

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.5 - phase: 0 /pseudo
(637 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
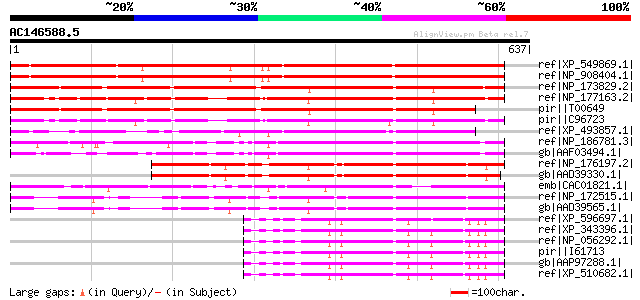
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_549869.1| transcriptional co-repressor -like [Oryza sativ... 574 e-162
ref|NP_908404.1| putative co-repressor protein [Oryza sativa (ja... 574 e-162
ref|NP_173829.2| paired amphipathic helix repeat-containing prot... 523 e-147
ref|NP_177163.2| paired amphipathic helix repeat-containing prot... 490 e-137
pir||T00649 hypothetical protein F3I6.12 - Arabidopsis thaliana ... 486 e-136
pir||C96723 hypothetical protein F20P5.21 [imported] - Arabidops... 483 e-134
ref|XP_493857.1| Similar to Arabidopsis chromosome I BAC genomic... 422 e-116
ref|NP_186781.3| paired amphipathic helix repeat-containing prot... 399 e-109
gb|AAF03494.1| unknown protein [Arabidopsis thaliana] 379 e-103
ref|NP_176197.2| paired amphipathic helix repeat-containing prot... 375 e-102
gb|AAD39330.1| Hypothetical protein [Arabidopsis thaliana] gi|25... 370 e-101
emb|CAC01821.1| transcriptional regulatory-like protein [Arabido... 360 8e-98
ref|NP_172515.1| paired amphipathic helix repeat-containing prot... 323 1e-86
gb|AAD39565.1| T10O24.5 [Arabidopsis thaliana] gi|25402600|pir||... 322 3e-86
ref|XP_596697.1| PREDICTED: similar to mSin3A, partial [Bos taurus] 102 3e-20
ref|XP_343396.1| PREDICTED: similar to mSin3A [Rattus norvegicus] 101 6e-20
ref|NP_056292.1| transcriptional co-repressor Sin3A [Homo sapien... 101 8e-20
pir||I61713 co-repressor protein - mouse gi|642619|gb|AAA69773.1... 101 8e-20
gb|AAP97288.1| MSIN3A [Homo sapiens] 101 8e-20
ref|XP_510682.1| PREDICTED: similar to transcriptional co-repres... 101 8e-20
>ref|XP_549869.1| transcriptional co-repressor -like [Oryza sativa (japonica
cultivar-group)] gi|52076211|dbj|BAD44865.1|
transcriptional co-repressor -like [Oryza sativa
(japonica cultivar-group)]
Length = 1243
Score = 574 bits (1479), Expect = e-162
Identities = 333/635 (52%), Positives = 426/635 (66%), Gaps = 33/635 (5%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAK-GDSSPGVGATVMSPK 59
++IWTTFLEP+LGV R ED DAVK K + K+G A++ + ++ G A +
Sbjct: 565 VRIWTTFLEPILGVQPRTHGAEDA-DAVKPKSRTTKSGLATVGEINTTAAGAVAKHGHDE 623
Query: 60 NT-FEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSR 118
N EQ+ S NGV ++ DR+ + E ++ + G V + +E+
Sbjct: 624 NIPQEQTPSSLARMVNGVATDTQNGFHDVDRTARRAEEPSNTAVNGRVQGASPGTNEIPA 683
Query: 119 VNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSR-----PGYVYREGGLDL 173
V+ Q+ E+ N+ ++ + N + N+ SG+ A+ S R G L
