

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146651.7 - phase: 0 /pseudo
(394 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
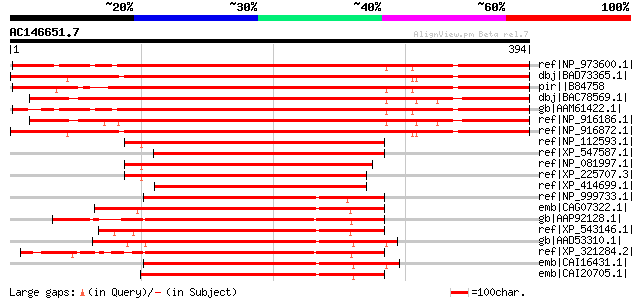
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_973600.1| katanin, putative [Arabidopsis thaliana] 451 e-125
dbj|BAD73365.1| vacuolar protein sorting factor 4B-like [Oryza s... 442 e-123
pir||B84758 probable katanin [imported] - Arabidopsis thaliana 438 e-121
dbj|BAC78569.1| katanin [Oryza sativa (japonica cultivar-group)]... 433 e-120
gb|AAM61422.1| putative katanin [Arabidopsis thaliana] gi|201970... 431 e-119
ref|NP_916186.1| katanin p60 subunit A 1-like [Oryza sativa (jap... 421 e-116
ref|NP_916872.1| putative katanin [Oryza sativa (japonica cultiv... 389 e-107
ref|NP_112593.1| hypothetical protein LOC83473 [Homo sapiens] gi... 286 8e-76
ref|XP_547587.1| PREDICTED: similar to RIKEN cDNA 3110023G01 [Ca... 274 3e-72
ref|NP_081997.1| hypothetical protein LOC71206 [Mus musculus] gi... 267 5e-70
ref|XP_225707.3| PREDICTED: similar to RIKEN cDNA 3110023G01 [Ra... 263 9e-69
ref|XP_414699.1| PREDICTED: similar to RIKEN cDNA 4933439B08 [Ga... 259 1e-67
ref|NP_999733.1| katanin p60 [Strongylocentrotus purpuratus] gi|... 242 2e-62
emb|CAG07322.1| unnamed protein product [Tetraodon nigroviridis] 240 5e-62
gb|AAP92128.1| putative ATPase ATP1 [Oryza sativa (japonica cult... 240 6e-62
ref|XP_543146.1| PREDICTED: similar to katanin p60 subunit A-lik... 239 1e-61
gb|AAD53310.1| katanin p60 [Xenopus laevis] gi|60390218|sp|Q9PUL... 239 1e-61
ref|XP_321284.2| ENSANGP00000018492 [Anopheles gambiae str. PEST... 239 1e-61
emb|CAI16431.1| katanin p60 (ATPase-containing) subunit A 1 [Hom... 239 1e-61
emb|CAI20705.1| novel protein similar to vertebrate katanin p60 ... 239 1e-61
>ref|NP_973600.1| katanin, putative [Arabidopsis thaliana]
Length = 393
Score = 451 bits (1160), Expect = e-125
Identities = 253/403 (62%), Positives = 290/403 (71%), Gaps = 23/403 (5%)
Query: 3 ADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDRAI 62
A DEP TRWSF EFK +YD + GRKKL E VSN + S+ ++G V ++ +
Sbjct: 2 ATDEPSQTRWSFLEFKTFYDAKFGRKKLPEED---VSNKDQPEDGSSNGNNGDVNNNSS- 57
Query: 63 YDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWE 122
+Q N NG N + EKPKKS+ PPFESAE RTLAESLSRDIIRG+PN+KWE
Sbjct: 58 --PVTNQDGNTALANG---NVIREKPKKSMFPPFESAETRTLAESLSRDIIRGNPNIKWE 112
Query: 123 SIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTT 182
SIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKAVATECNTT
Sbjct: 113 SIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKAVATECNTT 172
Query: 183 FFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR-GEGRSEHEASR 241
FFNISASS+VSKWRGDSEKL++VLF+LARHHAP+TIFLDEIDAIISQR GEGRSEHEASR
Sbjct: 173 FFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEGRSEHEASR 232
Query: 242 RLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR--NQKQEEQCLRNSFLC 299
RLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKR + + R F
Sbjct: 233 RLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPEARRGMFEM 292
Query: 300 SLMRN-------RCPMIYWWTGQKDTQARIFAYFVKKQQCSH*DA**HNLNKNQMWCPK- 351
+ ++ G + RI Q A L + P+
Sbjct: 293 LIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLA---ILEDREDVVPED 349
Query: 352 KLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
+LPK+GP++PED++ AL NTRPSAHL AH YD FN DYGSQIL
Sbjct: 350 ELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQIL 392
>dbj|BAD73365.1| vacuolar protein sorting factor 4B-like [Oryza sativa (japonica
