

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146664.7 - phase: 0
(2305 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
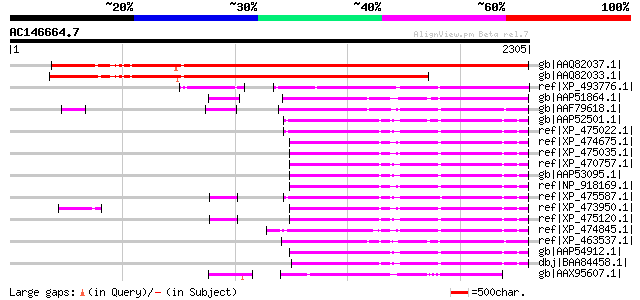
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum] 2361 0.0
gb|AAQ82033.1| gag/pol polyprotein [Pisum sativum] 1659 0.0
ref|XP_493776.1| unnamed protein product [Oryza sativa (japonica... 889 0.0
gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sati... 770 0.0
gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||... 761 0.0
gb|AAP52501.1| putative gag-pol precursor [Oryza sativa (japonic... 708 0.0
ref|XP_475022.1| OSJNBb0093G06.3 [Oryza sativa (japonica cultiva... 708 0.0
ref|XP_474675.1| OSJNBa0027O01.4 [Oryza sativa (japonica cultiva... 707 0.0
ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultiva... 705 0.0
ref|XP_470757.1| putative gag-pol precursor [Oryza sativa] gi|18... 705 0.0
gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cul... 703 0.0
ref|NP_918169.1| putative gag-pol polyprotein [Oryza sativa (jap... 703 0.0
ref|XP_475587.1| putative polyprotein [Oryza sativa (japonica cu... 702 0.0
ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultiv... 701 0.0
ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cu... 700 0.0
ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultiva... 697 0.0
ref|XP_463537.1| putative polyprotein [Oryza sativa (japonica cu... 696 0.0
gb|AAP54912.1| gag-pol precursor [Oryza sativa (japonica cultiva... 694 0.0
dbj|BAA84458.1| GAG-POL precursor [Oryza sativa (japonica cultiv... 691 0.0
gb|AAX95607.1| RNase H, putative [Oryza sativa (japonica cultiva... 688 0.0
>gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 2361 bits (6119), Expect = 0.0
Identities = 1226/2194 (55%), Positives = 1567/2194 (70%), Gaps = 137/2194 (6%)
Query: 186 EERFGALEKKMSNMRGKETVVQSIYDLCLVPDVNIPPKFKMPVFEKYQGDTCPQNHLTMY 245
+ + AL +K+ M + ++ + ++ LV + IP KFK P F+KY G +CP+ H+ Y
Sbjct: 130 DRKVDALAEKIRAMECQNSLGFDVTNMGLVEGLRIPYKFKAPSFDKYNGTSCPRTHVQAY 189
Query: 246 IRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMGLKG--VTTFEQLAEAFMQQYKYNTYLAP 303
R++ AY ++ + ++ FQDSL+G + W+M LK + + L EAF++QYK+N +AP
Sbjct: 190 YRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKSDSIRCWRDLGEAFLRQYKHNMDMAP 249
Query: 304 SRKELQSLTQKDKESFKEYAQRFIQKAAQIRPPLDERELSELFYETLSPCYSEKMIVCAS 363
SR +LQSL QK ESFKEYAQR+ + AA+++PP+ EREL+++F TL + ++M C
Sbjct: 250 SRTQLQSLCQKSNESFKEYAQRWRELAARVQPPMLERELTDMFIGTLQGVFMDRMGSCPF 309
Query: 364 QKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGVSSNGNKKFGNGYPKRNAQEVGMVAHG 423
F+D+V G R E + +G V S+ +KK G P+R E V H
Sbjct: 310 GSFSDVVICGERTESLIK---------TGKIQDVGSSSSKKPFAGAPRRREGETNAVQHR 360
Query: 424 GPQPVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQQPYNPQQLSQPPYYPQ 483
Q A P APQ P QQ QQ PQ
Sbjct: 361 RDQNRIEYRQAAAVTIP---APQ-----------------PRQQQQQRVQQ-------PQ 393
Query: 484 QPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQNLVQ--TIPPPRIPDSL 541
Q QQRP QPR ++P ++FD LPM+Y LLP LL LV+ T+ PP + L
Sbjct: 394 QQQQQRP-YQPRQRMPD------RRFDSLPMSYAELLPELLRLGLVELCTMAPPTV---L 443
Query: 542 PRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKLINSKELTFTDPDAVAQNNPLPTH-GP 600
P Y ++ C +H GAPGH E+C AL+ +VQ LI++ + F P NNP+P H G
Sbjct: 444 PPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDANAINFA-PVPNVVNNPMPQHGGH 502
Query: 601 AVNMIQDDQEEARILSVGDIKTPLVPIHVKMCKATLFNHNHEACDICLMDPRGCIQVQND 660
VN I+ + E + +V D++T L+ + ++ +++ E C C GC Q++
Sbjct: 503 RVNNIEGKEAEDLVDNVDDVQTSLLVVKSRLLNEGVYSGCDEDCLGCAESENGCDQLRAG 562
Query: 661 MQGLLNRRELVVTREPESK-DVCVVTPVFR-----ARRPLVINPNSTKPVGTPLVICVPR 714
+QG+++ L +R + + V +T F+ +R + + PV P+ + +
Sbjct: 563 IQGMMDEGCLQFSRAVKDRGTVSTITIYFKPSEGHGQRVVSAPATNGTPVTIPVPVTISA 622
Query: 715 PTPTTA--------QKAVPYKYEGTI---------LEPGSET------------TSPVAV 745
PT A +AVP+KY+ +P ++T T AV
Sbjct: 623 PTTIVASGRRAVENSRAVPWKYDNAYRSNRRVESQTKPVNQTPVTIGLANRVPATVGPAV 682
Query: 746 DNIAENSRILRSGRIFPTVGPKSVSVPVDEPVKERNA-----GKGKAG--------EQAK 792
DN+ RSGR+F +P+++ NA KGK ++A
Sbjct: 683 DNVGGPGGFTRSGRLF-----------APQPLRDNNAEALAKAKGKQAVVEEEPVQKEAP 731
Query: 793 EFDFE-DADEVLKLIKKSEYRVVDQLLQTPAKISIMALLSSSGAHRDALRKVLDQAFVDY 851
E FE D +E +K+IKKS+Y++VDQL QTP+KISI++LL S AHR+AL K+L+ A+V
Sbjct: 732 EGSFEKDVEEFMKMIKKSDYKIVDQLNQTPSKISILSLLLCSEAHRNALLKMLNLAYVPQ 791
Query: 852 DVTLGQFESIVGNVTACNSLTFSDEDLPAEGNKHNRALLISVLCRTDSLSNVLIDTGSAL 911
++++ Q E ++ NV+ + + F++ DLP EG HN+AL I++ C+ LS+VL+DTGS+L
Sbjct: 792 EISVNQLEGVMANVSTRHGVGFTNLDLPPEGRNHNKALHITMECKGAVLSHVLVDTGSSL 851
Query: 912 NVMPKSTLDQLAYSEAPLRLSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQ 971
NV+PK L ++ L S + VRAFD ++RSV GEV LP+ +GP F + F VM+IQ
Sbjct: 852 NVLPKQILKKIDVEGFVLTPSDLIVRAFDRSKRSVCGEVTLPVKIGPEVFDIIFYVMDIQ 911
Query: 972 ASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRGKLITVSGESAFLISNLSAFSVIGGSGSD 1031
++SCLLGRPWIH AGAV+STLHQKLK+V G+++TV GE L+S+LS+F + G
Sbjct: 912 PAYSCLLGRPWIHAAGAVSSTLHQKLKYVWNGQIVTVCGEEEILVSHLSSFKYVEVDGEI 971
Query: 1032 GPSF-QGFS--AEESVGKIE-----TCMASLKDARRVIQEGKTEGWGQLVELPENKRKEG 1083
+ Q F A E V E + S K A+ V+ GK EGWG++V+LP + K G
Sbjct: 972 HETLCQVFETVALEKVAYAEQRKPGVSITSYKQAKEVVDSGKAEGWGKMVDLPVKEDKFG 1031
Query: 1084 IGF---LNSKPGMFDPTRGSFHSAGFI-HDSPETNAILD-----DAPGGVTPVFVTPGGA 1134
+G+ + G P+ +F SAG + H D D V P PGG+
Sbjct: 1032 VGYEPLQAEQNGQAGPS--TFTSAGLMNHGDVSATGSEDCDSDCDLDNWVRP--CAPGGS 1087
Query: 1135 CCNWIAVDIPSVTPCSKLNISESVEHSDPMLSPNFEVPVYEA--VAEEDEEIPNEIKWML 1192
NW A ++ VT ++ ++ + + +F+ P+Y+A +EED E+P E+ +L
Sbjct: 1088 INNWTAEEVVQVTLLTESDLMAFTMNGSVIAQYDFDNPIYQAEEESEEDCELPAELVRLL 1147
Query: 1193 EQERKTIQPHQEEIEIINLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDM 1252
+QE + IQPHQEE+E++NLGTE+ +EIKIGA+L+ S+K+R+IE++REYV+IFAWSY+DM
Sbjct: 1148 KQEERVIQPHQEELEVVNLGTEDATREIKIGAALEDSVKRRLIEMLREYVEIFAWSYQDM 1207
Query: 1253 PGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLAN 1312
PGLD D+V HRLPL+ CP VKQKLRR+ PDMA KIKEEV+KQ DAGFL + YP W+AN
Sbjct: 1208 PGLDTDIVVHRLPLREGCPSVKQKLRRTSPDMATKIKEEVQKQWDAGFLAVTSYPPWMAN 1267
Query: 1313 IVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIR 1372
IVPVPKKDGK+RMCVDYRDLN+ASPKD+FPLPHIDVLVDNTA+ VFSFMDGFSGYNQI+
Sbjct: 1268 IVPVPKKDGKVRMCVDYRDLNRASPKDDFPLPHIDVLVDNTAQSSVFSFMDGFSGYNQIK 1327
Query: 1373 MAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVK 1432
MAPED EKT+FITPWG FCY VMPFGL NAGATYQR MT +FHDM+HKEIEVYVDDMI K
Sbjct: 1328 MAPEDMEKTTFITPWGTFCYKVMPFGLKNAGATYQRAMTTLFHDMMHKEIEVYVDDMIAK 1387
Query: 1433 SGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIR 1492
S TEEEH+ L K+F RLRK+KLRLNPNKCTFGVRSGKLLGFIVS+KGIEVDP KV+AI+
Sbjct: 1388 SQTEEEHLVNLQKLFDRLRKFKLRLNPNKCTFGVRSGKLLGFIVSEKGIEVDPAKVKAIQ 1447
Query: 1493 EMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKE 1552
EMP PKTEKQVRGFLGRLNYI+RFISH+TATC PIFKLLRK+Q +KWNDDCQKAFD+IKE
Sbjct: 1448 EMPEPKTEKQVRGFLGRLNYIARFISHLTATCEPIFKLLRKNQAIKWNDDCQKAFDKIKE 1507
Query: 1553 YLLEPPILVPPVDGRPLIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCESRYS 1612
YL +PPIL+PPV GRPLIMYL+V E+SMGCVLG+ DE+G+KEH IYYLSKKFTDCE+RYS
Sbjct: 1508 YLQKPPILIPPVPGRPLIMYLSVTENSMGCVLGRHDESGRKEHAIYYLSKKFTDCETRYS 1567
Query: 1613 VLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIE 1672
+LEKTCCALAWAA+RLR YM+NHTT LISKMDP+KYIFEKPALTGR+ARWQM+L+EYDI+
Sbjct: 1568 LLEKTCCALAWAARRLRQYMLNHTTLLISKMDPVKYIFEKPALTGRVARWQMILTEYDIQ 1627
Query: 1673 YRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWG 1732
Y +QKA+KGSIL+++LA QPIEDYQP+ F+FPD+++MYLK KDC EP+ EGPDP+ +W
Sbjct: 1628 YTSQKAIKGSILSDYLAEQPIEDYQPMMFEFPDEDIMYLKMKDCKEPLVEEGPDPDDKWT 1687
Query: 1733 LIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKK 1792
L+FDGAVN+ G+G+GAVLI PKG H+PF+ARL FD TNN EYEACIMGIEEAIDLRIK
Sbjct: 1688 LMFDGAVNMNGNGVGAVLINPKGAHMPFSARLTFDVTNNEAEYEACIMGIEEAIDLRIKT 1747
Query: 1793 IVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALAT 1852
+ I+GDSALV+NQ+ G+W T P LIPYRDY RR+LTFF KV+L+HVPRDENQMADALAT
Sbjct: 1748 LDIFGDSALVVNQVNGDWNTNQPHLIPYRDYTRRILTFFKKVKLYHVPRDENQMADALAT 1807
Query: 1853 LSSMINVNGHNTVPVINVQFLDRPAYVFVAEA-IDDDKPWYHDIQVFLQTQKYPPGASNK 1911
LSSMI VN N VP + V L+RPAYVF AE+ + D+KPWY+DI+ FL+TQ+YP GAS
Sbjct: 1808 LSSMIKVNWWNHVPHVAVNRLERPAYVFAAESVVIDEKPWYYDIKNFLKTQEYPEGASKN 1867
Query: 1912 DKKTLRKLSSRFFLN-EDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMA 1970
DKKTLR+L+ F+LN +DVLYKRNFD VLLRC+D+ EA+ LM E+HEGSFGTH+ GHAMA
Sbjct: 1868 DKKTLRRLAGSFYLNQDDVLYKRNFDMVLLRCMDRPEADMLMQEVHEGSFGTHAGGHAMA 1927
Query: 1971 KKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGR 2030
KK+LRAGYYW+TM +DC+ +A++CHKCQIYAD++H+PPS LNV++SPWPF+MWGIDMIG+
Sbjct: 1928 KKLLRAGYYWMTMESDCFKYARKCHKCQIYADRVHVPPSPLNVMNSPWPFAMWGIDMIGK 1987
Query: 2031 IEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGT 2090
IEP ASNGHRFILVAIDYFTKWVEAASYAN+TKQVV RFIK +I RYGVP RIITDNG+
Sbjct: 1988 IEPTASNGHRFILVAIDYFTKWVEAASYANITKQVVTRFIKKEIICRYGVPERIITDNGS 2047
Query: 2091 NLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNIKKIIQKMVVTYKDWHEMLPYA 2150
NLNN MMKELC DFKI+HHNSSPYRP+MNGAVEAANKNIKKI++KMVVTYKDWHEMLP+A
Sbjct: 2048 NLNNKMMKELCKDFKIEHHNSSPYRPKMNGAVEAANKNIKKIVRKMVVTYKDWHEMLPFA 2107
Query: 2151 LYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLI 2210
L+GYRTSVRTSTGATP+SLVYGMEAVLPVEVEIPSLRVL++ +L EAEW ++R+++L+LI
Sbjct: 2108 LHGYRTSVRTSTGATPYSLVYGMEAVLPVEVEIPSLRVLLDVKLDEAEWIRTRFNELSLI 2167
Query: 2211 EEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWTPNYEGPY 2270
EE+R+A +CHGQLYQ RMK+AFD+ V PR ++ GDLVLK I D RGKWTPNYEGPY
Sbjct: 2168 EERRLAVVCHGQLYQRRMKRAFDQKVRPRSYQIGDLVLKRILPPGTDNRGKWTPNYEGPY 2227
Query: 2271 VVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
VVK+ FSGGAL+LT MDGE+ P PVNSD VKKYF
Sbjct: 2228 VVKKVFSGGALMLTTMDGEDFPSPVNSDVVKKYF 2261
>gb|AAQ82033.1| gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 1659 bits (4296), Expect = 0.0
Identities = 901/1763 (51%), Positives = 1178/1763 (66%), Gaps = 143/1763 (8%)
Query: 175 SIMGDDKAI-----DWEERFGALEKKMSNMRGKETVVQSIYDLCLVPDVNIPPKFKMPVF 229
S++ +D+ + + + + AL +K+ M + ++ + ++ LV + IP KFK P F
Sbjct: 115 SLLSEDEDVLGRVDERDRKVDALAEKIRAMECQNSLGFDVTNMGLVEGLRIPYKFKAPSF 174
Query: 230 EKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMGLK--GVTTFEQL 287
+KY G +CP+ H+ Y R++ AY ++ + ++ FQDSL+G + W+M LK + ++ L
Sbjct: 175 DKYNGTSCPRTHVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKRDSIRCWKDL 234
Query: 288 AEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRPPLDERELSELFY 347
EAF++QYK+N +APSR +LQSL QK ESFKEYAQR+ + AA+++PP+ EREL+++F
Sbjct: 235 GEAFLRQYKHNMDMAPSRTQLQSLCQKSGESFKEYAQRWRELAARVQPPMLERELTDMFI 294
Query: 348 ETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGVSSNGNKKFGN 407
TL + ++M C F+D+V G R E + G SS +KK
Sbjct: 295 GTLQGVFMDRMGSCPFVSFSDVVICGERTESLIKTGKIQDAGSS---------SSKKPFA 345
Query: 408 GYPKRNAQEVGMVAHGGPQPVYPNHPFVANITPQMTAPQNPNYQPPRPQGPSLYYPPLYQ 467
G P+R E V + Q A P APQ P Q
Sbjct: 346 GAPRRREGETNAVQYRRDQNRSQRCQVAAVTIP---APQ-----------------PRQQ 385
Query: 468 QPYNPQQLSQPPYYPQQPYQQRPQQQPRPQVPHNQQYQRQQFDPLPMTYGALLPSLLAQN 527
Q QQ PQQ QQ+ QPR ++P ++FD LPM+Y LLP LL
Sbjct: 386 QQQRVQQ-------PQQQQQQQRPYQPRQRMPD------RRFDSLPMSYAELLPELLRLG 432
Query: 528 LVQ--TIPPPRIPDSLPRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKLINSKELTFTD 585
+V+ T+ PP + LP Y ++ C +H GAPGH E+C AL+ +VQ LI++K + F
Sbjct: 433 MVELRTMAPPTV---LPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDAKAINFA- 488
Query: 586 PDAVAQNNPLPTH-GPAVNMIQDDQEEARILSVGDIKTPLVPIHVKMCKATLFNHNHEAC 644
P NNP+P H G VN I+ + E +++V D++T L+ + ++ +++ E C
Sbjct: 489 PVPNVVNNPMPQHGGHRVNNIEGKEAEDLVVNVDDVQTSLLVVKGRLLNGGVYSGCDEDC 548
Query: 645 DICLMDPRGCIQVQNDMQGLLNRRELVVTREPESK-DVCVVTPVFR---ARRPLVINPNS 700
C GC Q++ +QG+++ L +R + + V +T F+ R V++ +
Sbjct: 549 LGCAESENGCDQLRTGIQGMMDEGCLQFSRAVKDRGTVSTITIYFKPSEGRGQRVVSAPA 608
Query: 701 TKPVGTPLVICVP----RPTPTTA--------QKAVPYKYEGTIL--EPGSETTSPV--- 743
T GTP+ I VP PT A +AVP+KY+ T PV
Sbjct: 609 TN--GTPVTISVPVTISAPTTIAASGRRAVENSRAVPWKYDNAYRSNRRAESQTRPVNQA 666
Query: 744 ----------------AVDNIAENSRILRSGRIFPTVGPKSVSVPVDEPVKERNA----- 782
AVDN+ RSGR+F +P+++ NA
Sbjct: 667 PVTIGLPNRVPATVGPAVDNVGGPGGFTRSGRLF-----------APQPLRDNNAEALAK 715
Query: 783 GKGKAG--------EQAKEFDFE-DADEVLKLIKKSEYRVVDQLLQTPAKISIMALLSSS 833
KGK ++A E FE D +E +K+IKKS+Y++VDQL QTP+KISI++LL S
Sbjct: 716 AKGKQAVVEEEPVQKEAPEGSFEKDVEEFMKIIKKSDYKIVDQLNQTPSKISILSLLLCS 775
Query: 834 GAHRDALRKVLDQAFVDYDVTLGQFESIVGNVTACNSLTFSDEDLPAEGNKHNRALLISV 893
AHR+AL K+L+ A+V ++++ Q E ++ NV+ + + F++ DL EG HN+AL I++