Sbjct: 684 VSTQNMPTERSAE-NIPVARTEQHGNAKANLEPTSGVNASRSSHAGNDTAAEARAGNETL 742
Query: 174 PSSEGADSTRPDTSTNGA-IIEDTKAHRCHKESVGHF--KSEREEGELSPNGDFEEDNFA 230
PS EG ++ R ++ NG E K ++ S H K EREEGELSPNGDFEEDNFA
Sbjct: 743 PSVEGGETGRSGSTLNGGGASEGNKGRLFNEASASHNTPKVEREEGELSPNGDFEEDNFA 802
Query: 231 VYANAGLEAVHKGKDCNTSQHYQNRREEQ--ICGVAGGEND----DESDGSPHRSSDDSE 284
+ + ++ V K K+ +TS+ +Q R E C A GEND DE + S RS++DSE
Sbjct: 803 PFEDGAVDGVSKAKEGSTSRPFQGRSGEAQPSCAEAAGENDADADDEGEESAQRSTEDSE 862
Query: 285 NASENG-DVSGTESADGEECSREEH---EEDGDHDN---KVESEGEAEGMADANDVEGDG 337
NASE G D SG+ES DGEECSRE+H EED DHD+ K ESEGEAEG + +DVEG G
Sbjct: 863 NASEGGEDASGSESGDGEECSREDHDEEEEDMDHDDQDAKAESEGEAEGTTETHDVEG-G 921
Query: 338 ASLPYSERFLLTAKPLVKYVSPVFHGKEE-NVQIFYGNDSFYVLFRLHQTLYERIRSAKI 396
SLP SERFL + KPL K+V H ++E + +IFYGNDSFYVLFRLHQ LYER+ SAK
Sbjct: 922 ISLPLSERFLHSVKPLAKHVPTALHDRDEKSSRIFYGNDSFYVLFRLHQILYERLLSAKT 981
Query: 397 NSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLD 456
NSSSAEKKWR S DT+ D YA+F+++LY+LLDGSSDN+KFEDDCR+IIGTQSYVLFTLD
Sbjct: 982 NSSSAEKKWRTSKDTNPPDLYAKFISALYNLLDGSSDNTKFEDDCRSIIGTQSYVLFTLD 1041
Query: 457 KLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRI 516
KLIYK+VKQLQA+A TDEMDNKLLQLY YE+SR G FFD+VYHENARVLLH+E+IYR
Sbjct: 1042 KLIYKVVKQLQAIA--TDEMDNKLLQLYLYEKSRSPGRFFDLVYHENARVLLHEESIYRF 1099
Query: 517 EC--SPTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKL 574
EC +PT+LSIQLM+YGH+KPEVTAVSIDPNFS+YL N++LS + D K G+FL+RNK
Sbjct: 1100 ECCSNPTKLSIQLMEYGHEKPEVTAVSIDPNFSSYLFNEYLSSMSDRKLSEGVFLERNKR 1159
Query: 575 KCARSDELSS--QVMDGIQVINGLECKIACGSSKV 607
K + +DE S + MDG++V NGLECKI+C +SKV
Sbjct: 1160 KHSNNDEPSDSLKAMDGVKVANGLECKISCKTSKV 1194
>ref|NP_908404.1| putative co-repressor protein [Oryza sativa (japonica
cultivar-group)]
Length = 1589
Score = 574 bits (1479), Expect = e-162
Identities = 333/635 (52%), Positives = 426/635 (66%), Gaps = 33/635 (5%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAK-GDSSPGVGATVMSPK 59
++IWTTFLEP+LGV R ED DAVK K + K+G A++ + ++ G A +
Sbjct: 928 VRIWTTFLEPILGVQPRTHGAEDA-DAVKPKSRTTKSGLATVGEINTTAAGAVAKHGHDE 986
Query: 60 NT-FEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSR 118
N EQ+ S NGV ++ DR+ + E ++ + G V + +E+
Sbjct: 987 NIPQEQTPSSLARMVNGVATDTQNGFHDVDRTARRAEEPSNTAVNGRVQGASPGTNEIPA 1046
Query: 119 VNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSR-----PGYVYREGGLDL 173
V+ Q+ E+ N+ ++ + N + N+ SG+ A+ S R G L
Sbjct: 1047 VSTQNMPTERSAE-NIPVARTEQHGNAKANLEPTSGVNASRSSHAGNDTAAEARAGNETL 1105
Query: 174 PSSEGADSTRPDTSTNGA-IIEDTKAHRCHKESVGHF--KSEREEGELSPNGDFEEDNFA 230
PS EG ++ R ++ NG E K ++ S H K EREEGELSPNGDFEEDNFA