cultivar-group)] gi|56201862|dbj|BAD73312.1| vacuolar
protein sorting factor 4B-like [Oryza sativa (japonica
cultivar-group)]
Length = 410
Score = 442 bits (1138), Expect = e-123
Identities = 252/415 (60%), Positives = 290/415 (69%), Gaps = 27/415 (6%)
Query: 1 MAADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGN-----------SSGIASN 49
M DEP TRW+FE+F+ YY+VRLG ++ E+ +G S+G
Sbjct: 1 MTGADEPSITRWTFEDFEVYYEVRLGIRREPGGDEDGDGDGGGGRGYAPLGSGSAGSTRP 60
Query: 50 GNSHGKVTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLS 109
+H +D A+++QF+ + NG G P+KSLLP FESAEMR LAE+L
Sbjct: 61 SAAHANGGADLAVFEQFERLERKVELRNGAIEAG---PPQKSLLPSFESAEMRNLAETLL 117
Query: 110 RDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT 169
RDIIRGSP+VKWESIKGLENAKRLLKEAVVMPIKYPKYF GLLSPWKGILLFGPPGTGKT
Sbjct: 118 RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFKGLLSPWKGILLFGPPGTGKT 177
Query: 170 MLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQ 229
MLAKAVATEC TTFFNISASSIVSKWRGDSEKLVKVLFELARHHAP+TIFLDEIDAIISQ
Sbjct: 178 MLAKAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQ 237
Query: 230 RGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR-NQKQ 288
RGE RSEHEASRRLKTELLIQMDGL +TD+LVFVLAATNLPWELDAAMLRRLEKR
Sbjct: 238 RGEARSEHEASRRLKTELLIQMDGLTKTDDLVFVLAATNLPWELDAAMLRRLEKRILVPL 297
Query: 289 EEQCLRNSFLCSLMRN-----RCP---MIYWWTGQKDTQARIFAYFVKKQQCSH*DA**H 340
EQ R++ L+ + P ++ G + R+ Q +
Sbjct: 298 PEQEARHAMFEELLPSVPGTMNIPYDVLVEKTEGYSGSDIRLVCKEAAMQPLRRLMS--- 354
Query: 341 NLNKNQMWCPK-KLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
L Q P+ +LP+VGPV ED+E ALRNTRPSAHL H+Y+ FN DYGS +L
Sbjct: 355 VLEGRQEEVPEDELPEVGPVTTEDIELALRNTRPSAHLHVHRYEKFNQDYGSHVL 409
>pir||B84758 probable katanin [imported] - Arabidopsis thaliana
Length = 393
Score = 438 bits (1127), Expect = e-121
Identities = 249/410 (60%), Positives = 285/410 (68%), Gaps = 37/410 (9%)
Query: 3 ADDEPMPTRWSFEEFKKYYDVRLGRKKLVENG-------ENAVSNGNSSGIASNGNSHGK 55
A DEP TRWSF EFK +YD + GRKKL E E+ SNGN+ + +N +
Sbjct: 2 ATDEPSQTRWSFLEFKTFYDAKFGRKKLPEEDVSNKDQPEDGSSNGNNGDVNNNSSP--- 58
Query: 56 VTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRG 115
VT+ R + G + KKS+ PPFESAE RTLAESLSRDIIRG
Sbjct: 59 VTNQRC-------------YVYSLGSDECFFCRKKSMFPPFESAETRTLAESLSRDIIRG 105
Query: 116 SPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAV 175
+PN+KWESIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKAV
Sbjct: 106 NPNIKWESIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKAV 165
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR-GEGR 234
ATECNTTFFNISASS+VSKWRGDSEKL++VLF+LARHHAP+TIFLDEIDAIISQR GEGR
Sbjct: 166 ATECNTTFFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEGR 225
Query: 235 SEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR--NQKQEEQC 292
SEHEASRRLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKR + +
Sbjct: 226 SEHEASRRLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPEA 285
Query: 293 LRNSFLCSLMRN-------RCPMIYWWTGQKDTQARIFAYFVKKQQCSH*DA**HNLNKN 345
R F + ++ G + RI Q A L
Sbjct: 286 RRGMFEMLIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLA---ILEDR 342
Query: 346 QMWCPK-KLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
+ P+ +LPK+GP++PED++ AL NTRPSAHL AH YD FN DYGSQIL
Sbjct: 343 EDVVPEDELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQIL 392
>dbj|BAC78569.1| katanin [Oryza sativa (japonica cultivar-group)]
gi|57899262|dbj|BAD87507.1| katanin [Oryza sativa
(japonica cultivar-group)]
Length = 386
Score = 433 bits (1113), Expect = e-120