Sbjct: 776 EAHRNALLKMLNLAYVPQEISVNQLEGVMANVSTRHGVGFTNLDLTPEGRNHNKALHITM 835
Query: 894 LCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVRAFDGTRRSVYGEVDLP 953
C+ LS+VL+DTGS+LNV+PK L ++ L S + VRAFDG++RSV GEV LP
Sbjct: 836 ECKGAVLSHVLVDTGSSLNVLPKQILKKIDVEGFVLTPSDLIVRAFDGSKRSVCGEVTLP 895
Query: 954 ISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRGKLITVSGESA 1013
+ +GP F + F VM+IQ ++SCLLGRPWIH AGAV+STLHQKLK+V G+++TV GE
Sbjct: 896 VKIGPEVFDIIFYVMDIQPAYSCLLGRPWIHAAGAVSSTLHQKLKYVWNGQIVTVCGEEE 955
Query: 1014 FLISNLSAFSVIGGSGSDGPSF-QGFS--AEESVGKIE-----TCMASLKDARRVIQEGK 1065
L+S+LS+F + G + Q F A E V E + S K A+ V+ GK
Sbjct: 956 ILVSHLSSFKYVEVDGEIHETLCQAFETVALEKVAYAEQRKPGASITSYKQAKEVVDSGK 1015
Query: 1066 TEGWGQLVELPENKRKEGIGF---LNSKPGMFDPTRGSFHSAGFI-HDSPETNAILD--- 1118
EGWG++V LP + K G+G+ + G P+ +F SAG + H D
Sbjct: 1016 AEGWGKMVYLPVKEDKFGVGYEPLQAEQNGQAGPS--TFTSAGLMNHGDVSATGSEDCDS 1073
Query: 1119 --DAPGGVTPVFVTPGGACCNWIAVDIPSVTPCSKLNISESVEHSDPMLSPNFEVPVYEA 1176
D V P PGG+ NW A ++ VT ++ ++ ++ M +F P+Y+A
Sbjct: 1074 DCDLDNWVRP--CAPGGSINNWTAEEVVQVTLLTESDLMAFTMNNSVMAQYDFGNPIYQA 1131
Query: 1177 --VAEEDEEIPNEIKWMLEQERKTIQPHQEEIEIINLGTEEDKKEIKIGASLDVSIKKRV 1234
+EED E+P E+ +L QE + IQPHQEE+E++NLGTE+ K+EIKIGA+L+ S+K+R+
Sbjct: 1132 EEESEEDCELPAELVRLLRQEERVIQPHQEELEVVNLGTEDAKREIKIGAALEDSVKRRL 1191
Query: 1235 IELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRK 1294
IE++REYV+IFAWSY+DMPGLD D+V HRLPL+ CP VKQKLRR+ PDMA KIKEEV+K
Sbjct: 1192 IEMLREYVEIFAWSYQDMPGLDTDIVVHRLPLREGCPSVKQKLRRTSPDMATKIKEEVQK 1251
Query: 1295 QIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTA 1354
Q DAGFL + YP W+ANIVPVPKKDGK+RMCVDYRDLN+ASPKD+FPLPHIDVLVDNTA
Sbjct: 1252 QWDAGFLAVTSYPPWVANIVPVPKKDGKVRMCVDYRDLNRASPKDDFPLPHIDVLVDNTA 1311
Query: 1355 KCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIF 1414
+ VFSFMDGFSGYNQI+MAPED EKT+FITPWG F Y VMPFGL NAGATYQR MT +F
Sbjct: 1312 QSSVFSFMDGFSGYNQIKMAPEDMEKTTFITPWGTFYYKVMPFGLKNAGATYQRAMTTLF 1371
Query: 1415 HDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGF 1474
HDM+HKEIEVYVDDMI KS TEEEH+ L K+F RLRK+KLRLNPNKCTFGVRSGKLLGF
Sbjct: 1372 HDMMHKEIEVYVDDMIAKSQTEEEHLVNLQKLFDRLRKFKLRLNPNKCTFGVRSGKLLGF 1431
Query: 1475 IVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKD 1534
IVS+KGIEVDP KV+AI+EMP PKTEKQVRGFLGRLNYI+RFISH+TATC PIFKLLRK+
Sbjct: 1432 IVSEKGIEVDPAKVKAIQEMPEPKTEKQVRGFLGRLNYIARFISHLTATCEPIFKLLRKN 1491
Query: 1535 QGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVLGQQDETGKKE 1594
Q +KWNDDCQKAFD+IKEYL +PPIL PPV GRPLIMYL+V E+SMGCVLGQ DE+G+KE
Sbjct: 1492 QAIKWNDDCQKAFDKIKEYLQKPPILTPPVPGRPLIMYLSVTENSMGCVLGQHDESGRKE 1551
Query: 1595 HVIYYLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPA 1654
H IYYLSKKFTDCE+RYS+LEKTCCALAWAA+RLR YM+NHTT LISKMDP+KYIFEKPA
Sbjct: 1552 HAIYYLSKKFTDCETRYSLLEKTCCALAWAARRLRQYMLNHTTLLISKMDPVKYIFEKPA 1611
Query: 1655 LTGRIARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAK 1714
LTGR+ARWQM+L+EYDI+Y +QKA+KGSIL+++LA QPIEDYQP+ F+FPD+++MYLK K
Sbjct: 1612 LTGRVARWQMILTEYDIQYTSQKAIKGSILSDYLAEQPIEDYQPMMFEFPDEDIMYLKVK 1671
Query: 1715 DCDEPVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVE 1774
DC+EP+ EGPDP+ +W L+FDGAVN+ G+G+GAVLI PKG HIPF+ARL FD TNN E
Sbjct: 1672 DCEEPLVEEGPDPDDKWTLMFDGAVNMNGNGVGAVLINPKGAHIPFSARLTFDVTNNEAE 1731
Query: 1775 YEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKV 1834
YEACIMGIEEAIDLRIK + IYGDSALV+NQ+ G+W T P LIPYRDY RR+LTFF KV
Sbjct: 1732 YEACIMGIEEAIDLRIKTLDIYGDSALVVNQVNGDWNTNQPHLIPYRDYTRRILTFFKKV 1791
Query: 1835 ELHHVPRDENQMADALATLSSMI 1857
L+HVPRDENQMADALATLSSMI
Sbjct: 1792 RLYHVPRDENQMADALATLSSMI 1814
>ref|XP_493776.1| unnamed protein product [Oryza sativa (japonica cultivar-group)]
gi|9927274|dbj|BAB08213.2| Similar to Arabidopsis
thaliana chromosome II BAC F26H6; putative retroelement
pol polyprotein (AC006920) [Oryza sativa (japonica
cultivar-group)]
Length = 2876
Score = 889 bits (2297), Expect = 0.0
Identities = 483/1159 (41%), Positives = 700/1159 (59%), Gaps = 53/1159 (4%)
Query: 1172 PVYEAVAEEDEEIPNEIKWMLEQERKTIQPHQEEIEIINLGTEEDKKEIKIGASLDVSIK 1231
P A +EE+ ++ L+Q QP +E+ +NLGTE+D + I + L +
Sbjct: 1746 PFEVAAVGVEEEL--DVAGALKQLDDGGQPTIDELVEMNLGTEDDPRPIFVSGMLTEEER 1803
Query: 1232 KRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEE 1291
+ + E+ D FAW+YK+MPGLD V H+L + P+ PVKQ RR P+ ++ E
Sbjct: 1804 EDYRSFLMEFRDCFAWTYKEMPGLDSRVATHKLAIDPQFRPVKQPPRRLRPEFQDQVIAE 1863
Query: 1292 VRKQIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVD 1351
V + I+ GF+ +YP+WLANIVPV KK+G++R+CVD+RDLN+A PKD+FPLP +++VD
Sbjct: 1864 VDRLINVGFIKEIQYPRWLANIVPVEKKNGQVRVCVDFRDLNRACPKDDFPLPITEMVVD 1923
Query: 1352 NTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMT 1411
+T SGYNQI+M D T+F TP G F Y VMPFGL NAGATYQR M
Sbjct: 1924 STTG------YGALSGYNQIKMDLLDAFDTAFRTPKGNFYYTVMPFGLKNAGATYQRAMQ 1977
Query: 1412 KIFHDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKL 1471
+ D+IH +E YVDDM+VK+ E H E L +F+RLR+++L++NP KC F V+SG
Sbjct: 1978 FVLDDLIHHSVECYVDDMVVKTKDHEHHQEDLRIVFERLRRHQLKMNPLKCAFAVQSGVF 2037
Query: 1472 LGFIVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLL 1531
LGF++ +GIE++P K++AI MP P+ K +R G+L YI RFIS+++ P KL+
Sbjct: 2038 LGFVIRHRGIEIEPKKIKAILNMPPPQELKDLRKLQGKLAYIRRFISNLSGRIQPFSKLM 2097
Query: 1532 RKDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVLGQQDETG 1591
+K W+++CQ FD IK YLL PP+L PV GRPLI+Y+ S+G +L Q ++ G
Sbjct: 2098 KKGTPFVWDEECQNGFDSIKRYLLNPPVLAAPVKGRPLILYIATQPASIGALLAQHNDEG 2157
Query: 1592 KKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFE 1651
KE YYLS+ E YS +EK C AL +A K+LRHYM+ H LI++ DPI+Y+
Sbjct: 2158 -KEVACYYLSRTMVGAEQNYSPIEKLCLALIFALKKLRHYMLAHQIQLIARADPIRYVLS 2216
Query: 1652 KPALTGRIARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYL 1711
+P LTGR+ +W +L+ EYDI + QKA+KG LAE LA P+ D P+ + PD+E+
Sbjct: 2217 QPVLTGRLGKWALLMMEYDITFVPQKAIKGQALAEFLATHPMPDDSPLIANLPDEEIFTA 2276
Query: 1712 KAKDCDEPVFGEGPDPESEWGLIFDGA----VNVYG-----SGIGAVLITPKGTHIPFT- 1761
+ ++ +W L FDGA +N G +G G V TP+G I +
Sbjct: 2277 ELQE--------------QWELYFDGASRKDINPDGTPRRRAGAGLVFKTPQGGVIYHSF 2322
Query: 1762 ARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYR 1821
+ L+ +C+NN EYEA I G+ A+ + ++ + +GDS L+I QI +E R P L+PY
Sbjct: 2323 SLLKEECSNNEAEYEALIFGLLLALSMEVRSLRAHGDSRLIIRQINNIYEVRKPELVPYY 2382
Query: 1822 DYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVNGHNTVPVINVQFLDRPAYV-F 1880
ARRL+ F +E+ HVPR +N ADALA L++ + G N ++ + PA +
Sbjct: 2383 TVARRLMDKFEHIEVIHVPRSKNAPADALAKLAAALVFQGDNPAQIVVEERWLLPAVLEL 2442
Query: 1881 VAEAID-------DDKPWYHDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVL 1930
+ E ++ +++ W Q FL K+ G+ +D R+L R + VL
Sbjct: 2443 IPEEVNIIITNSAEEEDWR---QPFLDYFKH--GSLPEDPVERRQLQRRLPSYIYKAGVL 2497
Query: 1931 YKRNF-DGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYN 1989
YKR++ VLLRCVD+ EA +++ E+H G G H G M I GYYW + ADC
Sbjct: 2498 YKRSYGQEVLLRCVDRSEANRVLQEVHHGVCGGHQSGPKMYHSIRLVGYYWPGIMADCLK 2557
Query: 1990 HAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYF 2049
AK CH CQI+ + H PP+ L+ WPF WGID+IG I P +S GHRFIL A DYF
Sbjct: 2558 TAKTCHGCQIHDNFKHQPPAPLHPTVPSWPFDAWGIDVIGLINPPSSRGHRFILTATDYF 2617
Query: 2050 TKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHH 2109
+KW EA V V+ F++ ++I R+GVP+RI +DN + + + +KI+ +
Sbjct: 2618 SKWAEAVPLREVKSSDVINFLERHIIYRFGVPHRITSDNAKAFKSQKIYRFMEKYKIKWN 2677
Query: 2110 NSSPYRPQMNGAVEAANKNIKKIIQKMVVTY-KDWHEMLPYALYGYRTSVRTSTGATPFS 2168
S+ Y PQ NG EA NK + KI++K V + +DWH+ L AL+ YR +VRT T ATP+S
Sbjct: 2678 YSTGYYPQANGMAEAFNKTLGKILKKTVDKHRRDWHDRLYEALWAYRVTVRTPTQATPYS 2737
Query: 2169 LVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRM 2228
LVYG EAVLP+E+++PSLRV + EL++ E + R+ +L+ +EE+R+ AL + +LY+ M
Sbjct: 2738 LVYGNEAVLPLEIQLPSLRVAIHDELTKDEQIRLRFQELDAVEEERLGALQNLELYRQNM 2797
Query: 2229 KQAFDKGVHPREFKEGDLVLKCIKSF--QPDPRGKWTPNYEGPYVVKRAFSGGALILTNM 2286
+A+DK V R F++G+LVL + +GK+ P +EGPYV+++A+ GGA L +
Sbjct: 2798 VRAYDKLVKQRVFRKGELVLVLRRPIVVTHKMKGKFEPKWEGPYVIEQAYDGGAYQLIDH 2857
Query: 2287 DGEELPRPVNSDAVKKYFV 2305
G + P+N +KKYFV
Sbjct: 2858 QGSQPMPPINGRFLKKYFV 2876
Score = 117 bits (292), Expect = 6e-24
Identities = 91/290 (31%), Positives = 136/290 (46%), Gaps = 20/290 (6%)
Query: 755 LRSGRIFPTVGPKSVSVP-VDEPVKERNAGKGKAGEQAKEFDFEDADEVLKLIKKSEYRV 813
LR G+ P P VP VD+P K KA + + + K +Y+V
Sbjct: 1258 LRGGKTLPD--PHKSKVPNVDKPAK-------KASPPGEAPEAPETKTGSKEKPAVDYKV 1308
Query: 814 VDQLLQTPAKISIMALLSSSGAHRDALRKVLDQAFVDYDVTLGQFESIVGNVTACNSLTF 873
+ L + PA +S+ L R+AL K L QA Y+V + + + N N +TF
Sbjct: 1309 LAHLKRIPALLSVYDALMMVPDLREALIKAL-QAPEVYEVDMAKHR-LYDNPLFVNEITF 1366
Query: 874 SDEDLPAEGNKHNRALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSK 933
+DED +G HNR L I + L +LID GSA+N++P +L + ++ L
Sbjct: 1367 ADEDNIIKGGDHNRPLYIEGNIGSAHLRRILIDLGSAVNILPVRSLTRAGFTTKDLEPID 1426
Query: 934 VTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTL 993
V + FD + G + + I + F+V F V+E S+S LLGRPWIH V STL
Sbjct: 1427 VVICGFDNQGKPTLGAITIKIQMSTFSFKVRFFVIEANTSYSALLGRPWIHKYRVVPSTL 1486
Query: 994 HQKLKFVSRGKLITVSGESAFLISNLSAFSVIGGSGSDGPSFQGFSAEES 1043
HQ LKF+ +G + SN S +++ +D + F EE+
Sbjct: 1487 HQCLKFLDG------NGVQQRITSNFSPYTIQESYHADAKYY--FPVEEN 1528
Score = 42.0 bits (97), Expect = 0.25
Identities = 29/82 (35%), Positives = 43/82 (52%), Gaps = 9/82 (10%)
Query: 524 LAQNLVQTIPPPRIPDSLPRWYRPDLHCIYHQGAPGHDVERCFALKKEVQKLINSKELTF 583
L ++ +P PR PD + P L+C YH+ GH +E C A K+ +Q+ +N K +
Sbjct: 993 LMEHRALNLPEPRRPDQVTMTDNP-LYCPYHRYI-GHAIEDCIAFKEWLQRAVNEKRINL 1050
Query: 584 TDPDAVAQNNPLPTHGPAVNMI 605
D DA+ NP AVNM+
Sbjct: 1051 -DADAI---NP---DYHAVNMV 1065
>gb|AAP51864.1| putative retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|37530550|ref|NP_919577.1| putative
retroelement pol polyprotein [Oryza sativa (japonica
cultivar-group)] gi|18581535|gb|AAK52540.2| Putative
retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 770 bits (1989), Expect = 0.0
Identities = 425/1101 (38%), Positives = 634/1101 (56%), Gaps = 58/1101 (5%)
Query: 1211 LGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKPEC 1270
L + +D +EI I ++ + +IEL++E+ D F W Y +MPG +VEHRLPLKP
Sbjct: 524 LMSADDLEEIDISPAIGQG-RHLLIELLKEFRDCFTWEYYEMPGHSRSIVEHRLPLKPGV 582
Query: 1271 PPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVDYR 1330
P +Q RR DM +K E+++ DAGF+ Y +W+++IVPV KK+GK+R+C+D+R
Sbjct: 583 RPHQQLPRRCKADMLEPVKAEIKRLYDAGFIRPCRYAEWVSSIVPVIKKNGKVRVCIDFR 642
Query: 1331 DLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPW--G 1388
LNKA+PKD +P+ D LVD + K+ SFMDG +GYNQI MA ED KT+F P G
Sbjct: 643 YLNKATPKDEYPMLVADQLVDAASGHKILSFMDGNAGYNQIFMAEEDIHKTTFRCPGAIG 702
Query: 1389 AFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQ 1448
F +VV+ F L +AGATYQR M I+HD+I +EVY+DD++VKS E+H+ L K+F+
Sbjct: 703 LFEWVVITFVLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIADLRKVFE 762
Query: 1449 RLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLG 1508
R RKY L++NP KC FGV +G+ LGF+V ++GIEV + AI+++ P+ + +++ +G
Sbjct: 763 RTRKYGLKMNPTKCAFGVSAGQFLGFLVHERGIEVTQRSINAIKKIKPPEDKTELQEMIG 822
Query: 1509 RLNYISRFISHMTATCGPIFKLLR--KDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDG 1566
++N++ RFIS ++ P LLR DQ W + QKA D IKEYL PP+L+PP G
Sbjct: 823 KINFVRRFISILSGKLEPFTPLLRLKADQQFAWGAEQQKALDNIKEYLSSPPVLIPPQKG 882
Query: 1567 RPLIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAK 1626
P +YL+ + S+ VL Q+ E +KE V++YLS++ + E+RYS +EK C L ++
Sbjct: 883 IPFRLYLSAGDKSISSVLIQELE--RKERVVFYLSRRLLNAETRYSPVEKLCLCLYFSCT 940
Query: 1627 RLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQKAVKGSILAE 1686
RLRHY++++ +I K D +KY+ P L GRI +W L+E+D+ Y + KA+KG +A
Sbjct: 941 RLRHYLLSNECTVICKADVVKYMLSAPILKGRIGKWIFSLTEFDLRYESPKAIKGQAIAN 1000
Query: 1687 HLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDGAVNVYGSGI 1746
+ D+ D + G W L FDG+V +G GI
Sbjct: 1001 FIV------------DYRDDSI---------------GSVEVVPWTLFFDGSVCTHGCGI 1033
Query: 1747 GAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQI 1806
G V+I+P+G F ++ TNN EYEA + G++ ++ I I GDS LVI+Q+
Sbjct: 1034 GLVIISPRGACFEFAYTIKPYATNNQAEYEAVLKGLQLLKEVEADTIEIMGDSLLVISQL 1093
Query: 1807 KGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVNGHNTVP 1866
GE+E ++ LI Y + + L+ F V L HV R++N A+ LA +S
Sbjct: 1094 AGEYECKNDTLIVYNEKCQELMREFRLVTLKHVSREQNIEANDLAQGAS----------- 1142
Query: 1867 VINVQFLDRPAYVFVAEAIDDDKPWYHDIQVFLQTQKYPPGASNKDKKTLRKLSSRFFLN 1926
+ + + + VA DD W + + +LQ S + LR + ++ L
Sbjct: 1143 --GYKLMIKHVQIEVAVITADD--WRYYVYQYLQDP------SQSASRKLRYKALKYILL 1192