Sbjct: 1106 PSVEGGETGRSGSTLNGGGASEGNKGRLFNEASASHNTPKVEREEGELSPNGDFEEDNFA 1165
Query: 231 VYANAGLEAVHKGKDCNTSQHYQNRREEQ--ICGVAGGEND----DESDGSPHRSSDDSE 284
+ + ++ V K K+ +TS+ +Q R E C A GEND DE + S RS++DSE
Sbjct: 1166 PFEDGAVDGVSKAKEGSTSRPFQGRSGEAQPSCAEAAGENDADADDEGEESAQRSTEDSE 1225
Query: 285 NASENG-DVSGTESADGEECSREEH---EEDGDHDN---KVESEGEAEGMADANDVEGDG 337
NASE G D SG+ES DGEECSRE+H EED DHD+ K ESEGEAEG + +DVEG G
Sbjct: 1226 NASEGGEDASGSESGDGEECSREDHDEEEEDMDHDDQDAKAESEGEAEGTTETHDVEG-G 1284
Query: 338 ASLPYSERFLLTAKPLVKYVSPVFHGKEE-NVQIFYGNDSFYVLFRLHQTLYERIRSAKI 396
SLP SERFL + KPL K+V H ++E + +IFYGNDSFYVLFRLHQ LYER+ SAK
Sbjct: 1285 ISLPLSERFLHSVKPLAKHVPTALHDRDEKSSRIFYGNDSFYVLFRLHQILYERLLSAKT 1344
Query: 397 NSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLD 456
NSSSAEKKWR S DT+ D YA+F+++LY+LLDGSSDN+KFEDDCR+IIGTQSYVLFTLD
Sbjct: 1345 NSSSAEKKWRTSKDTNPPDLYAKFISALYNLLDGSSDNTKFEDDCRSIIGTQSYVLFTLD 1404
Query: 457 KLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRI 516
KLIYK+VKQLQA+A TDEMDNKLLQLY YE+SR G FFD+VYHENARVLLH+E+IYR
Sbjct: 1405 KLIYKVVKQLQAIA--TDEMDNKLLQLYLYEKSRSPGRFFDLVYHENARVLLHEESIYRF 1462
Query: 517 EC--SPTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKL 574
EC +PT+LSIQLM+YGH+KPEVTAVSIDPNFS+YL N++LS + D K G+FL+RNK
Sbjct: 1463 ECCSNPTKLSIQLMEYGHEKPEVTAVSIDPNFSSYLFNEYLSSMSDRKLSEGVFLERNKR 1522
Query: 575 KCARSDELSS--QVMDGIQVINGLECKIACGSSKV 607
K + +DE S + MDG++V NGLECKI+C +SKV
Sbjct: 1523 KHSNNDEPSDSLKAMDGVKVANGLECKISCKTSKV 1557
>ref|NP_173829.2| paired amphipathic helix repeat-containing protein [Arabidopsis
thaliana]
Length = 1353
Score = 523 bits (1346), Expect = e-147
Identities = 311/621 (50%), Positives = 390/621 (62%), Gaps = 32/621 (5%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MKIWTTF+E + GVPSR + ED ED VK+ + K+G++S + + SP A+V +
Sbjct: 710 MKIWTTFVEQIFGVPSRPQGAEDQEDVVKSMNQNVKSGSSSAGESEGSPHNYASVADSRR 769
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVN 120
+ + S E G E D S ET + NV + P+ +
Sbjct: 770 S-KSSRKANEHSQLGQTSNSERDGAAGRTSDALCETAQHEKMLKNVVTSDEKPE-----S 823
Query: 121 KQDHSIEQLVNAN-VSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGA 179
KQ SIE+ ++ +++ ++QSNG ++I + +G +P E L + G
Sbjct: 824 KQAVSIERAHDSTALAVDGLLDQSNGGSSIVHMTGHCNNNLKPVTCGTELELKMNDGNGP 883
Query: 180 --DSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGL 237
+ TNG +E T +E G K EREEGELSPNGDFEEDNFAVYA
Sbjct: 884 KLEVGNKKLLTNGIAVEITS----DQEMAGTSKVEREEGELSPNGDFEEDNFAVYAKTDF 939
Query: 238 EAVHKGKDC--NTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGT 295
E K D N ++R E C END E D + RSS+DS N ENGDVSGT