Identities = 244/392 (62%), Positives = 287/392 (72%), Gaps = 25/392 (6%)
Query: 16 EFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDRAIYDQFQSQGQNPTH 75
+FK +++ R G KK E +N +NG ++G S K TSD A+Y+QF+ Q +
Sbjct: 2 DFKGFWESRFGGKKEQEPEQNGHANGVANG------SVRKRTSDLAVYEQFEQQARQTEV 55
Query: 76 TNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLK 135
+G + +K LLP FESAEMR LAE+L RDIIRGSP+VKWESIKGLENAKRLLK
Sbjct: 56 RAAAIRDGNADAIQKPLLPSFESAEMRNLAETLLRDIIRGSPDVKWESIKGLENAKRLLK 115
Query: 136 EAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKW 195
EAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATEC TTFFNISASSIVSKW
Sbjct: 116 EAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSIVSKW 175
Query: 196 RGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLA 255
RGDSEKLVKVLFELARHHAP+TIFLDEIDAIISQRGE RSEHEASRRLKTELLIQMDGL
Sbjct: 176 RGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLT 235
Query: 256 RTDELVFVLAATNLPWELDAAMLRRLEKR-----NQKQEEQCLRNSFLCS-LMRNRCP-- 307
+T++LVFVLAATNLPWELDAAMLRRLEKR + + + L S + P
Sbjct: 236 KTNDLVFVLAATNLPWELDAAMLRRLEKRILVPLPEAEARHAMFEELLPSTTSKLEVPYD 295
Query: 308 -MIYWWTGQKDTQARIF----AYFVKKQQCSH*DA**HNLNKNQMWCPKKLPKVGPVVPE 362
++ G + R+ A ++ S +A ++++ ++LP+VGP+ PE
Sbjct: 296 TLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEA------RDELVPEEELPEVGPLKPE 349
Query: 363 DVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
D+E ALRNTRPSAHL AH+Y+ FN DYGSQIL
Sbjct: 350 DIEVALRNTRPSAHLHAHRYEKFNQDYGSQIL 381
>gb|AAM61422.1| putative katanin [Arabidopsis thaliana] gi|20197082|gb|AAC26698.2|
putative katanin [Arabidopsis thaliana]
gi|18403587|ref|NP_565791.1| katanin, putative
[Arabidopsis thaliana]
Length = 384
Score = 431 bits (1108), Expect = e-119
Identities = 248/403 (61%), Positives = 283/403 (69%), Gaps = 32/403 (7%)
Query: 3 ADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDRAI 62
A DEP TRWSF GRKKL E VSN + S+ ++G V ++ +
Sbjct: 2 ATDEPSQTRWSF---------LFGRKKLPEED---VSNKDQPEDGSSNGNNGDVNNNSS- 48
Query: 63 YDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWE 122
+Q N NG N + EKPKKS+ PPFESAE RTLAESLSRDIIRG+PN+KWE
Sbjct: 49 --PVTNQDGNTALANG---NVIREKPKKSMFPPFESAETRTLAESLSRDIIRGNPNIKWE 103
Query: 123 SIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTT 182
SIKGLENAK+LLKEAVVMPIKYP YF GLL+PWKGILLFGPPGTGKTMLAKAVATECNTT
Sbjct: 104 SIKGLENAKKLLKEAVVMPIKYPTYFNGLLTPWKGILLFGPPGTGKTMLAKAVATECNTT 163
Query: 183 FFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR-GEGRSEHEASR 241
FFNISASS+VSKWRGDSEKL++VLF+LARHHAP+TIFLDEIDAIISQR GEGRSEHEASR
Sbjct: 164 FFNISASSVVSKWRGDSEKLIRVLFDLARHHAPSTIFLDEIDAIISQRGGEGRSEHEASR 223
Query: 242 RLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR--NQKQEEQCLRNSFLC 299
RLKTELLIQMDGL +T+ELVFVLAATNLPWELDAAMLRRLEKR + + R F
Sbjct: 224 RLKTELLIQMDGLQKTNELVFVLAATNLPWELDAAMLRRLEKRILVPLPDPEARRGMFEM 283
Query: 300 SLMRN-------RCPMIYWWTGQKDTQARIFAYFVKKQQCSH*DA**HNLNKNQMWCPK- 351
+ ++ G + RI Q A L + P+
Sbjct: 284 LIPSQPGDEPLPHDVLVEKSEGYSGSDIRILCKEAAMQPLRRTLA---ILEDREDVVPED 340
Query: 352 KLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
+LPK+GP++PED++ AL NTRPSAHL AH YD FN DYGSQIL
Sbjct: 341 ELPKIGPILPEDIDRALSNTRPSAHLHAHLYDKFNDDYGSQIL 383
>ref|NP_916186.1| katanin p60 subunit A 1-like [Oryza sativa (japonica
cultivar-group)]
Length = 428
Score = 421 bits (1081), Expect = e-116
Identities = 246/414 (59%), Positives = 289/414 (69%), Gaps = 47/414 (11%)
Query: 16 EFKKYYDVRLGRKKLVENGENAVSNGNSSGIASNGNSHGKVTSDRAIYDQFQSQG----- 70
+FK +++ R G KK E +N +NG ++G S K TSD A+Y+QF+ Q