Query: 1927 EDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHAD 1986
+D LY R DGVLL+C+ +A+ + E+HEG GTH H M + AGY+W TM D
Sbjct: 1193 DDELYYRTIDGVLLKCLSTDQAKVAIGEVHEGICGTHQSAHKMKWLLRCAGYFWPTMLED 1252
Query: 1987 CYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAI 2046
C+ + K C CQ + P S +N I PWPF WGIDMIG I P +S GH+FILVA
Sbjct: 1253 CFRYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMINPPSSKGHKFILVAT 1312
Query: 2047 DYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKI 2106
DYFTKWVEA V ++F++ ++I R G+P I TD G+ ++ + D I
Sbjct: 1313 DYFTKWVEAIPLKKVDSGDAIQFVQEHIIYRVGIPQTITTDQGSIFVSDEFIQFADSMGI 1372
Query: 2107 QHHNSSPYRPQMNGAVEAANKNIKKIIQKMVVTY-KDWHEMLPYALYGYRTSVRTSTGAT 2165
+ NSSPY Q NG EA+NK++ K+I++ + Y + WH L AL+ YR + S
Sbjct: 1373 KLLNSSPYYAQANGQAEASNKSLIKLIKRKISDYPRQWHTRLAEALWSYRMACHGSIQVP 1432
Query: 2166 PFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQ 2225
P+ LVY +A+LP EV I S R ++ +L+ E+ D+ + + R+ AL +
Sbjct: 1433 PYKLVYRHKAILPWEVRIGSRRTELQNDLTADEYSNLMADEREDLVQSRLRALAKVIKDK 1492
Query: 2226 SRMKQAFDKGVHPREFKEGDLVLKCIKSF--QPDPRGKWTPNYEGPYVVKRAFSGGALIL 2283
R+ + ++K V P++F EG+L+ K I + GKW+PN+EGP+ + + S GA +L
Sbjct: 1493 ERVTRHYNKKVVPKDFSEGELIWKLILPIGTRDSKFGKWSPNWEGPFQIHKVVSKGAYML 1552
Query: 2284 TNMDGEELPRPVNSDAVKKYF 2304
+DGE R +N +KKY+
Sbjct: 1553 QGLDGEVYDRALNGKYLKKYY 1573
Score = 72.0 bits (175), Expect = 2e-10
Identities = 41/138 (29%), Positives = 72/138 (51%)
Query: 883 NKHNRALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVRAFDGT 942
N+H + L I+ +S +++D G+A+N+MP +T +L + L + + ++ F G
Sbjct: 358 NRHLKPLYINGYVNGKPMSKMMVDGGAAVNLMPYATFRKLGRNAEDLIKTNMVLKDFGGN 417
Query: 943 RRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSR 1002
G +++ ++VG TF V++ + S+S LLGR WIH + ST+HQ L
Sbjct: 418 PSETMGVLNVELTVGSKTIPTTFFVIDGKGSYSLLLGRDWIHANCCIPSTIHQCLIQWQG 477
Query: 1003 GKLITVSGESAFLISNLS 1020
K+ V +S + N S
Sbjct: 478 DKIEIVPADSQLKMENPS 495
>gb|AAF79618.1| F5M15.26 [Arabidopsis thaliana] gi|25402907|pir||H86337 protein
F5M15.26 [imported] - Arabidopsis thaliana
Length = 1838
Score = 761 bits (1964), Expect = 0.0
Identities = 434/1131 (38%), Positives = 644/1131 (56%), Gaps = 54/1131 (4%)
Query: 1194 QERKTIQPHQEEIEI-----INLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWS 1248
+E KT EE E+ +N+ + + + +GA + SI+ +I L++ FAWS
Sbjct: 741 KEAKTPSRSYEESELCRTEMVNIDESDPTRCVGVGAEISPSIRLELIALLKRNSKTFAWS 800
Query: 1249 YKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQ 1308
+DM G+DP + H L + P PVKQK R+ P+ A + EEV K + AG ++ +YP+
Sbjct: 801 IEDMKGIDPAITAHELNVDPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPE 860
Query: 1309 WLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGY 1368
WLAN V V KK+GK R+CVDY DLNKA PKD++PLPHID LV+ T+ + SFMD FSGY
Sbjct: 861 WLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGY 920
Query: 1369 NQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDD 1428
NQI M +D+EKTSF+T G +CY VM FGL NAGATYQR + K+ D I + +EVY+DD
Sbjct: 921 NQILMHKDDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDD 980
Query: 1429 MIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKV 1488
M+VKS E+HVE+L K F L Y ++LNP KCTFGV SG+ LG++V+++GIE +P ++
Sbjct: 981 MLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQI 1040
Query: 1489 RAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFD 1548
RAI E+P P+ ++V+ GR+ ++RFIS T C P + LL++ W+ D ++AF+
Sbjct: 1041 RAILELPSPRNAREVQRLTGRIAALNRFISRSTDKCLPFYNLLKRRAQFDWDKDSEEAFE 1100
Query: 1549 QIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCE 1608
++K+YL PPILV P G L +Y+ V + ++ VL ++D ++ I+Y SK + E
Sbjct: 1101 KLKDYLSTPPILVKPEVGETLYLYIAVSDHAVSSVLVREDR--GEQRPIFYTSKSLVEAE 1158
Query: 1609 SRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSE 1668
+RY V+EK A+ +A++LR Y +HT +++ P++ P+ +GR+ +W + LSE
Sbjct: 1159 TRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTD-QPLRVALHSPSQSGRMTKWAVELSE 1217
Query: 1669 YDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPE 1728
YDI++R + A+K +LA+ L P++ + V G +
Sbjct: 1218 YDIDFRPRPAMKSQVLADFLIELPLQ--------------------SAERAVSGNRGE-- 1255
Query: 1729 SEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDL 1788
EW L DG+ + GSGIG L++P + + RLRF TNN+ EYE I G+ A +
Sbjct: 1256 -EWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRFVATNNVAEYEVLIAGLRLAAGM 1314
Query: 1789 RIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMAD 1848
+I I + DS L+ Q+ GE+E ++ + Y + + F +L +PR +N AD
Sbjct: 1315 QITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQLMTKDFENFKLSKIPRGDNAPAD 1374
Query: 1849 ALATLSSMINVNGHNTVPV--INVQFLDRPAYVFVAEAI--------DDDKPWYHDIQVF 1898
ALA L+ + + +PV I+ +D V + I D W +I+ +
Sbjct: 1375 ALAALALTSDSDLRRIIPVESIDKPSIDSTDAVEIVNTIRSSNAPDPADPTDWRVEIRDY 1434
Query: 1899 LQTQKYPPGASNKDKKTLRKL---SSRFFLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEI 1955
L P DK T R+L ++++ L ++ L K + G +L C+ E ++M E
Sbjct: 1435 LSDGTLP-----SDKWTARRLRIKAAKYTLMKEHLLKVSAFGAMLNCLHGTEINEIMKET 1489
Query: 1956 HEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPSMLNVIS 2015
HEG+ G HS G A+A K+ + G+YW TM +DC +C +CQ +A IH P +L
Sbjct: 1490 HEGAAGNHSGGRALALKLKKLGFYWPTMISDCKTFTAKCEQCQRHAPTIHQPTELLRAGV 1549
Query: 2016 SPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLI 2075
+P+PF W +D++G + AS RFILV DYFTKWVEA SYA + V F+ +I
Sbjct: 1550 APYPFMRWAMDIVGPM--PASRQKRFILVMTDYFTKWVEAESYATIRANDVQNFVWKFII 1607
Query: 2076 SRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNIKKIIQK 2135
R+G+P IITDNG+ + + C +KI+ + S+P PQ NG EA NK I ++K
Sbjct: 1608 CRHGLPYEIITDNGSQFISLSFENFCASWKIRLNKSTPRYPQGNGQAEATNKTILSGLKK 1667
Query: 2136 MVVTYKD-WHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIPSLRVLMEAEL 2194
+ K W + L L+ YRT+ R++T TPF+ YGMEA+ P EV SLR M +
Sbjct: 1668 RLDEKKGAWADELDGVLWSYRTTPRSATDQTPFAHAYGMEAMAPAEVGYSSLRRSMMVKN 1727
Query: 2195 SEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEGDLVL-KCIKS 2253
E + D+L+ +EE R AALC Q YQ + +++ VH R F GDLVL K ++
Sbjct: 1728 PELN-DRMMLDRLDDLEEIRNAALCRIQNYQLAAAKHYNQKVHNRHFDVGDLVLRKVFEN 1786
Query: 2254 FQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
GK N+EG Y V + G L M G +PR NS +K+Y+
Sbjct: 1787 TAEINAGKLGANWEGSYQVSKIVRPGDYELLTMSGTAVPRTWNSMHLKRYY 1837
Score = 78.6 bits (192), Expect = 2e-12
Identities = 43/138 (31%), Positives = 73/138 (52%)
Query: 871 LTFSDEDLPAEGNKHNRALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLR 930
++F+D DL HN L++ ++ ++ VLIDTGS+++++ K L + ++ ++
Sbjct: 600 ISFTDVDLEGLDTPHNDPLVVELIISDSRVTRVLIDTGSSVDLIFKDVLTAMNITDRQIK 659
Query: 931 LSKVTVRAFDGTRRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVT 990
+ FDG G + LPI VG V F V+ A ++ +LG PWIH A+
Sbjct: 660 PVSKPLAGFDGDFVMTIGTIKLPIFVGGLIAWVKFVVIGKPAVYNVILGTPWIHQMQAIP 719
Query: 991 STLHQKLKFVSRGKLITV 1008
ST HQ +KF + + T+
Sbjct: 720 STYHQCVKFPTHNGIFTL 737
Score = 50.8 bits (120), Expect = 5e-04
Identities = 35/113 (30%), Positives = 49/113 (42%), Gaps = 6/113 (5%)
Query: 230 EKYQGDTCPQNHLTMYI----RRMIAYKNNVPLLIHCFQDSLTGPAHTWFMGLK--GVTT 283
E Y G P+ LT + R + N F + LTGPAH WF LK + +
Sbjct: 222 ESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIEHLTGPAHNWFSRLKPNSIDS 281
Query: 284 FEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRPP 336
F QL +F++ Y S +L S++Q KES + + RF I P
Sbjct: 282 FHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFVDRFKLVVTNITVP 334
>gb|AAP52501.1| putative gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37531824|ref|NP_920214.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
gi|22128685|gb|AAM92798.1| putative gag-pol precursor
[Oryza sativa (japonica cultivar-group)]
Length = 1986
Score = 708 bits (1828), Expect = 0.0
Identities = 401/1109 (36%), Positives = 612/1109 (55%), Gaps = 68/1109 (6%)
Query: 1217 KKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQK 1276
++EI++ A+ + ++ I ++ DIFAW DMPG+ +V+EH L +K + P+KQ+
Sbjct: 924 REEIRLAATTESAL----ITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQR 979
Query: 1277 LRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKAS 1336
LRR D IKEE+ K + AGF+ +P WLAN V V KK G+ RMCVDY DLNK+
Sbjct: 980 LRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSC 1039
Query: 1337 PKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMP 1396
PKD F LP ID +VD+TA C++ SF+D +SGY+QIR+ D KTSFITP+GA+CYV MP
Sbjct: 1040 PKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMP 1099
Query: 1397 FGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLR 1456
FGL NAGATYQR + + F I + +E YVDD++VK+ +++ + L + F +R ++++
Sbjct: 1100 FGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMK 1159
Query: 1457 LNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRF 1516
LNP KCTFGV SGKLLGF+VS +GI+ +P+KV AI M P T+K V+ G + +SRF
Sbjct: 1160 LNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRF 1219
Query: 1517 ISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVL 1576
+S + P FKLL+K +W + QKAF+ K+ L +PP+L P PL++Y++
Sbjct: 1220 VSRLGERGMPFFKLLKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSAT 1279
Query: 1577 EDSMGCVL-GQQDETG---KKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYM 1632
+ VL +++E G K + IY++S+ D ++RY ++K + ++L HY
Sbjct: 1280 SQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKARYPQVQKLLYGILITTRKLSHYF 1339
Query: 1633 INHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQP 1692
H+ +++ P+ + GRIA+W + L DI ++ + ++K LA+ +A
Sbjct: 1340 QGHSVTVVTSF-PLGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVA--- 1395
Query: 1693 IEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLIT 1752
++ + D P +++ W + FDG+ + G+G G VLI+
Sbjct: 1396 --EWTECQEDTPVEKI--------------------EHWTMHFDGSKRLSGTGAGVVLIS 1433
Query: 1753 PKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWET 1812
P G + + + F ++N+ EYEA + G+ AI L IK++++ GDS LV+NQ+ EW
Sbjct: 1434 PTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSC 1493
Query: 1813 RHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVNGHNTVPV-INVQ 1871
++ YR R+L F+ +EL HV R N+ AD LA S T P + V+
Sbjct: 1494 IDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGSK-----RETAPSDVFVE 1548
Query: 1872 FLDRPAYVFVAEAIDDD--------KPWYHDIQVFLQTQKYPPGASNKDKKTLRKLSSR- 1922
L P D D W FL +Q+ P +DK ++S R
Sbjct: 1549 HLYTPTVPHKDTTQDADTRNIAMVEADWREPFIRFLTSQELP-----QDKDEAERISRRS 1603
Query: 1923 --FFLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYW 1980
+ L+E LYK++ G+L RCV E +L+ +IH G G H+ + K R G++W
Sbjct: 1604 KLYVLHESELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFW 1663
Query: 1981 ITMHADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHR 2040
T +D + C CQ +A +IH+P L I WPF++WG+DM+G + KA G+
Sbjct: 1664 PTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYT 1722
Query: 2041 FILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKEL 2100
+ VAID F+KW+EA +T F N++ R+GVPNRIITDNGT + K+
Sbjct: 1723 HLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGTQFTGGVFKDF 1781
Query: 2101 CDDFKIQHHNSSPYRPQMNGAVEAANKNI-----KKIIQKMVVTYKDWHEMLPYALYGYR 2155
C+DF I+ +S P NG VE AN I ++ ++ W + LP L+ R
Sbjct: 1782 CEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLR 1841
Query: 2156 TSVRTSTGATPFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLIEEKRM 2215
T+ +TG +PF LVYG EA+LP EVE SLR E + + R D L+ +EE R
Sbjct: 1842 TTPSRATGQSPFFLVYGAEAMLPSEVEFESLRF---RNFREERYEEDRVDDLHRLEEVRE 1898
Query: 2216 AALCHGQLYQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWTPNYEGPYVVKRA 2275
AAL Y +++ ++ V R F GDLVL+ I++ + R K +P +EGP+++
Sbjct: 1899 AALIQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEV 1956
Query: 2276 FSGGALILTNMDGEELPRPVNSDAVKKYF 2304
G+ L DG + N + +++++
Sbjct: 1957 TRPGSYRLKREDGTLVDNSWNIEHLRRFY 1985
Score = 48.9 bits (115), Expect = 0.002
Identities = 31/128 (24%), Positives = 59/128 (45%), Gaps = 5/128 (3%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 773 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKSSNAPFHGVIPGLSATPL 832
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRG 1003
G++ LP++ G E ++F+V + + ++ +LG P + AV + +K
Sbjct: 833 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGHPALAKFMAVPHYTYMMMKMPGPR 892
Query: 1004 KLITVSGE 1011
++T+ +
Sbjct: 893 GVLTLRSD 900
Score = 48.1 bits (113), Expect = 0.003
Identities = 53/221 (23%), Positives = 92/221 (40%), Gaps = 19/221 (8%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 547
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 548 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 600
Query: 392 GGSSGVSSNGNKKFGNGY-----PKRNAQE-VGMVAHGGPQ 426
G S N G G KR +E V AH Q