Sbjct: 940 ETFSKANDSTGNNISGDRSREGEPSCLETRAENDAEGDENAARSSEDSRNEYENGDVSGT 999
Query: 296 ESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPLVK 355
ES GE+ E+D D++NK ESEGEAE MADA+D E +G++LP S RFLL KPLVK
Sbjct: 1000 ESGGGED-----PEDDLDNNNKGESEGEAECMADAHDAEENGSALPVSARFLLHVKPLVK 1054
Query: 356 YVSPVF--HGKEE----NVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASN 409
YV H K++ N Q+FYGNDSFYVLFRLH+ LYERI SAK+NSSS E KWR SN
Sbjct: 1055 YVPSAIALHDKDKDSLKNSQVFYGNDSFYVLFRLHRILYERILSAKVNSSSPEGKWRTSN 1114
Query: 410 DTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAV 469
+ TD YARFM +LY+LLDG+SDN+KFEDDCRAIIGTQSY+LFTLDKLI+K +K LQ V
Sbjct: 1115 TKNPTDSYARFMTALYNLLDGTSDNAKFEDDCRAIIGTQSYILFTLDKLIHKFIKHLQVV 1174
Query: 470 ASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIEC---SPTRLSIQ 526
+ DEMDNKLLQLY YE+SR+ + FD VY++N RVLL DENIYRIEC +P +LSIQ
Sbjct: 1175 VA--DEMDNKLLQLYFYEKSRRPETIFDAVYYDNTRVLLPDENIYRIECRLSTPAKLSIQ 1232
Query: 527 LMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQV 586
LM G DKP+VT+VSIDP F+ YLHNDFLS+ P+++E I+L RNK K R DE
Sbjct: 1233 LMCNGLDKPDVTSVSIDPTFAAYLHNDFLSIQPNAREDRRIYLNRNKRKVCREDE-QLYS 1291
Query: 587 MDGIQVINGLECKIACGSSKV 607
D +++ NGLECKIACGSSKV
Sbjct: 1292 TDEVKIKNGLECKIACGSSKV 1312
>ref|NP_177163.2| paired amphipathic helix repeat-containing protein [Arabidopsis
thaliana]
Length = 1362
Score = 490 bits (1261), Expect = e-137
Identities = 309/633 (48%), Positives = 389/633 (60%), Gaps = 74/633 (11%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MK+WT FLEP+ GVPSR + ED EDAVK+ + ++ SP GA++ N
Sbjct: 726 MKVWTEFLEPIFGVPSRPQGAEDREDAVKSTNHDREDQEDAV-----SPQNGASIA---N 777
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGS--STLQGNVHINASIPDEVSR 118
+ + K ++N V E D K S ++ L S +T + N + PDE +
Sbjct: 778 SMRSNGPRKVNESNQVRQASELD--KDVTSSKTSDALLSCDNTQNDKMPKNLTTPDERAE 835
Query: 119 VNKQDHSIEQLVNAN-VSMSSRVEQSNGRTNINNAS------------GLAATLSRPGYV 165
KQ SIE+ N+N + + + Q NG+ + + + GL+ + +P
Sbjct: 836 T-KQAVSIERAHNSNALPLDGLLPQRNGKISSLSVADEEWYPFLLYSPGLSNSNPKPALT 894
Query: 166 YREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFE 225
G + + R + N I T A + G K EREEGELSP GDFE
Sbjct: 895 ---SGTEELKPNYVNGPRVEIGDNPVIPNGTVA----EWFAGEAKVEREEGELSPTGDFE 947
Query: 226 EDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESD-GSPHRSSDDSE 284
EDN+AV+ +EA+ K K END +D S RSSD S
Sbjct: 948 EDNYAVHGENDMEALSKSK----------------------ENDATADDASAPRSSDGSG 985
Query: 285 NASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAE-GMADAND-VEGDGASLPY 342
N S NGDVSGT+S DGE+C RE+ D DH NKVESEGEAE GM+D +D EGD L
Sbjct: 986 NTSHNGDVSGTDSGDGEDCYRED---DIDH-NKVESEGEAEEGMSDGHDDTEGDMPVLSI 1041