Sbjct: 22 DFKGFWESRFGGKKEQEPEQNGHANGVANG------SVRKRTSDLAVYEQFEQQVFLDRR 75
Query: 71 ------QNPTHTNGFGP-----------NGVDEKPKKSLLPPFESAEMRTLAESLSRDII 113
+P F +G + +K LLP FESAEMR LAE+L RDII
Sbjct: 76 LVSCVVLDPRSVLLFQARQTEVRAAAIRDGNADAIQKPLLPSFESAEMRNLAETLLRDII 135
Query: 114 RGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAK 173
RGSP+VKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAK
Sbjct: 136 RGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAK 195
Query: 174 AVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEG 233
AVATEC TTFFNISASSIVSKWRGDSEKLVKVLFELARHHAP+TIFLDEIDAIISQRGE
Sbjct: 196 AVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGEA 255
Query: 234 RSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR-----NQKQ 288
RSEHEASRRLKTELLIQMDGL +T++LVFVLAATNLPWELDAAMLRRLEKR + +
Sbjct: 256 RSEHEASRRLKTELLIQMDGLTKTNDLVFVLAATNLPWELDAAMLRRLEKRILVPLPEAE 315
Query: 289 EEQCLRNSFLCS-LMRNRCP---MIYWWTGQKDTQARIF----AYFVKKQQCSH*DA**H 340
+ L S + P ++ G + R+ A ++ S +A
Sbjct: 316 ARHAMFEELLPSTTSKLEVPYDTLVEKTEGYSGSDIRLVCKEAAMQPLRRLMSVLEA--- 372
Query: 341 NLNKNQMWCPKKLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
++++ ++LP+VGP+ PED+E ALRNTRPSAHL AH+Y+ FN DYGSQIL
Sbjct: 373 ---RDELVPEEELPEVGPLKPEDIEVALRNTRPSAHLHAHRYEKFNQDYGSQIL 423
>ref|NP_916872.1| putative katanin [Oryza sativa (japonica cultivar-group)]
Length = 411
Score = 389 bits (999), Expect = e-107
Identities = 232/416 (55%), Positives = 275/416 (65%), Gaps = 28/416 (6%)
Query: 1 MAADDEPMPTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGN-----------SSGIASN 49
M DEP TRW+FE+F+ YY+VRLG ++ E+ +G S+G
Sbjct: 1 MTGADEPSITRWTFEDFEVYYEVRLGIRREPGGDEDGDGDGGGGRGYAPLGSGSAGSTRP 60
Query: 50 GNSHGKVTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLS 109
+H +D A+++QF+ + NG G P+KSLLP FESAEMR LAE+L
Sbjct: 61 SAAHANGGADLAVFEQFERLERKVELRNGAIEAG---PPQKSLLPSFESAEMRNLAETLL 117
Query: 110 RDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFT-GLLSPWKGILLFGPPGTGK 168
RDIIRGSP+VKWESIKGLENAKRLLKEAVVMPIKYPK ++ + I + +
Sbjct: 118 RDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKVLQRSAITLERYIAFWSTRNRKE 177
Query: 169 TMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIIS 228
TMLAKAVATEC TTFFNISASSIVSKWRGDSEKLVKVLFELARHHAP+TIFLDEIDAIIS
Sbjct: 178 TMLAKAVATECKTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIIS 237
Query: 229 QRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR-NQK 287
QRGE RSEHEASRRLKTELLIQMDGL +TD+LVFVLAATNLPWELDAAMLRRLEKR
Sbjct: 238 QRGEARSEHEASRRLKTELLIQMDGLTKTDDLVFVLAATNLPWELDAAMLRRLEKRILVP 297
Query: 288 QEEQCLRNSFLCSLMRN-----RCP---MIYWWTGQKDTQARIFAYFVKKQQCSH*DA** 339
EQ R++ L+ + P ++ G + R+ Q +
Sbjct: 298 LPEQEARHAMFEELLPSVPGTMNIPYDVLVEKTEGYSGSDIRLVCKEAAMQPLRRLMS-- 355
Query: 340 HNLNKNQMWCPK-KLPKVGPVVPEDVEAALRNTRPSAHLLAHKYDTFNADYGSQIL 394
L Q P+ +LP+VGPV ED+E ALRNTRPSAHL H+Y+ FN DYGS +L
Sbjct: 356 -VLEGRQEEVPEDELPEVGPVTTEDIELALRNTRPSAHLHVHRYEKFNQDYGSHVL 410
>ref|NP_112593.1| hypothetical protein LOC83473 [Homo sapiens]
gi|34190544|gb|AAH34999.2| Similar to mouse
4933439B08Rik protein [Homo sapiens]
Length = 466
Score = 286 bits (732), Expect = 8e-76
Identities = 142/203 (69%), Positives = 172/203 (83%), Gaps = 6/203 (2%)
Query: 88 PKKSLLPPFES-----AEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPI 142
P + LL P + +EMR LA +SRDI +PN+KW I GL+ AK+L+KEAVV PI
Sbjct: 143 PSERLLKPLSAFIGMNSEMRELAAVVSRDIYLHNPNIKWNDIIGLDAAKQLVKEAVVYPI 202
Query: 143 KYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKL 202
