLINKPTIPG--L 56
M H L+ T+P QGHINP+L+F KRL++ G +VTF T+++ R+ T P +
Sbjct: 1 MVNKRHILLATFPAQGHINPSLEFAKRLLNTGYVDQVTFFTSVYALRRM-RFETDPSSRI 59
Query: 57 SFATFSDGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTC------LI 110
F D YDDG K +D +YMSE +RG++ L + ++ C ++
Sbjct: 60 DFVAXXDSYDDGLKK--GDDGKNYMSEMRKRGTKALKDTLIKLNDAAMGSECYNRVSFVV 117
Query: 111 YTLILSWAPKVAHELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGL 170
Y+ + SWA +VA E+ +PS LLWI+ ATVFD++Y+YF+ + D I + D+ L +LP L
Sbjct: 118 YSHLFSWAAEVAREVDVPSALLWIEPATVFDVYYFYFNGYADDI-DAGSDQIQLPNLPQL 176
Query: 171 SFSLKSRDLPSFLLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVG 230
S +DLPSFLL S+ F +KE+ L++E +VL+NT + E + L +D
Sbjct: 177 S----KQDLPSFLLPSSPARFRT-LMKEKFDTLDKEPKAKVLINTFDALETEQLKAID-- 229
Query: 231 KIKMIPIGPLIPSA-FLDGKDPTDN--SFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFG 287
+ ++I IGPLIPS+ F DG DP+ + S+GGD+ R + + Y+ WL+SK E SVVYVSFG
Sbjct: 230 RYELISIGPLIPSSIFSDGNDPSSSNKSYGGDLFR-KADETYMDWLNSKPESSVVYVSFG 288
Query: 288 TLAVLSKRQMEEIARALLDSGFSFLWVIR-DKKLQQQKEEEVDDDELSCREELENNMNGK 346
+L L K QMEEIA L D+ LWVIR +++ +Q++ E ++ LS + GK
Sbjct: 289 SLLRLPKPQMEEIAIGLSDTKSPVLWVIRRNEEGDEQEQAEEEEKLLSFFDRHGTERLGK 348
Query: 347 IVKWCSQVEVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKT 406
IV WCSQ++VL+H+S+GCF+THCGWNS +ESL GVP+V FPQW DQ TNAK+IEDVW++
Sbjct: 349 IVTWCSQLDVLTHKSVGCFVTHCGWNSAIESLACGVPVVCFPQWFDQGTNAKMIEDVWRS 408
Query: 407 GLRMEHDEE-GMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAV-KEGGSSNRN 464
G+R+ +EE G+V EI++C+ V+ K ELR +A WK LA+ A+ +E GSS N
Sbjct: 409 GVRVRVNEEGGVVDRREIKRCVSEVI----KSRELRESAMMWKGLAKEAMDEERGSSMNN 464
Query: 465 LRSYLNDI 472
L++++ I
Sbjct: 465 LKNFITRI 472
>gb|AAF61647.1| UDP-glucose:salicylic acid glucosyltransferase [Nicotiana tabacum]
Length = 459
Score = 326 bits (835), Expect = 1e-87
Identities = 191/477 (40%), Positives = 284/477 (59%), Gaps = 24/477 (5%)
Query: 3 QNHHFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFS 62
Q H LI+ YP QGHINP LQF+KRL S G K+T A T + T +S S
Sbjct: 4 QKAHCLILPYPAQGHINPMLQFSKRLQSKGVKITIAATKSFLKTMQELST--SVSVEAIS 61
Query: 63 DGYDDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVA 122
DGYDDG + V+Y++ F GS+ L+ +I P +C++Y L WA +V
Sbjct: 62 DGYDDGGREQAGT-FVAYITRFKEVGSDTLSQLIGKLTNCGCPVSCIVYDPFLPWAVEVG 120
Query: 123 HELHLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSF 182
+ + + + Q+ V +I+Y H H + D IS+PGL ++++ D+PSF
Sbjct: 121 NNFGVATAAFFTQSCAVDNIYY---HVHKGVLKLPPTDVDKEISIPGL-LTIEASDVPSF 176
Query: 183 LLASNTYTFALPSLKEQIQLLNEEINPRVLVNTVEEFELDALNKVDVGKIKMIP-IGPLI 241
+ SN + + + Q N E VL+N+ E E + ++ + KI I IGP I
Sbjct: 177 V--SNPESSRILEMLVN-QFSNLENTDWVLINSFYELEKEVIDWM--AKIYPIKTIGPTI 231
Query: 242 PSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQMEEIA 301
PS +LD + P D +G V + + + WL+ + SVVYVSFG+LA L QMEE+A
Sbjct: 232 PSMYLDKRLPDDKEYGLSVFK-PMTNACLNWLNHQPVSSVVYVSFGSLAKLEAEQMEELA 290
Query: 302 RALLDSGFSFLWVIRDKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQVEVLSHRS 361
L +S +FLWV+R + E ++ ++ L EEL + G +V WC Q++VL H+S