Query: 343 SERFLLTAKPLVKYVSPVFHGKE-----ENVQIFYGNDSFYVLFRLHQTLYERIRSAKIN 397
S + LL KPL KYV P + K+ +N Q+FYGNDSFYVLFRLHQ LY+RI SAKIN
Sbjct: 1042 SVKNLLHVKPLAKYVPPALYDKDNDDSRKNSQVFYGNDSFYVLFRLHQILYDRILSAKIN 1101
Query: 398 SSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDK 457
SSS ++KW+ SN T+ D YAR M++LY+LLDG+SDNSKFEDDCRAIIGTQSYVLFTLDK
Sbjct: 1102 SSSPDRKWKTSNPTNPADSYARIMDALYNLLDGTSDNSKFEDDCRAIIGTQSYVLFTLDK 1161
Query: 458 LIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIE 517
LIYKL+K LQAVA+ DEMDNKL QLYAYE+SRK F D VY+ENA VLL DE+IYRIE
Sbjct: 1162 LIYKLIKHLQAVAA--DEMDNKLQQLYAYEKSRKPEKFLDAVYYENALVLLPDEDIYRIE 1219
Query: 518 C---SPTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKL 574
C +P++LSIQL+DYGHDKP+VT++S+DP F+ YLHN FLS P++KE I+LKRNK
Sbjct: 1220 CEQSTPSKLSIQLLDYGHDKPDVTSISMDPTFAAYLHNVFLSYQPNAKENPRIYLKRNKR 1279
Query: 575 KCARSDELSSQVMDGIQVINGLECKIACGSSKV 607
K DEL + D +++INGLECKI C SSKV
Sbjct: 1280 KNGGDDELCT--TDEVKIINGLECKITCSSSKV 1310
>pir||T00649 hypothetical protein F3I6.12 - Arabidopsis thaliana
gi|2829870|gb|AAC00578.1| Hypothetical protein
[Arabidopsis thaliana]
Length = 1263
Score = 486 bits (1252), Expect = e-136
Identities = 290/585 (49%), Positives = 366/585 (61%), Gaps = 31/585 (5%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MKIWTTF+E + GVPSR + ED ED VK+ + K+G++S + + SP A+V +
Sbjct: 696 MKIWTTFVEQIFGVPSRPQGAEDQEDVVKSMNQNVKSGSSSAGESEGSPHNYASVADSRR 755
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVN 120
+ + S E G E D S ET + NV + P+ +
Sbjct: 756 S-KSSRKANEHSQLGQTSNSERDGAAGRTSDALCETAQHEKMLKNVVTSDEKPE-----S 809
Query: 121 KQDHSIEQLVNAN-VSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGA 179
KQ SIE+ ++ +++ ++QSNG ++I + +G +P E L + G
Sbjct: 810 KQAVSIERAHDSTALAVDGLLDQSNGGSSIVHMTGHCNNNLKPVTCGTELELKMNDGNGP 869
Query: 180 --DSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGL 237
+ TNG +E T +E G K EREEGELSPNGDFEEDNFAVYA
Sbjct: 870 KLEVGNKKLLTNGIAVEITS----DQEMAGTSKVEREEGELSPNGDFEEDNFAVYAKTDF 925
Query: 238 EAVHKGKDC--NTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGT 295
E K D N ++R E C END E D + RSS+DS N ENGDVSGT
Sbjct: 926 ETFSKANDSTGNNISGDRSREGEPSCLETRAENDAEGDENAARSSEDSRNEYENGDVSGT 985
Query: 296 ESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPLVK 355
ES GE+ E+D D++NK ESEGEAE MADA+D E +G++LP S RFLL KPLVK
Sbjct: 986 ESGGGED-----PEDDLDNNNKGESEGEAECMADAHDAEENGSALPVSARFLLHVKPLVK 1040
Query: 356 YVSPVF--HGKEE----NVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASN 409
YV H K++ N Q+FYGNDSFYVLFRLH+ LYERI SAK+NSSS E KWR SN