+YP+ FTG+LSPWKG+LL+GPPGTGKT+LAKAVATEC TTFFNISAS+IVSKWRGDSEKL
Sbjct: 203 RYPQLFTGILSPWKGLLLYGPPGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKL 262
Query: 203 VKVLFELARHHAPATIFLDEIDAIISQRGEGR-SEHEASRRLKTELLIQMDGLARTDELV 261
V+VLFELAR+HAP+TIFLDE+++++SQRG EHE S R+KTELL+QMDGLAR+++LV
Sbjct: 263 VRVLFELARYHAPSTIFLDELESVMSQRGTASGGEHEGSLRMKTELLVQMDGLARSEDLV 322
Query: 262 FVLAATNLPWELDAAMLRRLEKR 284
FVLAA+NLPWELD AMLRRLEKR
Sbjct: 323 FVLAASNLPWELDCAMLRRLEKR 345
>ref|XP_547587.1| PREDICTED: similar to RIKEN cDNA 3110023G01 [Canis familiaris]
Length = 567
Score = 274 bits (701), Expect = 3e-72
Identities = 131/176 (74%), Positives = 159/176 (89%), Gaps = 1/176 (0%)
Query: 110 RDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKT 169
+DI +PN+KW+ I GL+ AK+L+KEAVV PI+YP+ FTG+LSPWKG+LL+GPPGTGKT
Sbjct: 281 KDIYLHNPNIKWDDIIGLDTAKQLVKEAVVYPIRYPQLFTGILSPWKGLLLYGPPGTGKT 340
Query: 170 MLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQ 229
+LAKAVATEC TTFFNISAS+IVSKWRGDSEKLV+VLFELAR+HAP+TIFLDE+++++SQ
Sbjct: 341 LLAKAVATECKTTFFNISASTIVSKWRGDSEKLVRVLFELARYHAPSTIFLDELESVMSQ 400
Query: 230 RGEG-RSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR 284
RG EHE S R+KTELL+QMDGLAR+++LVFVLAA+NLPWELD AMLRRLEKR
Sbjct: 401 RGTAPGGEHEGSLRMKTELLVQMDGLARSEDLVFVLAASNLPWELDCAMLRRLEKR 456
>ref|NP_081997.1| hypothetical protein LOC71206 [Mus musculus]
gi|12856210|dbj|BAB30604.1| unnamed protein product [Mus
musculus]
Length = 409
Score = 267 bits (682), Expect = 5e-70
Identities = 133/194 (68%), Positives = 163/194 (83%), Gaps = 6/194 (3%)
Query: 88 PKKSLLPPFES-----AEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPI 142
P + LL P + +EMR LA +SRDI +PN+KW I GL+ AK+L+KEAVV PI
Sbjct: 216 PSERLLKPLSAFIGMNSEMRELAAVVSRDIYLHNPNIKWNDIIGLDAAKQLVKEAVVYPI 275
Query: 143 KYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKL 202
+YP+ FTG+LSPWKG+LL+GPPGTGKT+LAKAVATEC TTFFNISAS+IVSKWRGDSEKL
Sbjct: 276 RYPQLFTGILSPWKGLLLYGPPGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKL 335
Query: 203 VKVLFELARHHAPATIFLDEIDAIISQRG-EGRSEHEASRRLKTELLIQMDGLARTDELV 261
V+VLFELAR+HAP+TIFLDE+++++SQRG EHE S R+KTELL+QMDGLAR+++LV
Sbjct: 336 VRVLFELARYHAPSTIFLDELESVMSQRGMVPGGEHEGSLRMKTELLVQMDGLARSEDLV 395
Query: 262 FVLAATNLPWELDA 275
FVLAA+NLPWE DA
Sbjct: 396 FVLAASNLPWEGDA 409
>ref|XP_225707.3| PREDICTED: similar to RIKEN cDNA 3110023G01 [Rattus norvegicus]
Length = 526
Score = 263 bits (671), Expect = 9e-69
Identities = 130/190 (68%), Positives = 160/190 (83%), Gaps = 6/190 (3%)
Query: 88 PKKSLLPPFES-----AEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPI 142
P + LL P + +EMR LA +SRDI +PN+KW I GL+ AK+L+KEAVV PI
Sbjct: 337 PSERLLKPLSAFIGMNSEMRELAAVVSRDIYLHNPNIKWNDIIGLDAAKQLVKEAVVYPI 396
Query: 143 KYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKL 202
+YP+ FTG+LSPWKG+LL+GPPGTGKT+LAKAVATEC TTFFNISAS+IVSKWRGDSEKL
Sbjct: 397 RYPQLFTGILSPWKGLLLYGPPGTGKTLLAKAVATECKTTFFNISASTIVSKWRGDSEKL 456
Query: 203 VKVLFELARHHAPATIFLDEIDAIISQRG-EGRSEHEASRRLKTELLIQMDGLARTDELV 261
V+VLFELAR+HAP+TIFLDE+++++SQRG EHE S R+KTELL+QMDGLAR+++LV
Sbjct: 457 VRVLFELARYHAPSTIFLDELESVMSQRGMVPGGEHEGSLRMKTELLVQMDGLARSEDLV 516
Query: 262 FVLAATNLPW 271
FVLAA+NLPW
Sbjct: 517 FVLAASNLPW 526
>ref|XP_414699.1| PREDICTED: similar to RIKEN cDNA 4933439B08 [Gallus gallus]
Length = 448
Score = 259 bits (662), Expect = 1e-67
Identities = 124/162 (76%), Positives = 148/162 (90%), Gaps = 1/162 (0%)
Query: 111 DIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTM 170
DI +PNVKW+ I GL+ AKRL+KEAVV PI+YP+ FTG+LSPWKG+LL+GPPGTGKT+