Sbjct: 601 SGPSWKPKNTPATGGGGSNNHKDRKRKPEELVATAAHSSRQ 641
>ref|XP_475022.1| OSJNBb0093G06.3 [Oryza sativa (japonica cultivar-group)]
gi|38347562|emb|CAE04995.2| OSJNBb0093G06.3 [Oryza sativa
(japonica cultivar-group)]
Length = 1986
Score = 708 bits (1827), Expect = 0.0
Identities = 397/1106 (35%), Positives = 612/1106 (54%), Gaps = 62/1106 (5%)
Query: 1217 KKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQK 1276
++EI++ A+ + ++ I ++ DIFAW DMPG+ +V+EH L +K + P+KQ+
Sbjct: 924 REEIRLAATTESAL----ITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQR 979
Query: 1277 LRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKAS 1336
LRR D IKEE+ K + AGF+ +P WLAN V V KK G+ RMCVDY DLNK+
Sbjct: 980 LRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSC 1039
Query: 1337 PKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMP 1396
PKD F LP ID +VD+TA C++ SF+D +SGY+QIR+ D KTSFITP+GA+CYV MP
Sbjct: 1040 PKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMP 1099
Query: 1397 FGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLR 1456
FGL NAGATYQR + + F I + +E YVDD++VK+ +++ + L + F +R ++++
Sbjct: 1100 FGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLISDLEETFASIRAFRMK 1159
Query: 1457 LNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRF 1516
LNP KCTFGV SGKLLGF+VS +GI+ +P+KV AI M P T+K V+ G + +SRF
Sbjct: 1160 LNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRF 1219
Query: 1517 ISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVL 1576
+S + P FKLL+K +W + QKAF+ K+ L +PP+L P PL++Y++
Sbjct: 1220 VSRLGERGMPFFKLLKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSAT 1279
Query: 1577 EDSMGCVL-GQQDETG---KKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYM 1632
+ VL +++E G K + IY++S+ D ++RY ++K + ++L HY
Sbjct: 1280 SQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYF 1339
Query: 1633 INHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQP 1692
H+ +++ P+ + GRIA+W + L DI ++ + ++K LA+ +A
Sbjct: 1340 QGHSVTVVTSF-PLGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVA--- 1395
Query: 1693 IEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLIT 1752
++ + D P +++ W + FDG+ + G+G G VLI+
Sbjct: 1396 --EWTECQEDTPVEKI--------------------EHWTMHFDGSKRLSGTGAGVVLIS 1433
Query: 1753 PKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWET 1812
P G + + + F ++N+ EYEA + G+ AI L IK++++ GDS LV+NQ+ EW
Sbjct: 1434 PTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSC 1493
Query: 1813 RHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVN------GHNTVP 1866
++ YR R+L F+ +EL HV R N+ AD LA S H P
Sbjct: 1494 IDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKREAAPSDVFVEHLYTP 1553
Query: 1867 VINVQFLDRPAYVFVAEAIDDDKPWYHDIQVFLQTQKYPPGASNKDKKTLRKLSSR---F 1923
+ + + A ++ D W FL +Q+ P +DK ++S R +
Sbjct: 1554 TVPHKDTTQDADTHDVVMVEAD--WREPFIRFLSSQELP-----QDKDEAERISRRSKLY 1606
Query: 1924 FLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITM 1983
++E LYK++ G+L RCV E +L+ +IH G G H+ + K R G++W T
Sbjct: 1607 VMHESELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTA 1666
Query: 1984 HADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHRFIL 2043
+D + C CQ +A +IH+P L I WPF++WG+DM+G + KA G+ +
Sbjct: 1667 VSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLF 1725
Query: 2044 VAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELCDD 2103
VAID F+KW+EA +T F N++ R+GVPNRIITDNGT + K+ C+D
Sbjct: 1726 VAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGTQFTGGVFKDFCED 1784
Query: 2104 FKIQHHNSSPYRPQMNGAVEAANKNI-----KKIIQKMVVTYKDWHEMLPYALYGYRTSV 2158
F I+ +S P NG VE AN I ++ ++ W + LP L+ RT+
Sbjct: 1785 FGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTP 1844
Query: 2159 RTSTGATPFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAAL 2218
+TG +PF LVYG EA+LP EVE SLR E + + R D L+ +EE R AAL
Sbjct: 1845 SRATGQSPFFLVYGAEAMLPSEVEFESLRF---RNFREERYEEDRVDDLHRLEEAREAAL 1901
Query: 2219 CHGQLYQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSG 2278
Y +++ ++ V R F GDLVL+ I++ + R K +P +EGP+++
Sbjct: 1902 IQSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRP 1959
Query: 2279 GALILTNMDGEELPRPVNSDAVKKYF 2304
G+ L DG + N + +++++
Sbjct: 1960 GSYRLKREDGTLVDNSWNIEHLRRFY 1985
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 773 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 832
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 833 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 887
Score = 48.9 bits (115), Expect = 0.002
Identities = 54/221 (24%), Positives = 92/221 (41%), Gaps = 19/221 (8%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES +EY +RF ++ +I
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLREYIRRFSEQRNKISD 547
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 548 ISDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 600
Query: 392 GGSSGVSSNGNKKFGNGY-----PKRNAQE-VGMVAHGGPQ 426
G S N G G KR +E V AH Q
Sbjct: 601 SGPSWKPKNTPATGGGGSNNHKDRKRKPEELVATAAHSSRQ 641
>ref|XP_474675.1| OSJNBa0027O01.4 [Oryza sativa (japonica cultivar-group)]
gi|38346188|emb|CAD39529.2| OSJNBa0027O01.4 [Oryza sativa
(japonica cultivar-group)]
Length = 2013
Score = 707 bits (1824), Expect = 0.0
Identities = 398/1080 (36%), Positives = 599/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 973 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 1032
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLNK+ PKD F LP ID +VD+TA C++ SF+
Sbjct: 1033 EVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFL 1092
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1093 DCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1152
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 1153 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 1212
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K +W +
Sbjct: 1213 ANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPE 1272
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPP+L P PL++Y++ + VL +++E G K + IY
Sbjct: 1273 AQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIY 1332
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H+ +++ P+ I + GR
Sbjct: 1333 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREVNGR 1391
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A + +C E
Sbjct: 1392 IAKWALELMSLDISFKPRISIKSQALADFVA----------------------EWTECQE 1429
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
E + W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1430 DTPAENME---HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1486
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1487 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSYLDDNMMAYRQEVRKLEDKFDGLELSH 1546
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S H P + + + A A ++ D W
Sbjct: 1547 VLRHNNEAADRLANFGSKREAAPSDVFVEHLYTPTVPHKDTTQVAGTHDAAMVEVD--WR 1604
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
+ FL +Q+ P +DK ++S R + L+E LYK++ G+L RCV E
Sbjct: 1605 EPLIRFLTSQELP-----QDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGR 1659
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1660 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1719
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1720 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1778
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNGT + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1779 F-INIVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1837
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1838 LQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1897
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1898 SLRF---RNFREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVG 1954
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1955 DLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2012
Score = 50.1 bits (118), Expect = 0.001
Identities = 46/197 (23%), Positives = 85/197 (42%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 547
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 548 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 600
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 601 SGPSWKQNKGTPATGGG 617
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 774 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 833
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 834 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 888
>ref|XP_475035.1| OSJNBa0042F21.5 [Oryza sativa (japonica cultivar-group)]
gi|38347303|emb|CAE02298.2| OSJNBa0042F21.5 [Oryza sativa
(japonica cultivar-group)]
Length = 1950
Score = 705 bits (1820), Expect = 0.0
Identities = 397/1080 (36%), Positives = 597/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 910 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 969
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLNK+ PKD F LP ID +VD+TA C++ SF+
Sbjct: 970 EVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFL 1029
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1030 DCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1089
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 1090 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 1149
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K +W +
Sbjct: 1150 ANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDNFQWGPE 1209
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPPIL P PL++Y++ + VL +++E G K + IY
Sbjct: 1210 AQKAFEDFKKLLTEPPILASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIY 1269
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H+ +++ P+ I GR
Sbjct: 1270 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREANGR 1328
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A + +C E
Sbjct: 1329 IAKWALELMSLDISFKPRISIKSQALADFVA----------------------EWTECQE 1366
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
E + W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1367 DTPAENME---HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1423
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1424 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1483
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S + H P + + + A ++ D W
Sbjct: 1484 VLRHNNEAADRLANFGSKREMAPSDVFVEHLYTPTVPHKDTTQDADTHDVALVEAD--WR 1541
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
FL +Q+ P +DK ++S R + ++E LYK++ G+L RCV E
Sbjct: 1542 EPFIRFLTSQELP-----QDKDEAERISRRSKLYAMHEAELYKKSPSGILQRCVSLEEGR 1596
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1597 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1656
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1657 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1715
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNGT + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1716 F-INIVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1774
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1775 LQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1834
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1835 SLRF---RNFREERYEEDRVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVG 1891
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1892 DLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 1949
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 711 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 770
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 771 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 825
Score = 48.5 bits (114), Expect = 0.003
Identities = 45/197 (22%), Positives = 84/197 (41%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 365 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 424
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 425 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 484
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+ + AV+ S
Sbjct: 485 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYTKAEDAVTASKQ 537
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 538 SGPSWKQNKGTPATGGG 554
>ref|XP_470757.1| putative gag-pol precursor [Oryza sativa] gi|18071370|gb|AAL58229.1|
putative gag-pol precursor [Oryza sativa]
Length = 2026
Score = 705 bits (1819), Expect = 0.0
Identities = 396/1080 (36%), Positives = 599/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 977 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 1036
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLNK+ PKD F LP ID +VD+TA C++ SF+
Sbjct: 1037 EVHHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFL 1096