Sbjct: 1041 YVPSAIALHDKDKDSLKNSQVFYGNDSFYVLFRLHRILYERILSAKVNSSSPEGKWRTSN 1100
Query: 410 DTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAV 469
+ TD YARFM +LY+LLDG+SDN+KFEDDCRAIIGTQSY+LFTLDKLI+K +K LQ V
Sbjct: 1101 TKNPTDSYARFMTALYNLLDGTSDNAKFEDDCRAIIGTQSYILFTLDKLIHKFIKHLQVV 1160
Query: 470 ASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIEC---SPTRLSIQ 526
+ DEMDNKLLQLY YE+SR+ + FD VY++N RVLL DENIYRIEC +P +LSIQ
Sbjct: 1161 VA--DEMDNKLLQLYFYEKSRRPETIFDAVYYDNTRVLLPDENIYRIECRLSTPAKLSIQ 1218
Query: 527 LMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKR 571
LM G DKP+VT+VSIDP F+ YLHNDFLS+ P+++E I+L R
Sbjct: 1219 LMCNGLDKPDVTSVSIDPTFAAYLHNDFLSIQPNAREDRRIYLNR 1263
>pir||C96723 hypothetical protein F20P5.21 [imported] - Arabidopsis thaliana
gi|2194132|gb|AAB61107.1| F20P5.21 gene product
[Arabidopsis thaliana]
Length = 1383
Score = 483 bits (1242), Expect = e-134
Identities = 310/646 (47%), Positives = 390/646 (59%), Gaps = 87/646 (13%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MK+WT FLEP+ GVPSR + ED EDAVK+ + ++ SP GA++ N
Sbjct: 737 MKVWTEFLEPIFGVPSRPQGAEDREDAVKSTNHDREDQEDAV-----SPQNGASIA---N 788
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGS--STLQGNVHINASIPDEVSR 118
+ + K ++N V E D K S ++ L S +T + N + PDE +
Sbjct: 789 SMRSNGPRKVNESNQVRQASELD--KDVTSSKTSDALLSCDNTQNDKMPKNLTTPDERAE 846
Query: 119 VNKQDHSIEQLVNAN-VSMSSRVEQSNGRTNINNAS------------GLAATLSRPGYV 165
KQ SIE+ N+N + + + Q NG+ + + + GL+ + +P
Sbjct: 847 T-KQAVSIERAHNSNALPLDGLLPQRNGKISSLSVADEEWYPFLLYSPGLSNSNPKPALT 905
Query: 166 YREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFE 225
G + + R + N I T A + G K EREEGELSP GDFE
Sbjct: 906 ---SGTEELKPNYVNGPRVEIGDNPVIPNGTVA----EWFAGEAKVEREEGELSPTGDFE 958
Query: 226 EDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESD-GSPHRSSDDSE 284
EDN+AV+ +EA+ K K END +D S RSSD S
Sbjct: 959 EDNYAVHGENDMEALSKSK----------------------ENDATADDASAPRSSDGSG 996
Query: 285 NASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAE-GMADAND-VEGDGASLPY 342
N S NGDVSGT+S DGE+C RE+ D DH NKVESEGEAE GM+D +D EGD L
Sbjct: 997 NTSHNGDVSGTDSGDGEDCYRED---DIDH-NKVESEGEAEEGMSDGHDDTEGDMPVLSI 1052
Query: 343 SERFLLTAKPLVKYVSPVFHGKE-----ENVQIFYGNDSFYVLFRLHQTLYERIRSAKIN 397
S + LL KPL KYV P + K+ +N Q+FYGNDSFYVLFRLHQ LY+RI SAKIN
Sbjct: 1053 SVKNLLHVKPLAKYVPPALYDKDNDDSRKNSQVFYGNDSFYVLFRLHQILYDRILSAKIN 1112
Query: 398 SSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDK 457
SSS ++KW+ SN T+ D YAR M++LY+LLDG+SDNSKFEDDCRAIIGTQSYVLFTLDK
Sbjct: 1113 SSSPDRKWKTSNPTNPADSYARIMDALYNLLDGTSDNSKFEDDCRAIIGTQSYVLFTLDK 1172
Query: 458 LIYKLVK-------------QLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENA 504