Sbjct: 287 DIYLHNPNVKWDDIIGLDAAKRLVKEAVVYPIRYPQLFTGILSPWKGLLLYGPPGTGKTL 346
Query: 171 LAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQR 230
LAKAVATECNTTFFNISAS+IVSKWRGDSEKLV+VLFELAR+HAP+TIFLDE+++++SQR
Sbjct: 347 LAKAVATECNTTFFNISASTIVSKWRGDSEKLVRVLFELARYHAPSTIFLDELESVMSQR 406
Query: 231 GE-GRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPW 271
G EHE SRR+KTELL+QMDGLAR+D+LVFVLAA+NLPW
Sbjct: 407 GTISGGEHEGSRRMKTELLVQMDGLARSDDLVFVLAASNLPW 448
>ref|NP_999733.1| katanin p60 [Strongylocentrotus purpuratus]
gi|3098603|gb|AAC15706.1| katanin p60 subunit
[Strongylocentrotus purpuratus]
gi|60390159|sp|O61577|KTNA1_STRPU Katanin p60
ATPase-containing subunit (Katanin p60 subunit) (p60
katanin)
Length = 516
Score = 242 bits (617), Expect = 2e-62
Identities = 118/189 (62%), Positives = 153/189 (80%), Gaps = 7/189 (3%)
Query: 102 RTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLF 161
+ L E+L RDI++ +PNV W I GL AKRLL+EAVV+P+ P YF G+ PWKG+L+
Sbjct: 214 KDLVENLERDIVQRNPNVHWADIAGLTEAKRLLEEAVVLPLWMPDYFKGIRRPWKGVLMV 273
Query: 162 GPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLD 221
GPPGTGKTMLAKAVATEC TTFFN+S++S+ SK+ G+SEKLV++LFE+AR +AP+TIF+D
Sbjct: 274 GPPGTGKTMLAKAVATECGTTFFNVSSASLTSKYHGESEKLVRLLFEMARFYAPSTIFID 333
Query: 222 EIDAIISQRGEGRSEHEASRRLKTELLIQMDGLA------RTDELVFVLAATNLPWELDA 275
EID+I S+RG G SEHEASRR+K+ELLIQMDG++ + ++V VLAATN PW++D
Sbjct: 334 EIDSICSKRGTG-SEHEASRRVKSELLIQMDGVSGPSAGEESSKMVMVLAATNFPWDIDE 392
Query: 276 AMLRRLEKR 284
A+ RRLEKR
Sbjct: 393 ALRRRLEKR 401
>emb|CAG07322.1| unnamed protein product [Tetraodon nigroviridis]
Length = 486
Score = 240 bits (613), Expect = 5e-62
Identities = 125/233 (53%), Positives = 165/233 (70%), Gaps = 14/233 (6%)
Query: 65 QFQSQGQNPTHTNGFGPNGVDEKPKKSLLPP-------FESAEMRT-LAESLSRDIIRGS 116
Q ++ + P + G DEK KK P F+ A + L + L RDI+ +
Sbjct: 140 QTNAKAERPGLKDARGVRAKDEKGKKGASEPGDGELKKFDGAGHDSDLVDLLERDIVSRN 199
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVA 176
PNV W+ I LE+AK+LL+EAVV+P+ P +F G+ PWKG+L+ GPPGTGKTMLAKAVA
Sbjct: 200 PNVHWDDIADLEDAKKLLREAVVLPMWMPDFFKGIRRPWKGVLMVGPPGTGKTMLAKAVA 259
Query: 177 TECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSE 236
TEC TTFFN+S+S++ SK+RG+SEKLV+VLFE+AR +AP TIF+DEID+I S+RG E
Sbjct: 260 TECGTTFFNVSSSTLTSKYRGESEKLVRVLFEMARFYAPTTIFIDEIDSICSRRGTS-DE 318
Query: 237 HEASRRLKTELLIQMDGLART-----DELVFVLAATNLPWELDAAMLRRLEKR 284
HEASRR+K+E L+QMDG+ T ++V VLAATN PW++D A+ RRLEKR
Sbjct: 319 HEASRRVKSEFLVQMDGMGNTPDEDPSKMVMVLAATNFPWDIDEALRRRLEKR 371
>gb|AAP92128.1| putative ATPase ATP1 [Oryza sativa (japonica cultivar-group)]
gi|34911210|ref|NP_916952.1| putative CAD ATPase [Oryza
sativa (japonica cultivar-group)]
gi|19386661|dbj|BAB86043.1| putative katanin [Oryza
sativa (japonica cultivar-group)]
gi|21644706|dbj|BAC01262.1| putative katanin [Oryza
sativa (japonica cultivar-group)]
Length = 519
Score = 240 bits (612), Expect = 6e-62
Identities = 125/259 (48%), Positives = 173/259 (66%), Gaps = 27/259 (10%)
Query: 33 NGENAVSNGNSSGIASNGNSHGKVTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKPKKSL 92
NG S SS +S G GK +S +A D S D + KS
Sbjct: 165 NGSKGNSGARSSTASSTGGRKGKSSSSKA--DSVSS----------------DAEEGKSK 206
Query: 93 LPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLL 152
+E +M LA L RD++ +P V+W+ + GL AKRLL+EAVV+P+ P+YF G+
Sbjct: 207 KGQYEGPDM-DLAAMLERDVLDSTPGVRWDDVAGLSEAKRLLEEAVVLPLWMPEYFQGIR 265
Query: 153 SPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARH 212
PWKG+L+FGPPGTGKT+LAKAVATEC TTFFN+S++++ SKWRG+SE++V+ LF+LAR