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1097 DCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1156
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 1157 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 1216
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K +W +
Sbjct: 1217 ANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPE 1276
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPP+L P PL++Y++ + VL +++E G K + IY
Sbjct: 1277 AQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIY 1336
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H+ +++ P+ I GR
Sbjct: 1337 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREANGR 1395
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A ++ + D P +++
Sbjct: 1396 IAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPAEKM---------- 1440
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1441 ----------EHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1490
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1491 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1550
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S V H P + + + A ++ D W
Sbjct: 1551 VLRHNNEAADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVEAD--WR 1608
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
+ FL +Q+ P +DK ++S R + ++E LYK++ G+L RCV E
Sbjct: 1609 EPLIRFLTSQELP-----QDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGR 1663
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1664 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1723
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1724 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNAPDF 1782
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNG + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1783 F-INIVHRFGVPNRIITDNGRQFTGGVFKDCCEDFGIKICYASVAHPMSNGQVERANGMI 1841
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W E LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1842 LQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1901
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1902 SLRF---RNFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVG 1958
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1959 DLVLRKIQTTR--DRHKLSPLWEGPFIISAVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2016
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 778 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 837
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 838 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 892
Score = 47.4 bits (111), Expect = 0.006
Identities = 32/124 (25%), Positives = 61/124 (48%), Gaps = 2/124 (1%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 432 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDSKAMANYLPVALADSARSWLHG 491
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 492 LPRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNVVQKSGESLRDYIRRFSEQRNKISD 551
Query: 336 PLDE 339
D+
Sbjct: 552 ITDD 555
>gb|AAP53095.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533012|ref|NP_920808.1| putative retroelement [Oryza
sativa (japonica cultivar-group)]
gi|19881590|gb|AAM00991.1| Putative retroelement [Oryza
sativa]
Length = 2017
Score = 703 bits (1815), Expect = 0.0
Identities = 397/1080 (36%), Positives = 596/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 977 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 1036
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLNK PKD F LP ID +VD+TA C++ SF+
Sbjct: 1037 EVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSFL 1096
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1097 DCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1156
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 1157 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 1216
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K W +
Sbjct: 1217 ANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFHWGPE 1276
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPP+L P PL++Y++ + VL +++E G K + IY
Sbjct: 1277 AQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEDGHVQKVQRPIY 1336
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H +++ P+ I GR
Sbjct: 1337 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHLVTVVTSF-PLGDILHNREANGR 1395
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A ++ + D P ++V
Sbjct: 1396 IAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPAEKV---------- 1440
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1441 ----------EHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1490
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1491 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1550
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S V H P + + + A ++ D W
Sbjct: 1551 VLRHNNKAADRLANFGSKREVAPSDVFVEHLYAPTVPHKDTTQVAGTHDVAMVEAD--WR 1608
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
+ FL +Q+ P +DK ++S R + ++E LYK++ G+L RCV E
Sbjct: 1609 EPLIRFLTSQELP-----QDKDEAERISRRSRLYVMHEAELYKKSPSGILQRCVSLEEGR 1663
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1664 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1723
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1724 ELQTIPLSWPFAVWGLDMVGPFK-KAFGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1782
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNG + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1783 F-INIVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1841
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W E LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1842 LQGIKARVFDRLKPYAGKWVEQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1901
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1902 SLRF---RNFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVG 1958
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1959 DLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLIDNSWNIEHLRRFY 2016
Score = 50.1 bits (118), Expect = 0.001
Identities = 45/197 (22%), Positives = 85/197 (42%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 432 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDSKAMANYLPVALADSARSWLHG 491
Query: 278 LK--GVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L + ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 492 LPRGTIESWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 551
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 552 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 604
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 605 SGPSWKPNKGTPATGGG 621
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 778 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 837
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 838 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 892
>ref|NP_918169.1| putative gag-pol polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2012
Score = 703 bits (1815), Expect = 0.0
Identities = 396/1080 (36%), Positives = 596/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 972 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 1031
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ MCVDY DLNK+ PKD F LP ID +VD+TA C++ SF+
Sbjct: 1032 EVLHPDWLANPVLVRKKTGQWLMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFL 1091
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1092 DCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1151
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 1152 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 1211
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K +W +
Sbjct: 1212 ANPEKVTAILNMKSPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDSFRWGPE 1271
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPP+L P PL++Y++ + VL +++E G K + IY
Sbjct: 1272 AQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIY 1331
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H+ +++ P+ I GR
Sbjct: 1332 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREANGR 1390
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A + +C E
Sbjct: 1391 IAKWALELMSLDISFKPRISIKSQALADFVA----------------------EWTECQE 1428
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
E + W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1429 DTPAENME---HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1485
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1486 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1545
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S H P + + + A ++ D W
Sbjct: 1546 VLRHNNEAADRLANFGSKREAAPSDVFVEHLYSPTVPHKDATQAAGAHDVAMVEAD--WR 1603
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
+ FL +Q+ P +DK ++S R + L+E LYK++ G+L RCV E
Sbjct: 1604 EPLIRFLTSQELP-----QDKDEAERISRRSKLYVLHEAELYKKSPSGILQRCVSLEEGR 1658
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1659 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1718
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1719 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1777
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNGT + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1778 F-INIVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1836
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1837 LQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTSSRATGQSPFFLVYGAEAMLPSEVEFE 1896
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1897 SLRF---RNFREERYEEDRVDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVG 1953
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1954 DLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2011
Score = 50.1 bits (118), Expect = 0.001
Identities = 46/197 (23%), Positives = 85/197 (42%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 427 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 486
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 487 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 546
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 547 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 599
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 600 SGPSWKQNKGTPATGGG 616
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 773 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 832
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 833 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 887
>ref|XP_475587.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46391119|gb|AAS90646.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1991
Score = 702 bits (1813), Expect = 0.0
Identities = 393/1103 (35%), Positives = 610/1103 (54%), Gaps = 56/1103 (5%)
Query: 1217 KKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQK 1276
++EI++ A+ + ++ I ++ DIFAW DMPG+ +V+EH L +K + P+KQ+
Sbjct: 929 REEIRLAATTESAL----ITFLQNNKDIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQR 984
Query: 1277 LRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKAS 1336
LRR D IKEE+ K + AGF+ +P WLAN V V KK G+ RMCVDY DLNK+
Sbjct: 985 LRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSC 1044
Query: 1337 PKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMP 1396
PKD F LP ID +VD+TA C++ SF+D +SGY+QIR+ D KTSFITP+GA+CYV MP
Sbjct: 1045 PKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMP 1104
Query: 1397 FGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLR 1456
FGL NAGATYQR + + F I + +E YVDD++VK+ +++ + L + F +R ++++
Sbjct: 1105 FGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLILDLEETFASIRAFRMK 1164
Query: 1457 LNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRF 1516
LNP KCTFGV SGKLLGF+VS +GI+ +P+KV AI M P T+K V+ G + +SRF
Sbjct: 1165 LNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRF 1224
Query: 1517 ISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVL 1576
+S + P FKLL+K +W + QKAF+ K+ L +PP+L P PL++Y++
Sbjct: 1225 VSRLGERGMPFFKLLKKTDNFQWGPEAQKAFEDFKQLLTKPPVLASPHPQEPLLLYVSAT 1284
Query: 1577 EDSMGCVL-GQQDETG---KKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYM 1632
+ VL +++E G K + IY++S+ D ++RY ++K + ++L HY
Sbjct: 1285 SQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQKLLYGILITTRKLSHYF 1344
Query: 1633 INHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQP 1692
H+ +++ P+ + GRIA+W + L DI ++ + ++K LA+ +A
Sbjct: 1345 QGHSVTVVTSF-PLGDVLHNREANGRIAKWALELMSLDISFKPRTSIKSQALADFVA--- 1400
Query: 1693 IEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLIT 1752
++ + D P +++ W + FDG+ + G+G G VLI+