LIYKL+K QLQAVA+ DEMDNKL QLYAYE+SRK F D VY+ENA
Sbjct: 1173 LIYKLIKHSSHMIQLTNVILQLQAVAA--DEMDNKLQQLYAYEKSRKPEKFLDAVYYENA 1230
Query: 505 RVLLHDENIYRIEC---SPTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDS 561
VLL DE+IYRIEC +P++LSIQL+DYGHDKP+VT++S+DP F+ YLHN FLS P++
Sbjct: 1231 LVLLPDEDIYRIECEQSTPSKLSIQLLDYGHDKPDVTSISMDPTFAAYLHNVFLSYQPNA 1290
Query: 562 KEKSGIFLKRNKLKCARSDELSSQVMDGIQVINGLECKIACGSSKV 607
KE I+LKRNK K DEL + D +++INGLECKI C SSKV
Sbjct: 1291 KENPRIYLKRNKRKNGGDDELCT--TDEVKIINGLECKITCSSSKV 1334
>ref|XP_493857.1| Similar to Arabidopsis chromosome I BAC genomic sequence (AC002396);
unknown protein [Oryza sativa]
Length = 1056
Score = 422 bits (1086), Expect = e-116
Identities = 276/586 (47%), Positives = 345/586 (58%), Gaps = 92/586 (15%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MKIW TFLEP+LGV R ED K S K G A++ ++S
Sbjct: 522 MKIWATFLEPILGVHPRGHGVEDE----KHNSRSTKAGPANVEINNAS------------ 565
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGS-STLQGNVHINA--SIPDEVS 117
TNG VK SD VPK + S + L G V +A S+ D
Sbjct: 566 ------------TNGTVTVKH---AHSDEIVPKEQASCSRAILVGGVAADAQNSLQDAER 610
Query: 118 RVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSE 177
V + + + +++ + ++ S G + +P S
Sbjct: 611 TVCRDEERPKTMLDRRLQNTTPAVDS-------------------------GEIRIPGSF 645
Query: 178 GADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGL 237
+ + + I E +H H K EREEGELSPNGD E NF + +
Sbjct: 646 NSKDNK-----HCPINEYCGSHN-------HSKVEREEGELSPNGDVGE-NFGPFDGVSV 692
Query: 238 EAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSS---DDSENASENG-DVS 293
+ V K K+ +T + Q R + GEND ++D S+ +DSENASE G D S
Sbjct: 693 DGVSKAKEDSTRRLLQGRPMDAT--EFAGENDVDADDEGEESAQMMEDSENASEAGEDAS 750
Query: 294 GTESADGEECSREEHEEDGDHDN-----KVESEGEAEGMADANDVEGDGASLPYSERFLL 348
G+ES DGEECSRE+HE++ D D K ESEGEA +A D + G SLP+SER
Sbjct: 751 GSESGDGEECSREDHEDEDDMDQDDPDAKAESEGEAAENTEAQDADA-GISLPFSERSHN 809
Query: 349 TAKPLVKYVSPVFHGKEENVQ-IFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRA 407
KPL K+V + EE IFYGNDSFYVLFRLHQ LYERI SAK NSSSAEKKW+A
Sbjct: 810 AVKPLAKHVPRALNDHEEKFSCIFYGNDSFYVLFRLHQILYERILSAKTNSSSAEKKWKA 869
Query: 408 SNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQ 467
S DT+ D+Y++FM++LY+LLDGSSDN+KFEDDCR+IIGTQSYVLFTLDKLIYK+ LQ
Sbjct: 870 SKDTNLPDQYSKFMSALYNLLDGSSDNTKFEDDCRSIIGTQSYVLFTLDKLIYKV---LQ 926
Query: 468 AVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIE--CSPTRLSI 525
A+AS DEMDNKLLQLY YE+SR G FFD+VYHENARVLLHDE+IYR E +PTRLSI
Sbjct: 927 AIAS--DEMDNKLLQLYIYEKSRSPGRFFDLVYHENARVLLHDESIYRFERRSNPTRLSI 984
