

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
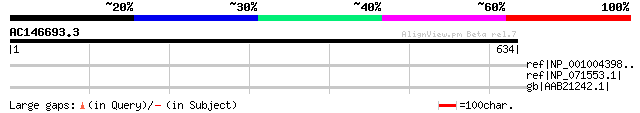
Query= AC146693.3 - phase: 0 /pseudo
(634 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_001004398.1| Amphiphysin [Gallus gallus] gi|62843|emb|CAA... 36 3.3
ref|NP_071553.1| amphiphysin 1 [Rattus norvegicus] gi|2145467|em... 35 9.7
gb|AAB21242.1| uroporphyrinogen III synthase, UROIIIS [human, Pe... 35 9.7
>ref|NP_001004398.1| Amphiphysin [Gallus gallus] gi|62843|emb|CAA42953.1| Amphiphysin
[Gallus gallus] gi|86184|pir||S22700 amphiphysin -
chicken gi|1703280|sp|P50478|AMPH_CHICK Amphiphysin
Length = 682
Score = 36.2 bits (82), Expect = 3.3
Identities = 21/55 (38%), Positives = 27/55 (48%), Gaps = 7/55 (12%)
Query: 318 PNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTP 372
P +S SP + P +L+P +P RPKSP K PP+ P PK TP
Sbjct: 264 PEEVSPLPSPTASPNHMLAPASPAPARPKSPTQLRKGPPV-------PPLPKLTP 311
>ref|NP_071553.1| amphiphysin 1 [Rattus norvegicus] gi|2145467|emb|CAA73808.1|
amphiphysin [Rattus norvegicus]
gi|14916529|sp|O08838|AMPH_RAT Amphiphysin
Length = 683
Score = 34.7 bits (78), Expect = 9.7
Identities = 20/55 (36%), Positives = 26/55 (46%), Gaps = 7/55 (12%)
Query: 318 PNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTP 372
P S SP + P L+P +P +RP+SP K PP+ P PK TP
Sbjct: 264 PEEASPLPSPTASPNHTLAPASPAPVRPRSPSQTRKGPPV-------PPLPKVTP 311
>gb|AAB21242.1| uroporphyrinogen III synthase, UROIIIS [human, Peptide Mutant, 265
aa]
Length = 265
Score = 34.7 bits (78), Expect = 9.7
Identities = 15/33 (45%), Positives = 20/33 (60%)
Query: 324 RLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPP 356
RL NS P++I P PK L+P +P +WPP
Sbjct: 229 RLISNSWPQVIFPPWPPKLLQPSAPLRLARWPP 261
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.364 0.164 0.631
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 882,245,688
Number of Sequences: 2540612
Number of extensions: 31221747
Number of successful extensions: 145482
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 145481
Number of HSP's gapped (non-prelim): 3
length of query: 634
length of database: 863,360,394
effective HSP length: 134
effective length of query: 500
effective length of database: 522,918,386
effective search space: 261459193000
effective search space used: 261459193000
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146693.3