Sbjct: 266 RPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARF 325
Query: 213 HAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTD-------ELVFVLA 265
+AP+TIF+DEID++ + RG EHE+SRR+K+ELL+Q+DG+ + ++V VLA
Sbjct: 326 YAPSTIFIDEIDSLCTSRG-ASGEHESSRRVKSELLVQIDGVNNSSTTEDGQPKIVMVLA 384
Query: 266 ATNLPWELDAAMLRRLEKR 284
ATN PW++D A+ RRLEKR
Sbjct: 385 ATNFPWDIDEALRRRLEKR 403
>ref|XP_543146.1| PREDICTED: similar to katanin p60 subunit A-like 1 [Canis
familiaris]
Length = 582
Score = 239 bits (610), Expect = 1e-61
Identities = 123/234 (52%), Positives = 170/234 (72%), Gaps = 18/234 (7%)
Query: 68 SQGQNPTHTNG--FGPNGVDEKPKKSL--------LPPFESAEM-RTLAESLSRDIIRGS 116
S+G+ P+ + + G D+K +K++ +P F+ A + L E+L RDI+ +
Sbjct: 235 SKGEKPSTSRDKDYRARGRDDKGRKNMHDGASDGEIPKFDGAGYDKDLVEALERDIVSRN 294
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVA 176
P++ W+ I LE AK+LL+EAVV+P+ P +F G+ PWKG+L+ GPPGTGKTMLAKAVA
Sbjct: 295 PSIHWDDIADLEEAKKLLREAVVLPMWMPDFFKGIRRPWKGVLMVGPPGTGKTMLAKAVA 354
Query: 177 TECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSE 236
TEC TTFFN+S+S++ SK+RG+SEKLV++LFE+AR +AP TIF+DEID+I S+RG E
Sbjct: 355 TECGTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAPTTIFIDEIDSICSRRGTS-DE 413
Query: 237 HEASRRLKTELLIQMDGLART------DELVFVLAATNLPWELDAAMLRRLEKR 284
HEASRR+K+ELLIQMDG+ ++V VLAATN PW++D A+ RRLEKR
Sbjct: 414 HEASRRVKSELLIQMDGVGGALENDDPSKMVMVLAATNFPWDIDEALRRRLEKR 467
>gb|AAD53310.1| katanin p60 [Xenopus laevis] gi|60390218|sp|Q9PUL2|KTNA1_XENLA
Katanin p60 ATPase-containing subunit (Katanin p60
subunit) (p60 katanin)
Length = 486
Score = 239 bits (610), Expect = 1e-61
Identities = 130/263 (49%), Positives = 175/263 (66%), Gaps = 33/263 (12%)
Query: 64 DQFQSQGQNPTHTNGFGPNGVDEKP------KKSLLPPFESAEM--------------RT 103
++F + + P + N V KP K +L+ SA++ +
Sbjct: 128 NRFSAAAKGPNLPSARNANNVKMKPVRAREKKDALIKNKSSADVSETEVKRFDGSGYDKD 187
Query: 104 LAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGP 163
L E+L RDII +PN++W+ I LE AK+LLKEAVV+P+ P++F G+ PWKG+L+ GP
Sbjct: 188 LIEALERDIISQNPNIRWDDIADLEEAKKLLKEAVVLPMWMPEFFKGIRRPWKGVLMVGP 247
Query: 164 PGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEI 223
PGTGKT+LAKAVATEC TTFFNIS+S++ SK+RG+SEKLV++LFE+AR +AP TIF+DEI
Sbjct: 248 PGTGKTLLAKAVATECKTTFFNISSSTLTSKYRGESEKLVRLLFEMARFYAPTTIFIDEI 307
Query: 224 DAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDE------LVFVLAATNLPWELDAAM 277
D+I S+RG EHEASRR+K ELL+QMDG+ E +V VLAATN PW++D A+
Sbjct: 308 DSICSRRGTS-EEHEASRRVKAELLVQMDGVGGASENEDPSKMVMVLAATNFPWDIDEAL 366
Query: 278 LRRLEKR------NQKQEEQCLR 294
RRLEKR + K E+ LR
Sbjct: 367 RRRLEKRIYIPLPSAKGREELLR 389
>ref|XP_321284.2| ENSANGP00000018492 [Anopheles gambiae str. PEST]
gi|55233566|gb|EAA01173.2| ENSANGP00000018492 [Anopheles
gambiae str. PEST]
Length = 492
Score = 239 bits (609), Expect = 1e-61
Identities = 137/283 (48%), Positives = 184/283 (64%), Gaps = 27/283 (9%)
Query: 9 PTRWSFEEFKKYYDVRLGRKKLVENGENAVSNGNSSGI-----ASNGNSHGKVTSDRAIY 63
P RW + R R V G G S+G A+NGN G+ +
Sbjct: 115 PIRWPGRRRR-----RTDRSGRVGPGPRYAPAGPSAGAPGASDATNGN--GEKGDKEKLG 167
Query: 64 DQFQSQGQNPTHTNGFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWES 123
D +G N G P V+ K P A++ L + L RDI++ +PN+ W+
Sbjct: 168 DD--EEGNN----GGDTPEEVERK-----FEPASHADV-DLVDMLERDILQKNPNIHWDD 215
Query: 124 IKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTF 183
I L AKRLL+EAVV+P+ P YF G+ PWKG+L+ GPPGTGKTMLAKAVATEC TTF
Sbjct: 216 IADLHEAKRLLEEAVVLPMWMPDYFKGIRRPWKGVLMVGPPGTGKTMLAKAVATECGTTF 275