Sbjct: 1401 --EWTECQEDTPVEKI--------------------EHWTMHFDGSKRLSGTGAGVVLIS 1438
Query: 1753 PKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQIKGEWET 1812
P G + + + F ++N+ EYEA + G+ AI L IK++++ GDS LV+NQ+ EW
Sbjct: 1439 PTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSC 1498
Query: 1813 RHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVN------GHNTVP 1866
++ YR R+L F+ +EL HV R N+ AD LA S H P
Sbjct: 1499 IDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKREAAPSDVFVEHLYSP 1558
Query: 1867 VINVQFLDRPAYVFVAEAIDDDKPWYHDIQVFLQTQKYPPGASNKDKKTLRKLSSRFFLN 1926
+ + + A ++ D W + FL +Q+ P ++ + + S + ++
Sbjct: 1559 TVPHKDATQVADTHDIAMVEAD--WREPLIRFLTSQELPQYKDEAER--ISRRSKLYVMH 1614
Query: 1927 EDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHAD 1986
E LYK++ G+L RCV E +L+ +IH G G H+ + K R G++W T +D
Sbjct: 1615 EAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSD 1674
Query: 1987 CYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAI 2046
+ C CQ +A +IH+P L I WPF++WG+DM+G + KA G+ + +AI
Sbjct: 1675 ADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFMAI 1733
Query: 2047 DYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKI 2106
D F+KW+EA +T F N++ R+GVPNRIITDNGT + K+ C+DF I
Sbjct: 1734 DKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGI 1792
Query: 2107 QHHNSSPYRPQMNGAVEAANKNI-----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTS 2161
+ +S P NG VE AN I ++ ++ W LP L+ RT+ +
Sbjct: 1793 KICYASVAHPMSNGQVERANGMILQRIKARVFDRLKPYAGKWVSQLPSVLWSLRTTPSRA 1852
Query: 2162 TGATPFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHG 2221
TG +PF LVYG EA+LP EVE SLR E + + R D L+ +EE R AAL
Sbjct: 1853 TGQSPFFLVYGAEAMLPSEVEFESLRF---RNFREERYEEDRVDDLHRLEEVREAALIQS 1909
Query: 2222 QLYQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGAL 2281
Y +++ ++ V R F GDLVL+ I++ + R K +P +EGP+++ G+
Sbjct: 1910 ARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSY 1967
Query: 2282 ILTNMDGEELPRPVNSDAVKKYF 2304
L DG + N + +++++
Sbjct: 1968 RLKREDGTLVDNSWNIEHIRRFY 1990
Score = 50.8 bits (120), Expect = 5e-04
Identities = 32/128 (25%), Positives = 60/128 (46%), Gaps = 5/128 (3%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 778 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKSSNAPFHGVIPGLSATPL 837
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRG 1003
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 838 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPR 897
Query: 1004 KLITVSGE 1011
++T+ +
Sbjct: 898 GVLTLRSD 905
Score = 49.3 bits (116), Expect = 0.002
Identities = 54/221 (24%), Positives = 92/221 (41%), Gaps = 19/221 (8%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 433 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 492
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES +EY +RF ++ +I
Sbjct: 493 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLREYIRRFSEQRNKISD 552
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 553 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 605
Query: 392 GGSSGVSSNGNKKFGNGY-----PKRNAQE-VGMVAHGGPQ 426
G S N G G KR +E V AH Q
Sbjct: 606 SGPSWKPKNTPATGGGGSNNHKDRKRKPEELVATAAHSSRQ 646
>ref|XP_473950.1| OSJNBa0053K19.16 [Oryza sativa (japonica cultivar-group)]
gi|38344177|emb|CAE03508.2| OSJNBa0053K19.16 [Oryza
sativa (japonica cultivar-group)]
Length = 2010
Score = 701 bits (1809), Expect = 0.0
Identities = 395/1080 (36%), Positives = 597/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 970 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 1029
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLNK+ PKD F LP ID +VD+TA ++ SF+
Sbjct: 1030 EVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGRELLSFL 1089
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1090 DCYSGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1149
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKL+GF+VS +GI+
Sbjct: 1150 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLVGFMVSHRGIQ 1209
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K +W +
Sbjct: 1210 ANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDNFQWGPE 1269
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPP+L P PL++Y++ + VL +++E G K + IY
Sbjct: 1270 AQKAFEDFKKLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIY 1329
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H+ +++ P+ I GR
Sbjct: 1330 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREANGR 1388
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A + +C E
Sbjct: 1389 IAKWALELMSLDISFKPRISIKSQALADFVA----------------------EWTECQE 1426
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
E + W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1427 DTPAENME---HWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1483
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1484 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1543
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S V H P + + + A ++ D W
Sbjct: 1544 VLRHNNEAADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAMVETD--WR 1601
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
+ FL +Q+ P +DK ++S R + ++E LYK++ G+L RCV E
Sbjct: 1602 EPLIRFLTSQELP-----QDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGR 1656
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1657 QLLQDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1716
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1717 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1775
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNGT + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1776 F-INIVHRFGVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1834
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1835 LQGIKARVFDRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1894
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1895 SLRF---RNFREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVG 1951
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1952 DLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2009
Score = 51.6 bits (122), Expect = 3e-04
Identities = 46/197 (23%), Positives = 86/197 (43%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 425 VDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 484
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 485 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 544
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K + F + E+A+ AV+ S
Sbjct: 545 ITDDVIIAAFTKGIRHEELVGKFGRKPPMTVKLMFE-------KANEYAKAEDAVTASKQ 597
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 598 SGPSWKQNKGTPATGGG 614
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 771 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPHSELKPSNAPFHGVIPGLSATPL 830
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 831 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 885
>ref|XP_475120.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|46063437|gb|AAS79740.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1756
Score = 700 bits (1807), Expect = 0.0
Identities = 394/1080 (36%), Positives = 597/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 716 DIFAWKPSDMPGIPREVIEHSLHVKEDAKPIKQRLRRFVQDRKDAIKEELTKLLAAGFIK 775
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLNK PKD F LP ID +VD+TA C++ SF+
Sbjct: 776 EVLHPDWLANPVLVRKKTGQWRMCVDYTDLNKCCPKDPFGLPRIDQVVDSTAGCELLSFL 835
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D +SGY+QIR+ D KTSFITP GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 836 DCYSGYHQIRLKESDCLKTSFITPIGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 895
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD+++K+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 896 EAYVDDVVIKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 955
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+KV AI M P T+K V+ G + +SRF+S + P FKLL+K +W +
Sbjct: 956 ANPEKVTAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPE 1015
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K++L EPP+L P PL++Y++ + VL +++E G K + IY
Sbjct: 1016 AQKAFEDFKKFLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIY 1075
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY H+ +++ P+ I GR
Sbjct: 1076 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREANGR 1134
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A ++ + D P +++
Sbjct: 1135 IAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPAEKM---------- 1179
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1180 ----------EHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1229
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1230 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1289
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S V H P + + + A ++ D W
Sbjct: 1290 VLRHNNEAADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQIAGTHDVALVEAD--WR 1347
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
FL +Q+ P +DK ++S R + ++E LYK++ G+L RCV E
Sbjct: 1348 EPFIRFLTSQELP-----QDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGR 1402
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1403 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1462
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1463 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1521
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNG + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1522 F-INIVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1580
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1581 LQGIKARVFDRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1640
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1641 SLRF---RNFREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVG 1697
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1698 DLVLRKIQTTR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 1755
Score = 50.4 bits (119), Expect = 7e-04
Identities = 31/128 (24%), Positives = 61/128 (47%), Gaps = 5/128 (3%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 517 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 576
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSRG 1003
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 577 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMKMPGPR 636
Query: 1004 KLITVSGE 1011
+++++ +
Sbjct: 637 RVLSLRSD 644
Score = 47.8 bits (112), Expect = 0.005
Identities = 44/191 (23%), Positives = 82/191 (42%), Gaps = 13/191 (6%)
Query: 224 FKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMGLKGVT- 282
FK EKY G T P++ LT+Y + A + + + +L A +W GL T
Sbjct: 177 FKPTGIEKYDGTTNPESWLTVYGLAIRAAGGDSKAMANYLPVALADSARSWLHGLPRGTI 236
Query: 283 -TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRPPLDERE 341
++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I D+
Sbjct: 237 RSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISDITDDVI 296
Query: 342 LSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGV 397
++ + +E L + K F + E+A+ AV+ S G S
Sbjct: 297 IAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQSGPSWK 349
Query: 398 SSNGNKKFGNG 408
+ G G G
Sbjct: 350 PNKGTPATGGG 360
>ref|XP_474845.1| OSJNBa0035O13.3 [Oryza sativa (japonica cultivar-group)]
gi|21741410|emb|CAD40114.1| OSJNBa0035O13.3 [Oryza sativa
(japonica cultivar-group)]
Length = 2008
Score = 697 bits (1798), Expect = 0.0
Identities = 413/1181 (34%), Positives = 626/1181 (52%), Gaps = 92/1181 (7%)
Query: 1142 DIPSVTPCSKLNISESVEHSDPMLSPNFEVPVYEAVAEEDEEIPNEIKWMLEQERKTIQP 1201
DI C K + + H+ M S ++ + A A E E + E E KT +
Sbjct: 901 DIKQAVTCDKESCD--MAHTHEMASAREDIRLAAATASEGEIPVTKTSKSGESEAKTKK- 957
Query: 1202 HQEEIEIINLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVE 1261
I L + K + + +I ++ DIFAW DMPG+ +V+E
Sbjct: 958 -------IPLDPSDPTK----------TAESALITFLQNNKDIFAWKPSDMPGIPREVIE 1000
Query: 1262 HRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDG 1321
H L +K + P+KQ+LRR D IKEE+ K + AGF+ +P WLAN V V KK G
Sbjct: 1001 HSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLANPVLVRKKTG 1060
Query: 1322 KIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKT 1381
+ RMCVDY DLNK+ PKD F LP ID +VD+TA C++ SF+D +SGY+QIR+ D KT
Sbjct: 1061 QWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQIRLKESDCLKT 1120
Query: 1382 SFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVE 1441
SFITP+GA+CYV MPFGL NAGATYQR + + F I + +E YVDD++VK+ +++ +
Sbjct: 1121 SFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVVKTKQKDDLIS 1180
