

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146664.9 - phase: 0
(59 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
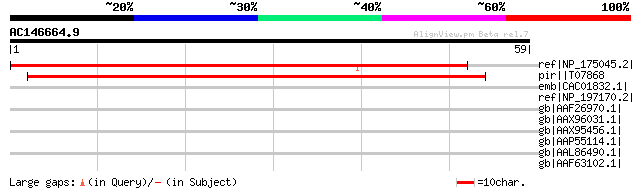
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_175045.2| PHD finger family protein [Arabidopsis thaliana] 53 2e-06
pir||T07868 DNA binding protein E4/E8BP-1 - tomato gi|2342860|gb... 46 3e-04
emb|CAC01832.1| putative protein [Arabidopsis thaliana] gi|11357... 44 0.001
ref|NP_197170.2| PHD finger family protein [Arabidopsis thaliana] 44 0.001
gb|AAF26970.1| unknown protein [Arabidopsis thaliana] gi|1523301... 40 0.013
gb|AAX96031.1| PHD-finger, putative [Oryza sativa (japonica cult... 40 0.016
gb|AAX95456.1| hypothetical protein [Oryza sativa (japonica cult... 37 0.18
gb|AAP55114.1| putative DNA-binding protein [Oryza sativa (japon... 36 0.31
gb|AAL86490.1| putative DNA-binding protein, 5'-partial [Oryza s... 36 0.31
gb|AAF63102.1| Hypothetical protein [Arabidopsis thaliana] gi|25... 34 0.90
>ref|NP_175045.2| PHD finger family protein [Arabidopsis thaliana]
Length = 371
Score = 52.8 bits (125), Expect = 2e-06
Identities = 24/53 (45%), Positives = 35/53 (65%), Gaps = 1/53 (1%)
Query: 1 MGHLSTLACPKVHEEARYLPNMISANFLQKSTVWPESF-KNSGTNNFSIGIYF 52
+ H+S+LACPKVHE A L +SA L + VWP++F KN G + S+ ++F
Sbjct: 311 VAHVSSLACPKVHETASSLKGRLSAEILPRLEVWPKTFLKNGGPKDESVALFF 363
>pir||T07868 DNA binding protein E4/E8BP-1 - tomato gi|2342860|gb|AAB67671.1|
E4/E8 binding protein-1 [Lycopersicon esculentum]
Length = 460
Score = 45.8 bits (107), Expect = 3e-04
Identities = 23/52 (44%), Positives = 32/52 (61%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLS+ AC KV A+ + +I FL K VWP+SFK++ + +I IYF S
Sbjct: 29 HLSSKACAKVWLAAKEMAEVIYPEFLPKRDVWPKSFKSAKLIDDNIAIYFFS 80
>emb|CAC01832.1| putative protein [Arabidopsis thaliana] gi|11357809|pir||T51500
hypothetical protein F5E19_20 - Arabidopsis thaliana
Length = 1280
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/52 (36%), Positives = 27/52 (51%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLSTLA P+V E P S N + + + WP F+ GT I ++F +
Sbjct: 829 HLSTLASPRVAEVVNKFPETFSLNEVPRKSTWPTQFEKLGTKEAHIALFFFA 880
>ref|NP_197170.2| PHD finger family protein [Arabidopsis thaliana]
Length = 1290
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/52 (36%), Positives = 27/52 (51%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLSTLA P+V E P S N + + + WP F+ GT I ++F +
Sbjct: 839 HLSTLASPRVAEVVNKFPETFSLNEVPRKSTWPTQFEKLGTKEAHIALFFFA 890
>gb|AAF26970.1| unknown protein [Arabidopsis thaliana] gi|15233015|ref|NP_186939.1|
PHD finger protein-related [Arabidopsis thaliana]
Length = 963
Score = 40.4 bits (93), Expect = 0.013
Identities = 16/52 (30%), Positives = 30/52 (56%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
+LSTLA PKV E + P ++ N + + + WP F+++G + ++F +
Sbjct: 692 YLSTLASPKVVEVVKQFPEKVTLNEVPRLSSWPAQFQDTGAKEQHVALFFFA 743
>gb|AAX96031.1| PHD-finger, putative [Oryza sativa (japonica cultivar-group)]
Length = 1056
Score = 40.0 bits (92), Expect = 0.016
Identities = 18/52 (34%), Positives = 28/52 (53%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLS A PKV E A+ P+ + L + +WP F ++G SI ++F +
Sbjct: 750 HLSCFASPKVLEVAKKFPSNVQLEELPRQNLWPPQFHDNGPTIDSIALFFFA 801
>gb|AAX95456.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 639
Score = 36.6 bits (83), Expect = 0.18
Identities = 21/53 (39%), Positives = 29/53 (54%), Gaps = 1/53 (1%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESF-KNSGTNNFSIGIYFLS 54
H S AC KV E + LP ++ L K WP+S+ K S + SIG++F S
Sbjct: 451 HFSNKACKKVCELSMSLPQIMKVTELPKLKAWPKSWEKASVPSAESIGLFFFS 503
>gb|AAP55114.1| putative DNA-binding protein [Oryza sativa (japonica
cultivar-group)] gi|37537050|ref|NP_922827.1| putative
DNA-binding protein [Oryza sativa (japonica
cultivar-group)]
Length = 982
Score = 35.8 bits (81), Expect = 0.31
Identities = 19/52 (36%), Positives = 26/52 (49%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLST A KV E + LP I + + + WP FK N +I +YF +
Sbjct: 499 HLSTCASLKVLEIVKQLPQRIQLVEVPRHSSWPLQFKEVKPNEDNIALYFFA 550
>gb|AAL86490.1| putative DNA-binding protein, 5'-partial [Oryza sativa (japonica
cultivar-group)]
Length = 948
Score = 35.8 bits (81), Expect = 0.31
Identities = 19/52 (36%), Positives = 26/52 (49%)
Query: 3 HLSTLACPKVHEEARYLPNMISANFLQKSTVWPESFKNSGTNNFSIGIYFLS 54
HLST A KV E + LP I + + + WP FK N +I +YF +
Sbjct: 465 HLSTCASLKVLEIVKQLPQRIQLVEVPRHSSWPLQFKEVKPNEDNIALYFFA 516
>gb|AAF63102.1| Hypothetical protein [Arabidopsis thaliana] gi|25405141|pir||E96501
hypothetical protein F28H19.3 [imported] - Arabidopsis
thaliana
Length = 419
Score = 34.3 bits (77), Expect = 0.90
Identities = 15/28 (53%), Positives = 19/28 (67%)
Query: 1 MGHLSTLACPKVHEEARYLPNMISANFL 28
+ H+S+LACPKVHE A L +SA L
Sbjct: 311 VAHVSSLACPKVHETASSLKGRLSAEIL 338
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.133 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 106,775,019
Number of Sequences: 2540612
Number of extensions: 3145561
Number of successful extensions: 5225
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 5215
Number of HSP's gapped (non-prelim): 10
length of query: 59
length of database: 863,360,394
effective HSP length: 35
effective length of query: 24
effective length of database: 774,438,974
effective search space: 18586535376
effective search space used: 18586535376
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146664.9