Query: 184 FNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRL 243
FN+S+S++ SK+RG+SEKLV++LFE+AR +AP+TIF+DEID++ S+RG SEHEASRR+
Sbjct: 276 FNVSSSTLTSKYRGESEKLVRLLFEMARFYAPSTIFIDEIDSLCSRRG-SESEHEASRRV 334
Query: 244 KTELLIQMDGLARTD--ELVFVLAATNLPWELDAAMLRRLEKR 284
K+ELL+QMDG++ + ++V VLAATN PW++D A+ RRLEKR
Sbjct: 335 KSELLVQMDGVSNDEATKIVMVLAATNFPWDIDEALRRRLEKR 377
>emb|CAI16431.1| katanin p60 (ATPase-containing) subunit A 1 [Homo sapiens]
gi|56417804|emb|CAI19505.1| katanin p60
(ATPase-containing) subunit A 1 [Homo sapiens]
gi|60390161|sp|O75449|KTNA1_HUMAN Katanin p60
ATPase-containing subunit A1 (Katanin p60 subunit A1)
(p60 katanin) gi|3283072|gb|AAC25114.1| p60 katanin
[Homo sapiens] gi|5901990|ref|NP_008975.1| katanin p60
subunit A 1 [Homo sapiens]
Length = 491
Score = 239 bits (609), Expect = 1e-61
Identities = 121/207 (58%), Positives = 158/207 (75%), Gaps = 13/207 (6%)
Query: 102 RTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLF 161
+ L E+L RDII +PNV+W+ I L AK+LLKEAVV+P+ P++F G+ PWKG+L+
Sbjct: 189 KDLVEALERDIISQNPNVRWDDIADLVEAKKLLKEAVVLPMWMPEFFKGIRRPWKGVLMV 248
Query: 162 GPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLD 221
GPPGTGKT+LAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LFE+AR ++PATIF+D
Sbjct: 249 GPPGTGKTLLAKAVATECKTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYSPATIFID 308
Query: 222 EIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDE------LVFVLAATNLPWELDA 275
EID+I S+RG EHEASRR+K ELL+QMDG+ T E +V VLAATN PW++D
Sbjct: 309 EIDSICSRRGTS-EEHEASRRVKAELLVQMDGVGGTSENDDPSKMVMVLAATNFPWDIDE 367
Query: 276 AMLRRLEKR------NQKQEEQCLRNS 296
A+ RRLEKR + K E+ LR S
Sbjct: 368 ALRRRLEKRIYIPLPSAKGREELLRIS 394
>emb|CAI20705.1| novel protein similar to vertebrate katanin p60 (ATPase-containing)
subunit A 1 (KATNA1) [Danio rerio]
gi|66472538|ref|NP_001018440.1| hypothetical protein
LOC553631 [Danio rerio] gi|63101878|gb|AAH95321.1|
Hypothetical protein LOC553631 [Danio rerio]
Length = 485
Score = 239 bits (609), Expect = 1e-61
Identities = 117/190 (61%), Positives = 151/190 (78%), Gaps = 6/190 (3%)
Query: 100 EMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGIL 159
E + L ++L RDII +PNV W+ I LE AK+LLKEAVV+P+ P++F G+ PWKG+L
Sbjct: 182 EDKDLIDALERDIISQNPNVTWDDIADLEEAKKLLKEAVVLPMWMPEFFKGIRRPWKGVL 241
Query: 160 LFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIF 219
+ GPPGTGKT+LAKAVATEC TTFFN+S+S++ SK+RG+SEKLV++LFE+AR +AP TIF
Sbjct: 242 MVGPPGTGKTLLAKAVATECRTTFFNVSSSTLTSKYRGESEKLVRLLFEMARFYAPTTIF 301
Query: 220 LDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDE-----LVFVLAATNLPWELD 274
+DEID+I S+RG EHEASRR+K ELL+QMDG+ T E +V VLAATN PW++D
Sbjct: 302 IDEIDSICSRRGTS-EEHEASRRVKAELLVQMDGVGGTSENDPSKMVMVLAATNFPWDID 360
Query: 275 AAMLRRLEKR 284
A+ RRLEKR
Sbjct: 361 EALRRRLEKR 370
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 678,639,580
Number of Sequences: 2540612
Number of extensions: 28183824
Number of successful extensions: 107443
Number of sequences better than 10.0: 4295
Number of HSP's better than 10.0 without gapping: 2689
Number of HSP's successfully gapped in prelim test: 1607
Number of HSP's that attempted gapping in prelim test: 99679
Number of HSP's gapped (non-prelim): 5488
length of query: 394
length of database: 863,360,394
effective HSP length: 130
effective length of query: 264
effective length of database: 533,080,834
effective search space: 140733340176
effective search space used: 140733340176
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146651.7