Query: 1442 YLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEK 1501
L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+ +P+KV AI M P T+K
Sbjct: 1181 DLEETFVSIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAILNMKPPSTQK 1240
Query: 1502 QVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPILV 1561
V+ G + +SRF+S + P FKLL+K +W + QKAF+ K+ L EPP+L
Sbjct: 1241 DVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDNFQWGPEAQKAFEDFKKLLTEPPVLA 1300
Query: 1562 PPVDGRPLIMYLTVLEDSMGCVLG-QQDETG---KKEHVIYYLSKKFTDCESRYSVLEKT 1617
P PL++Y++ + VL +++E G K + IY++S+ D ++RY ++K
Sbjct: 1301 SPHPQEPLLLYVSATSQVVSTVLAVEREEEGHVQKVQRPIYFVSEVLADSKTRYPQVQK- 1359
Query: 1618 CCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQK 1677
+ H+ +++ P+ I GRIA+W + L DI ++ +
Sbjct: 1360 -------------LLYGHSVTVVTSF-PLGDILHNREANGRIAKWALELMSLDISFKPRI 1405
Query: 1678 AVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDG 1737
++K LA+ +A + +C E E + W + FDG
Sbjct: 1406 SIKSQALADFVA----------------------EWTECQEDTPAENME---HWTMHFDG 1440
Query: 1738 AVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYG 1797
+ + G+G G VLI+P G + + + F ++N+ EYEA + G+ AI L IK++++ G
Sbjct: 1441 SKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEALLHGLRIAISLGIKRLIVRG 1500
Query: 1798 DSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMI 1857
DS LV+NQ+ EW ++ YR R+L F+ +EL HV R N+ AD LA S
Sbjct: 1501 DSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAADRLANFGSKR 1560
Query: 1858 NVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWYHDIQVFLQTQKYPPGASNK 1911
V H P + + + A ++ D W + FL +Q+ P +
Sbjct: 1561 EVAPSDVFVEHLYTPTVPHKDTTQVAGTHDVAVVETD--WREPLIRFLTSQELP-----Q 1613
Query: 1912 DKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHA 1968
DK ++S R + ++E LYK++ G+L RCV E +L+ +IH G G H+
Sbjct: 1614 DKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSGICGNHAAART 1673
Query: 1969 MAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMI 2028
+ K R G++W T +D + C CQ +A +IH+P L I WPF++WG+DM+
Sbjct: 1674 IVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSWPFAVWGLDMV 1733
Query: 2029 GRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDN 2088
G + KA G+ + VAID F+KW+EA +T F N++ R+GVPNRIITDN
Sbjct: 1734 GPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRFGVPNRIITDN 1791
Query: 2089 GTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI-----KKIIQKMVVTYKDW 2143
GT + K+ C+DF I+ +S P NG VE AN I ++ ++ W
Sbjct: 1792 GTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVFDRLKPYAGKW 1851
Query: 2144 HEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSR 2203
+ LP L+ RT+ +TG +PF LVYG EA+LP EVE SLR E + + R
Sbjct: 1852 VQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRF---RNFREERYEEDR 1908
Query: 2204 YDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWT 2263
D L+ +EE R AAL Y +++ ++ V R F GDLVL+ I++ + R K +
Sbjct: 1909 VDDLHRLEEAREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQTTR--DRHKLS 1966
Query: 2264 PNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
P +EGP+++ G+ L DG + N + +++++
Sbjct: 1967 PLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2007
Score = 49.7 bits (117), Expect = 0.001
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 774 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 833
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 834 GQITLPVTFGTRENFRTENISFEVADFETAYHAILGRPALAKFMAVPHYTYMMMK 888
Score = 48.5 bits (114), Expect = 0.003
Identities = 46/197 (23%), Positives = 84/197 (42%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P + LT+Y + A + + + +L A +W G
Sbjct: 428 VDWPAGFKPTGIEKYDGTTNPGSWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHG 487
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 488 LPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 547
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 548 ITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFE-------KANEYAKAEDAVTASKQ 600
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 601 SGPSWKQNKGTPATGGG 617
>ref|XP_463537.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 2001
Score = 696 bits (1797), Expect = 0.0
Identities = 394/1101 (35%), Positives = 612/1101 (54%), Gaps = 50/1101 (4%)
Query: 1209 INLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHRLPLKP 1268
+ L K I IG +L+ I++ ++++++E + +FAWS ++ G+D ++EH L +K
Sbjct: 945 VQLDEHNPGKFILIGENLEKHIEEEILKVVKENMAVFAWSPDELQGVDRSLIEHNLAIKS 1004
Query: 1269 ECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDGKIRMCVD 1328
P KQKLRR D K E+ K + A + +P+WLAN V V K +GK RMC+D
Sbjct: 1005 GYKPKKQKLRRMSTDRQQAAKIELEKLLKAKVIREVMHPEWLANPVLVKKANGKWRMCID 1064
Query: 1329 YRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSFITPWG 1388
+ DLNKA PKD+FPLP ID LVD TA C++ SF+D +SGY+Q+ M ED EKTSFITP+G
Sbjct: 1065 FTDLNKACPKDDFPLPRIDQLVDATAGCELMSFLDAYSGYHQVFMVKEDEEKTSFITPFG 1124
Query: 1389 AFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVEYLLKMFQ 1448
+C++ MPFGL NAGAT+ R + K+ + + +E Y+DD++VKS H + L + F+
Sbjct: 1125 TYCFIRMPFGLKNAGATFARLIGKVLAKQLGRNVEAYIDDIVVKSKQAFTHGKDLQETFE 1184
Query: 1449 RLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEKQVRGFLG 1508
LRK ++LNP KC FGVR+GKLLGF+VS++GIE +PDK+ AI +M PK ++V+ G
Sbjct: 1185 NLRKCSVKLNPEKCVFGVRAGKLLGFLVSKRGIEANPDKIAAIHQMEPPKNTREVQRLTG 1244
Query: 1509 RLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIKEYLLEPPILVPPVDGRP 1568
R+ +SRF+S P FK LR +W +CQ+AFD +K+YL E P L P G+P
Sbjct: 1245 RMASLSRFLSKSAEKGLPFFKTLRGANTFEWTAECQQAFDDLKKYLHEMPTLASPPKGQP 1304
Query: 1569 LIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCESRYSVLEKTCCALAWAAKRL 1628
L+MY+ ++ VL Q++E ++ +Y++S+ ++RYS +EK A+ A+++L
Sbjct: 1305 LLMYVAATPATVSAVLVQEEE--NRQVPVYFVSEALQGPKTRYSEVEKLIYAIVMASRKL 1362
Query: 1629 RHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQKAVKGSILAEHL 1688
RHY ++H + S PI + + GRIA+W M L +D++Y ++ A+K +LA+ +
Sbjct: 1363 RHYFLSHDITIPSAY-PIGEVLTNKEVAGRIAKWAMELLPFDLKYISRTAIKSQVLADFV 1421
Query: 1689 AHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDGAVNVYGSGIGA 1748
A ++ P + + + + + W + DGA N G+G A
Sbjct: 1422 A-----EWTPNEVE--------------------QQEEVKKPWIVFSDGACNAAGAGAAA 1456
Query: 1749 VLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDSALVINQIKG 1808
V+ TP + ++ +L F TNN EYE ++ + +A L +++++ DS LV
Sbjct: 1457 VVKTPMKQTLKYSVQLVFPSTNNTAEYEGVLLAMRKARALGARRLIVKTDSKLVAGHFSK 1516
Query: 1809 EWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINVNGHNTVPVI 1868
+E + + Y + AR F + + + R+EN AD LA ++ P+
Sbjct: 1517 SFEAKEETMAKYLEEARLNEKHFLGITVKAITREENGEADELAKAAA-------TGQPLE 1569
Query: 1869 NVQF--LDRPAYVFVAEA-IDDDKPWYHDIQVFLQTQKYPPGASNKDKKTLRKLSSRFFL 1925
N F + +P+Y A I + W I +L + + P ++ K ++ +S ++ +
Sbjct: 1570 NSFFDIITQPSYEKKEVACIQREGDWREPILKYLVSAQLP--EKEEEAKRIQLMSKKYKV 1627
Query: 1926 NEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHA 1985
E LYK LL+CV + E K++ EIHEG G H ++A K++R G YW T+
Sbjct: 1628 VEGQLYKSGVTAPLLKCVTREEGMKMVVEIHEGLCGAHQAPWSVASKVIRQGIYWPTIMK 1687
Query: 1986 DCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVA 2045
D + K C CQ + PP L I WPF WGID++G + P+A RF++VA
Sbjct: 1688 DTEKYIKTCKACQKFGPMTKAPPKELQPIPPVWPFYRWGIDIVGPL-PRAKGDLRFVIVA 1746
Query: 2046 IDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFK 2105
I+YF++W+EA + A +T V +F+ N+I R+G+P I+ DNG + +++C
Sbjct: 1747 IEYFSRWIEAEAVARITSAAVQKFVWKNIICRFGIPKEIVCDNGKQFESGKFQDMCKGLN 1806
Query: 2106 IQHHNSSPYRPQMNGAVEAANKNIKKIIQKMV--VTYKDWHEMLPYALYGYRTSVRTSTG 2163
+Q + +S PQ NG VE AN I + I+K + W E L L+ RT+V STG
Sbjct: 1807 LQINFASVGHPQTNGVVERANGKIMEAIKKRLEGSAKGKWPEDLLSVLWALRTTVVRSTG 1866
Query: 2164 ATPFSLVYGMEAVLPVEVEIPSLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQL 2223
TPF LVYG EA+ P EV S R++ + + E R L +++E R+ AL
Sbjct: 1867 MTPFRLVYGDEAMTPSEVGAHSPRMIFDQKDEE-----GREITLEMLDEIRVEALEKMAS 1921
Query: 2224 YQSRMKQAFDKGVHPREFKEGDLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALIL 2283
Y K +++ V R +EGDLVLK K GK +EGP++VK+ GA L
Sbjct: 1922 YTEGTKSYYNQKVKTRPIEEGDLVLK--KVLNEVAVGKLESKWEGPFIVKKKTETGAFKL 1979
Query: 2284 TNMDGEELPRPVNSDAVKKYF 2304
+DGEEL N+ ++KK++
Sbjct: 1980 AYLDGEELKHTWNAVSLKKFY 2000
Score = 46.2 bits (108), Expect = 0.013
Identities = 35/145 (24%), Positives = 66/145 (45%), Gaps = 4/145 (2%)
Query: 871 LTFSDEDLPAEGNKHNRALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLR 930
++F ED H L++SV + VLID GS+ +V+ ++ E L
Sbjct: 758 ISFGPEDAEGILFPHQDPLVVSVEIAQCEVQRVLIDGGSSADVLFYDAFKKMQIPEDRLT 817
Query: 931 LSKVTVRAFDGTRRSVYGEVDLPISVGP----HEFQVTFQVMEIQASFSCLLGRPWIHDA 986
+ V ++ F G + G++ L + G + ++ F V+++ ++ +LGR I+
Sbjct: 818 NAGVPLQGFGGQQVHAIGKISLQVVFGKGTNVRKEEIVFDVVDMPYQYNAILGRSTINIF 877
Query: 987 GAVTSTLHQKLKFVSRGKLITVSGE 1011
A+ + +K +ITV GE
Sbjct: 878 EAIIHHNYICMKLPGLRGVITVRGE 902
Score = 38.1 bits (87), Expect = 3.6
Identities = 30/122 (24%), Positives = 53/122 (42%), Gaps = 4/122 (3%)
Query: 221 PPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMGLKG 280
P FK+ KY+GDT P +L +Y + A + L G A +W+ +
Sbjct: 353 PINFKLSNITKYKGDTDPNEYLRVYETAVEAAGGDDTTKAKILPTMLEGVALSWYTTIPP 412
Query: 281 VT--TFEQLAEAFMQQYKYNTYLAPSR-KELQSLTQKDKESFKEYAQRFIQKAAQIRPPL 337
+T ++E + + F + Y P +L ++ Q E+ + + +F + QIR
Sbjct: 413 MTIYSWEHMRDTFRAGF-IGAYEEPKEADDLYAMKQLPGETLRSFIVKFSRVRCQIRHVD 471
Query: 338 DE 339
DE
Sbjct: 472 DE 473
>gb|AAP54912.1| gag-pol precursor [Oryza sativa (japonica cultivar-group)]
gi|37536646|ref|NP_922625.1| gag-pol precursor [Oryza
sativa (japonica cultivar-group)]
gi|13876521|gb|AAK43497.1| gag-pol precursor [Oryza
sativa (japonica cultivar-group)]
Length = 2017
Score = 694 bits (1791), Expect = 0.0
Identities = 392/1080 (36%), Positives = 596/1080 (54%), Gaps = 58/1080 (5%)
Query: 1243 DIFAWSYKDMPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLV 1302
DIFAW DMPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+
Sbjct: 977 DIFAWKPSDMPGIPREVIEHSLYVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIK 1036
Query: 1303 TSEYPQWLANIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFM 1362
+P WLAN V V KK G+ RMCVDY DLN + PKD F LP ID +VD+TA C++ SF+
Sbjct: 1037 EVLHPDWLANPVLVRKKTGQWRMCVDYTDLNISCPKDPFGLPRIDQVVDSTAGCELLSFL 1096
Query: 1363 DGFSGYNQIRMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEI 1422
D + GY+QIR+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +
Sbjct: 1097 DCYLGYHQIRLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNV 1156
Query: 1423 EVYVDDMIVKSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIE 1482
E YVDD++VK+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+
Sbjct: 1157 EAYVDDVVVKTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQ 1216
Query: 1483 VDPDKVRAIREMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDD 1542
+P+K+ AI M P T+K V+ G + +SRF+S + P FKLL+K +W D
Sbjct: 1217 ANPEKITAILNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDDFQWGPD 1276
Query: 1543 CQKAFDQIKEYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIY 1598
QKAF+ K+ L EPP+L P PL++Y+ + VL +++E G K + IY
Sbjct: 1277 AQKAFEDFKKLLTEPPVLASPHPQEPLLLYIAAASQVVSTVLVVEREEEGHVQKVQRPIY 1336
Query: 1599 YLSKKFTDCESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGR 1658
++S+ D ++RY ++K + ++L HY +H+ +++ + I GR
Sbjct: 1337 FVSEVLADSKTRYPQVQKLLYGILITTRKLSHYFQSHSVTVVTSFS-LGDILHNREANGR 1395
Query: 1659 IARWQMLLSEYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDE 1718
IA+W + L DI ++ + ++K LA+ +A ++ + D P +++
Sbjct: 1396 IAKWALELMSLDISFKPRISIKSQALADFVA-----EWTECQEDTPAEKM---------- 1440
Query: 1719 PVFGEGPDPESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEAC 1778
W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA
Sbjct: 1441 ----------EHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSASHNVAEYEAL 1490
Query: 1779 IMGIEEAIDLRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHH 1838
+ G+ AI L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL H
Sbjct: 1491 LHGLRIAISLGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSH 1550
Query: 1839 VPRDENQMADALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWY 1892
V R N+ AD LA S V H P + + + A ++ D W
Sbjct: 1551 VLRHNNEAADRLANFGSKREVAPSDVFVEHLYTPTVPHKDTTQIAGTHDVALVEAD--WR 1608
Query: 1893 HDIQVFLQTQKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAE 1949
+ FL +Q+ P +DK ++S R + ++E LYK++ G+L RCV E
Sbjct: 1609 EPLIRFLTSQELP-----QDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGR 1663
Query: 1950 KLMCEIHEGSFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPS 2009