Sbjct: 291 WGLSNSNKNFLWVVRSTE-----ESKLPNNFL---EELASE-KGLVVSWCPQLQVLEHKS 341
Query: 362 LGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDEEGMVKVE 421
+GCF+THCGWNSTLE++ GVPM+A P W+DQ TNAKL+EDVW+ G+R + DE+G+V+ E
Sbjct: 342 IGCFLTHCGWNSTLEAISLGVPMIAMPHWSDQPTNAKLVEDVWEMGIRPKQDEKGLVRRE 401
Query: 422 EIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIACITQI 478
I +C+++VM + +KG+++R NAKKWK+LAR AV EGGSS+RN+ +++ + I +
Sbjct: 402 VIEECIKIVM-EEKKGKKIRENAKKWKELARKAVDEGGSSDRNIEEFVSKLVTIASV 457
>emb|CAI62049.1| UDP-xylose phenolic glycosyltransferase [Lycopersicon esculentum]
Length = 456
Score = 319 bits (817), Expect = 1e-85
Identities = 185/482 (38%), Positives = 278/482 (57%), Gaps = 43/482 (8%)
Query: 6 HFLIITYPLQGHINPALQFTKRLISLGAKVTFATTIHLYSRLINKPTIPGLSFATFSDGY 65
H LI+ YP+QGHINP LQF+KRL S K+T A T + PT +S SDGY
Sbjct: 7 HCLILPYPVQGHINPMLQFSKRLRSKRVKITIALTKSFLKNMKELPT--SMSIEAISDGY 64
Query: 66 DDGQKSFGDEDIVSYMSEFTRRGSEFLTNIILSSKQENHPFTCLIYTLILSWAPKVAHEL 125
DDG + V+Y++ F GS+ L+ +I P C++Y L WA +VA +
Sbjct: 65 DDGGRDQAGT-FVAYITRFKEIGSDTLSQLIQKLAISGCPVNCIVYDPFLPWAVEVAKQF 123
Query: 126 HLPSTLLWIQAATVFDIFYYYFHEHGDYITNKSKDETCLISLPGLSFSLKSRDLPSFLLA 185
L S + Q V D YY+ H+ + DE LI PG S+ + D+PSF+++
Sbjct: 124 GLISAAFFTQNCVV-DNLYYHVHKGVIKLPPTQNDEEILI--PGFPNSIDASDVPSFVIS 180
Query: 186 SNTYTFALPSLKEQIQLLNEEINPR-----VLVNTVEEFELDALNKVDVGKIKMIPI--- 237
P + +++L + + VL+N+ E E + ++ + K+ PI
Sbjct: 181 --------PEAERIVEMLANQFSNLDKVDCVLINSFYELEKEVIDWMS----KIYPIKTI 228
Query: 238 GPLIPSAFLDGKDPTDNSFGGDVVRVDSKDDYIQWLDSKDEKSVVYVSFGTLAVLSKRQM 297
GP IPS +LD + D +G + + ++ + WL+ + SV+YVSFG+LA L QM
Sbjct: 229 GPTIPSMYLDKRLHDDKEYGLSMFK-PMTNECLNWLNHQPISSVLYVSFGSLAKLGSEQM 287
Query: 298 EEIARALLDSGFSFLWVIR---DKKLQQQKEEEVDDDELSCREELENNMNGKIVKWCSQV 354
EE+A L +S SFLWV+R + KL EE+ ++ G +V WC Q+
Sbjct: 288 EELAWGLKNSNKSFLWVVRSTEEPKLPNNFIEELTSEK------------GLVVSWCPQL 335
Query: 355 EVLSHRSLGCFMTHCGWNSTLESLGSGVPMVAFPQWTDQTTNAKLIEDVWKTGLRMEHDE 414
+VL H S+GCF+THCGWNSTLE++ GVPMVA PQW+DQ TNAKL++DVW+ G+R + DE
Sbjct: 336 QVLEHESIGCFLTHCGWNSTLEAISLGVPMVAMPQWSDQPTNAKLVKDVWEIGVRAKQDE 395
Query: 415 EGMVKVEEIRKCLEVVMGKGEKGEELRRNAKKWKDLARAAVKEGGSSNRNLRSYLNDIAC 474
+G+V+ E I +C+++VM + +KG+ +R NAKKWK++AR V EGGSS++N+ +++ +
Sbjct: 396 KGVVRREVIEECIKLVM-EEDKGKLIRENAKKWKEIARNVVNEGGSSDKNIEEFVSKLVT 454
Query: 475 IT 476
I+
Sbjct: 455 IS 456
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 837,928,234
Number of Sequences: 2540612
Number of extensions: 36244891
Number of successful extensions: 104666
Number of sequences better than 10.0: 1286
Number of HSP's better than 10.0 without gapping: 1205
Number of HSP's successfully gapped in prelim test: 81
Number of HSP's that attempted gapping in prelim test: 101502
Number of HSP's gapped (non-prelim): 1604
length of query: 478
length of database: 863,360,394
effective HSP length: 132
effective length of query: 346
effective length of database: 527,999,610
effective search space: 182687865060
effective search space used: 182687865060
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146706.4