+L+ +IH G G H+ + K R G++W T +D + C CQ +A +IH+P
Sbjct: 1664 QLLKDIHSGICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQ 1723
Query: 2010 MLNVISSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRF 2069
L I WPF++WG+DM+G + KA G+ + VAID F+KW+EA +T F
Sbjct: 1724 ELQTIPLSWPFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDF 1782
Query: 2070 IKNNLISRYGVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI 2129
N++ R+GVPNRIITDNG + K+ C+DF I+ +S P NG VE AN I
Sbjct: 1783 F-INIVHRFGVPNRIITDNGRQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMI 1841
Query: 2130 -----KKIIQKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIP 2184
++ ++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE
Sbjct: 1842 LQGIKARVFDRLKPYAGKWVKQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFE 1901
Query: 2185 SLRVLMEAELSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEG 2244
SLR E + + R D L+ +EE R AAL Y +++ ++ V R F G
Sbjct: 1902 SLRF---RNYREERYEEDRVDDLHRLEEVREAALIQSARYLQGLRRYHNRNVRSRAFLVG 1958
Query: 2245 DLVLKCIKSFQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
DLVL+ I++ + R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 1959 DLVLRKIQTTR--DRHKLSPLWEGPFIISDVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 2016
Score = 49.7 bits (117), Expect = 0.001
Identities = 46/197 (23%), Positives = 85/197 (42%), Gaps = 13/197 (6%)
Query: 218 VNIPPKFKMPVFEKYQGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWFMG 277
V+ P FK EKY G T P++ LT+Y + A + + + +L A +W G
Sbjct: 432 VDWPAGFKPTGIEKYDGTTNPESWLTIYGLAIRAAGGDSKAMANYLPVALADSARSWLHG 491
Query: 278 LKGVT--TFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQIRP 335
L T ++ +L + F+ ++ ++ +L ++ QK ES ++Y +RF ++ +I
Sbjct: 492 LPRGTIGSWAELRDHFIANFQGTFEHPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISD 551
Query: 336 PLDERELSE----LFYETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSS 391
D+ ++ + +E L + K F + E+A+ AV+ S
Sbjct: 552 ITDDVIIAAFTKGIRHEELVGKFGRKPPKTVKLMFE-------KANEYAKAEDAVTASKQ 604
Query: 392 GGSSGVSSNGNKKFGNG 408
G S + G G G
Sbjct: 605 SGPSWKPNKGTPAKGGG 621
Score = 48.9 bits (115), Expect = 0.002
Identities = 31/115 (26%), Positives = 55/115 (46%), Gaps = 5/115 (4%)
Query: 889 LLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVR-AFDGTRRSVY 947
L++ + R L LID GSALN++ TLD + + L+ S G +
Sbjct: 778 LVLDPVVRNVKLRRTLIDGGSALNILFAKTLDDMQIPRSELKPSNAPFHGVIPGLSATPL 837
Query: 948 GEVDLPISVGPHE----FQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLK 998
G++ LP++ G E ++F+V + + ++ +LGRP + AV + +K
Sbjct: 838 GQITLPVTFGTRENFRTENISFEVDDFETAYHAILGRPALAKFMAVPHYTYMMMK 892
>dbj|BAA84458.1| GAG-POL precursor [Oryza sativa (japonica cultivar-group)]
Length = 1032
Score = 691 bits (1782), Expect = 0.0
Identities = 389/1071 (36%), Positives = 593/1071 (55%), Gaps = 58/1071 (5%)
Query: 1252 MPGLDPDVVEHRLPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLA 1311
MPG+ +V+EH L +K + P+KQ+LRR D IKEE+ K + AGF+ +P WLA
Sbjct: 1 MPGIPREVIEHSLHVKEDAKPIKQRLRRFAQDRKDAIKEELTKLLAAGFIKEVLHPDWLA 60
Query: 1312 NIVPVPKKDGKIRMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQI 1371
N V V KK G+ RMCVDY DLNK+ PKD F LP ID +VD+TA C++ SF+D +SGY+QI
Sbjct: 61 NPVLVRKKTGQWRMCVDYTDLNKSCPKDPFGLPRIDQVVDSTAGCELLSFLDCYSGYHQI 120
Query: 1372 RMAPEDREKTSFITPWGAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIV 1431
R+ D KTSFITP+GA+CYV MPFGL NAGATYQR + + F I + +E YVDD++V
Sbjct: 121 RLKESDCLKTSFITPFGAYCYVTMPFGLKNAGATYQRMIQRCFSTQIGRNVEAYVDDVVV 180
Query: 1432 KSGTEEEHVEYLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAI 1491
K+ +++ + L + F +R ++++LNP KCTFGV SGKLLGF+VS +GI+ +P+KV AI
Sbjct: 181 KTKQKDDLISDLEETFASIRAFRMKLNPEKCTFGVPSGKLLGFMVSHRGIQANPEKVTAI 240
Query: 1492 REMPVPKTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKDQGVKWNDDCQKAFDQIK 1551
M P T+K V+ G + +SRF+S + P FKLL+K +W + QKAF+ K
Sbjct: 241 LNMKPPSTQKDVQKLTGCMAALSRFVSRLGERGMPFFKLLKKTDNFQWGPEAQKAFEDFK 300
Query: 1552 EYLLEPPILVPPVDGRPLIMYLTVLEDSMGCVL-GQQDETG---KKEHVIYYLSKKFTDC 1607
+ L EPP+L P PL++Y++ + VL +++E G K + IY++S+ D
Sbjct: 301 KLLTEPPVLASPHPQEPLLLYVSATSQVVSTVLVVEREEEGHVQKVQRPIYFVSEVLADS 360
Query: 1608 ESRYSVLEKTCCALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLS 1667
++RY ++K + ++L HY H+ +++ P+ I GRIA+W + L
Sbjct: 361 KTRYPQVQKLLYGILITTRKLSHYFQGHSVTVVTSF-PLGDILHNREANGRIAKWALELM 419
Query: 1668 EYDIEYRTQKAVKGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDP 1727
DI ++ + ++K LA+ +A ++ + D P +++
Sbjct: 420 SLDISFKPRISIKSQALADFVA-----EWTECQEDTPAEKM------------------- 455
Query: 1728 ESEWGLIFDGAVNVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAID 1787
W + FDG+ + G+G G VLI+P G + + + F ++N+ EYEA + G+ AI
Sbjct: 456 -EHWTMHFDGSKRLSGTGAGVVLISPTGERLSYVLWIHFSTSHNVAEYEALLHGLRIAIS 514
Query: 1788 LRIKKIVIYGDSALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMA 1847
L IK++++ GDS LV+NQ+ EW ++ YR R+L F+ +EL HV R N+ A
Sbjct: 515 LGIKRLIVRGDSQLVVNQVMKEWSCLDDNMMAYRQEVRKLEDKFDGLELSHVLRHNNEAA 574
Query: 1848 DALATLSSMINVN------GHNTVPVINVQFLDRPAYVFVAEAIDDDKPWYHDIQVFLQT 1901
D LA S H P + + + A ++ D W + FL +
Sbjct: 575 DRLANFGSKREAAPSDVFVEHLYTPTVPHKDTTQVADTHDIAMVEAD--WREPLIRFLTS 632
Query: 1902 QKYPPGASNKDKKTLRKLSSR---FFLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEG 1958
Q+ P +DK ++S R + ++E LYK++ G+L RCV E +L+ +IH G
Sbjct: 633 QELP-----RDKDEAERISRRSKLYVMHEAELYKKSPSGILQRCVSLEEGRQLLKDIHSG 687
Query: 1959 SFGTHSCGHAMAKKILRAGYYWITMHADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPW 2018
G H+ + K R G++W T +D + C CQ +A +IH+P L I W
Sbjct: 688 ICGNHAAARTIVGKAYRQGFFWPTAVSDADKIVRTCEGCQFFARQIHLPAQELQTIPLSW 747
Query: 2019 PFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRY 2078
PF++WG+DM+G + KA G+ + VAID F+KW+EA +T F N++ R+
Sbjct: 748 PFAVWGLDMVGPFK-KAVGGYTHLFVAIDKFSKWIEAKPVVTITADNARDFF-INIVHRF 805
Query: 2079 GVPNRIITDNGTNLNNNMMKELCDDFKIQHHNSSPYRPQMNGAVEAANKNI-----KKII 2133
GVPNRIITDNGT + K+ C+DF I+ +S P NG VE AN I ++
Sbjct: 806 GVPNRIITDNGTQFTGGVFKDFCEDFGIKICYASVAHPMSNGQVERANGMILQGIKARVF 865
Query: 2134 QKMVVTYKDWHEMLPYALYGYRTSVRTSTGATPFSLVYGMEAVLPVEVEIPSLRVLMEAE 2193
++ W + LP L+ RT+ +TG +PF LVYG EA+LP EVE SLR
Sbjct: 866 DRLKPYAGKWVQQLPSVLWSLRTTPSRATGQSPFFLVYGAEAMLPSEVEFESLRF---RN 922
Query: 2194 LSEAEWCQSRYDQLNLIEEKRMAALCHGQLYQSRMKQAFDKGVHPREFKEGDLVLKCIKS 2253
E + + R D L+ +EE R AAL Y +++ ++ V R F GDLVL+ I++
Sbjct: 923 FREERYEEDRVDDLHRLEEVREAALIRSARYLQGLRRYHNRNVRSRAFLVGDLVLRKIQT 982
Query: 2254 FQPDPRGKWTPNYEGPYVVKRAFSGGALILTNMDGEELPRPVNSDAVKKYF 2304
+ R K +P +EGP+++ G+ L DG + N + +++++
Sbjct: 983 TR--DRHKLSPLWEGPFIISEVTRPGSYRLKREDGTLVDNSWNIEHLRRFY 1031
>gb|AAX95607.1| RNase H, putative [Oryza sativa (japonica cultivar-group)]
Length = 2282
Score = 688 bits (1776), Expect = 0.0
Identities = 389/989 (39%), Positives = 562/989 (56%), Gaps = 65/989 (6%)
Query: 1204 EEIEIINLGTEEDKKEIKIGASLDVSIKKRVIELIREYVDIFAWSYKDMPGLDPDVVEHR 1263
+++E I++G + + I +L + ++IEL++E+ D FAW Y +MPGL +VEHR
Sbjct: 1334 DDLEEIDIGPGDRPRPTFICKNLSTEFRAKLIELLKEFRDCFAWEYYEMPGLSRSIVEHR 1393
Query: 1264 LPLKPECPPVKQKLRRSHPDMALKIKEEVRKQIDAGFLVTSEYPQWLANIVPVPKKDGKI 1323
LP+KP P +Q LRR DM +K E+++ DA F+ Y +W+++IVPV KK+
Sbjct: 1394 LPIKPGVRPHQQPLRRCKADMLEPVKVEIKRLYDACFIRPCRYAEWVSSIVPVIKKN--- 1450
Query: 1324 RMCVDYRDLNKASPKDNFPLPHIDVLVDNTAKCKVFSFMDGFSGYNQIRMAPEDREKTSF 1383
DLNKA+PKD +P+P D LVD + K+ S+MDG GYNQI MA ED KT+F
Sbjct: 1451 -------DLNKATPKDEYPMPVADQLVDAASGNKILSYMDGNVGYNQIFMAEEDIHKTAF 1503
Query: 1384 ITPW--GAFCYVVMPFGLINAGATYQRGMTKIFHDMIHKEIEVYVDDMIVKSGTEEEHVE 1441
P G F +VVM FGL +AGATYQR M I+HD+I +EVY+DD++VKS E+H+
Sbjct: 1504 RCPGAIGLFEWVVMTFGLKSAGATYQRAMNYIYHDLIGWLVEVYIDDVVVKSKEIEDHIA 1563
Query: 1442 YLLKMFQRLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPVPKTEK 1501
L K+F+R RKY L++NP KC FGV G+ LGF+V ++GIEV V AI+++ P+ +
Sbjct: 1564 DLRKVFERTRKYGLKMNPTKCAFGVSVGQFLGFLVHERGIEVTQRSVNAIKKIQPPENKT 1623
Query: 1502 QVRGFLGRLNYISRFISHMTATCGPIFKLLR--KDQGVKWNDDCQKAFDQIKEYLLEPPI 1559
+++ G++N++ RFIS+++ P LLR DQ W + QKA D IKEYL PP+
Sbjct: 1624 ELQEMNGKINFVRRFISYLSGRLEPFTPLLRLKADQQFTWGAEQQKALDDIKEYLSSPPV 1683
Query: 1560 LVPPVDGRPLIMYLTVLEDSMGCVLGQQDETGKKEHVIYYLSKKFTDCESRYSVLEKTCC 1619
L+PP G +YL+ E S+G VL Q E KE V++YLS++ D E+RYS +EK C
Sbjct: 1684 LIPPQKGISFRLYLSAGEKSIGSVLIQ--ELDGKERVVFYLSRRLLDVETRYSPMEKLCL 1741
Query: 1620 ALAWAAKRLRHYMINHTTWLISKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRTQKAV 1679
L ++ RL HY++++ +I K D IKY+ P L GR+ +W L+E+D+ Y + KA+
Sbjct: 1742 CLYFSCTRLSHYLLSNECTVICKADVIKYMLSAPILKGRVGKWIFSLTEFDLRYESPKAI 1801
Query: 1680 KGSILAEHLAHQPIEDYQPIKFDFPDKEVMYLKAKDCDEPVFGEGPDPESEWGLIFDGAV 1739
KG +A+ + CD+ + G W FDG+V
Sbjct: 1802 KGQAIADFIVEH------------------------CDDSI---GSVEVVPWTSFFDGSV 1834
Query: 1740 NVYGSGIGAVLITPKGTHIPFTARLRFDCTNNIVEYEACIMGIEEAIDLRIKKIVIYGDS 1799
+ GIG V+I+P+G F ++ TNN EYEA + G++ ++ I I GDS
Sbjct: 1835 CTHDCGIGLVIISPRGACFKFAYTIKPYATNNQAEYEAVLKGLQLLKEVEADAIEIMGDS 1894
Query: 1800 ALVINQIKGEWETRHPGLIPYRDYARRLLTFFNKVELHHVPRDENQMADALATLSSMINV 1859
LVI+Q+ GE+E + L+ Y + + L+ F V L HV R++N A+ LA +S
Sbjct: 1895 LLVISQLAGEYECKSDTLMVYNEKCQELMQEFRLVTLKHVSREQNIEANDLAQGAS---- 1950
Query: 1860 NGHNTVPVINVQFLDRPAYVFVAEAIDDDKPWYHDIQVFLQTQKYPPGASNKDKKTLRKL 1919
G+ P+I D A + A D W +D+ +L S + LR
Sbjct: 1951 -GYK--PMIK----DVKAEIAAITAGD----WRYDVHQYLHNP------SQSASRKLRYK 1993
Query: 1920 SSRFFLNEDVLYKRNFDGVLLRCVDKHEAEKLMCEIHEGSFGTHSCGHAMAKKILRAGYY 1979
+ ++ L +D LY R DGVLL+C+ +A+ + E+HEG GTH H M + RA Y+
Sbjct: 1994 ALKYTLLDDELYYRTIDGVLLKCLSADQAKVAIGEVHEGICGTHQSAHKMKWLLRRARYF 2053
Query: 1980 WITMHADCYNHAKRCHKCQIYADKIHIPPSMLNVISSPWPFSMWGIDMIGRIEPKASNGH 2039
W TM DC+ + K C CQ + P S +N I PWPF WGIDMIG I +S GH
Sbjct: 2054 WPTMLEDCFKYYKGCQDCQKFGAIQRAPASAMNPIIKPWPFRGWGIDMIGMISRPSSKGH 2113
Query: 2040 RFILVAIDYFTKWVEAASYANVTKQVVVRFIKNNLISRYGVPNRIITDNGTNLNNNMMKE 2099
+FILVA DYFTK VEA V ++F++ ++I R+G+P I TD G+ ++ +
Sbjct: 2114 KFILVATDYFTKLVEAIPLKKVDSGDAIQFVQEHIIYRFGIPQTITTDQGSIFVSDEFVQ 2173
Query: 2100 LCDDFKIQHHNSSPYRPQMNGAVEAANKNIKKIIQKMVVTY-KDWHEMLPYALYGYRTSV 2158
D I+ NSSPY Q NG EA+NK++ K+I++ Y + WH L AL+ YR +
Sbjct: 2174 FVDSMGIKLLNSSPYYAQANGQAEASNKSLIKLIKRKNSDYPRQWHTRLDEALWSYRMAC 2233
Query: 2159 RTSTGATPFSLVYGMEAVLPVEVEIPSLR 2187
S P+ LVYG EA+LP EV I S R
Sbjct: 2234 HGSIQVPPYKLVYGHEAILPWEVRIGSRR 2262
Score = 69.7 bits (169), Expect = 1e-09
Identities = 53/204 (25%), Positives = 92/204 (44%), Gaps = 11/204 (5%)
Query: 883 NKHNRALLISVLCRTDSLSNVLIDTGSALNVMPKSTLDQLAYSEAPLRLSKVTVRAFDGT 942
N+H + L I+ +S +++D G+ +N+MP +T +L + L + + + F G
Sbjct: 1164 NRHLKPLYINGYVNGKPMSKMMVDGGAEVNLMPYATFRKLGRNVEDLIKTNMVLIDFGGN 1223
Query: 943 RRSVYGEVDLPISVGPHEFQVTFQVMEIQASFSCLLGRPWIHDAGAVTSTLHQKLKFVSR 1002
G ++ ++VG TF V++ S+S LLGR WIH + ST+HQ L
Sbjct: 1224 LSETKGVSNVELTVGNKTIPTTFFVIDGNGSYSLLLGRDWIHANCCIPSTMHQCLIQWQD 1283
Query: 1003 GKLITVSGESAFLISNLSAF--SVIGGSG---------SDGPSFQGFSAEESVGKIETCM 1051
K+ V +S + N S + V+ GS D QGF + + + +I+
Sbjct: 1284 DKIEIVPADSQLKMENPSCYFEGVVEGSNVYIKDTVDDLDDKQGQGFISADDLEEIDIGP 1343
Query: 1052 ASLKDARRVIQEGKTEGWGQLVEL 1075
+ + TE +L+EL
Sbjct: 1344 GDRPRPTFICKNLSTEFRAKLIEL 1367
Score = 48.9 bits (115), Expect = 0.002
Identities = 41/157 (26%), Positives = 71/157 (45%), Gaps = 9/157 (5%)
Query: 218 VNIPPKFKMPVFEKY--QGDTCPQNHLTMYIRRMIAYKNNVPLLIHCFQDSLTGPAHTWF 275
V +P +FK+P F K+ Q T H++ ++ + L + F+ SL G TWF
Sbjct: 654 VPLPNRFKVPDFSKFSWQDSTSTYEHISRFLAQCGEASAVDALKVRLFRLSLAGSTFTWF 713
Query: 276 MGLK--GVTTFEQLAEAFMQQYKYNTYLAPSRKELQSLTQKDKESFKEYAQRFIQKAAQI 333
L + ++ L + F Y Y+ +L S+ QK E EY QRF + +
Sbjct: 714 SSLPYGSINSWADLEKQF-HSYFYSGIHEMKLSDLTSIKQKHDEPVHEYIQRFREMRNKC 772
Query: 334 RP-PLDERELSELFYETLSPCYSEKMIVCASQKFTDL 369
L + +L++L ++ + EK+ +S+ F L
Sbjct: 773 YSLSLTDAQLADLAFQGMIAPIREKL---SSEDFESL 806
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,184,554,853
Number of Sequences: 2540612
Number of extensions: 195554001
Number of successful extensions: 822571
Number of sequences better than 10.0: 38914
Number of HSP's better than 10.0 without gapping: 5253
Number of HSP's successfully gapped in prelim test: 33994
Number of HSP's that attempted gapping in prelim test: 683795
Number of HSP's gapped (non-prelim): 76853
length of query: 2305
length of database: 863,360,394
effective HSP length: 144
effective length of query: 2161
effective length of database: 497,512,266
effective search space: 1075124006826
effective search space used: 1075124006826
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 84 (37.0 bits)
Medicago: description of AC146664.7