Query: 526 QLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKR 571
QLM+YGH+KPEVTAVSIDPNFS+YL+N++LS + ++K IFL R
Sbjct: 985 QLMEYGHEKPEVTAVSIDPNFSSYLYNEYLSSISNTKLYDDIFLGR 1030
>ref|NP_186781.3| paired amphipathic helix repeat-containing protein [Arabidopsis
thaliana]
Length = 1378
Score = 399 bits (1024), Expect = e-109
Identities = 262/641 (40%), Positives = 373/641 (57%), Gaps = 71/641 (11%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKK------DSAKTGTASIAKGDSSPGVGAT 54
+++W FLE +LGVP R + + ED V K ++ G A+++ G + + +
Sbjct: 736 LRLWENFLEAVLGVPPRAKGTDLVEDVVINPKTLDVNHSTSPNGEAAVSSGGDTARLASR 795
Query: 55 VMSPKNTFEQSNSCKEWQTNGVGGVKEDDCLKS---DRSVPKTETLGSSTLQ-------G 104
+ ++++S ++ +G+G + +D K D + + + S ++ G
Sbjct: 796 KLKSAANGDENSSSGTFK-HGIGLLNKDSTGKENLEDVEIANRDGVACSAVKPQKEQETG 854
Query: 105 NV---HINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSR 161
N IP ++S E+ +++S+ S E ++ G ++
Sbjct: 855 NEAEKRFGKPIPMDIS---------ERAAISSISIPSGAENNHCVVGKEVLPGAHEIQAK 905
Query: 162 PGYVYREGGLDLPSSEGADSTRPDTSTNGAII------EDTKAHRCHKESVGHFKSEREE 215
P + D+ S E ST+ N ++ + +K R + G ++E+EE
Sbjct: 906 PSDTLTDIHHDVDSIETVHSTQGGDVGNSIVLANGLRSDSSKGTRNSDDPEGPSRNEKEE 965
Query: 216 GELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGS 275
GELSPNGDFE DNF VY + G+++ K +N E ++ A EN+D++D
Sbjct: 966 GELSPNGDFE-DNFGVYKDHGVKSTSKP---------ENSAEAEVEADAEVENEDDADDV 1015
Query: 276 PHRSSDDSENASENGDVSGTESADGEECSREE--HEEDGDHDN---KVESEGEAEGMADA 330
DSENASE SGTES G+ CS++E EE+G+HD K ESEGEAEGM D
Sbjct: 1016 ------DSENASE---ASGTESG-GDVCSQDEDREEENGEHDEIDGKAESEGEAEGM-DP 1064
Query: 331 NDVEGDGASLPYSERFLLTAKPLVKYVSPVF-HGKEENVQIFYGNDSFYVLFRLHQTLYE 389
+ +EG+ LP SER LL+ +PL K+V+ V + +++Q+FYGND FYVLFRLHQ LYE
Sbjct: 1065 HLLEGESELLPQSERVLLSVRPLSKHVAAVLCDERTKDLQVFYGNDDFYVLFRLHQILYE 1124
Query: 390 RIRSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQS 449
RI AK N S E K + DT++ D YARFM LY LLDGS++N+KFED+CRAIIG QS
Sbjct: 1125 RILYAKRNCSGGELKSKNLKDTNAGDPYARFMRVLYGLLDGSAENTKFEDECRAIIGNQS 1184
Query: 450 YVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLH 509
YVLFTLDKLIY+LVKQLQA+ + DEMDNKLLQLY YE+SRK G D VY+EN RVL+H
Sbjct: 1185 YVLFTLDKLIYRLVKQLQAIVA--DEMDNKLLQLYEYEKSRKPGRVIDSVYYENVRVLVH 1242
Query: 510 DENIYRIECS--PTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKS-G 566
+ENIYR+ECS P+RLSIQLMD +KPE AVS+DP F++Y+ + LSV KE+
Sbjct: 1243 EENIYRLECSSLPSRLSIQLMDNIIEKPEAYAVSMDPTFASYMQTELLSVSSGKKEEGHD 1302
Query: 567 IFLKRNKLKCARSDELSSQVMDGIQVINGLECKIACGSSKV 607
I L+RN + M+G++V+NGLECK++C S K+
Sbjct: 1303 IVLQRNLTGLYD----LCKAMEGVEVVNGLECKMSCSSYKI 1339
>gb|AAF03494.1| unknown protein [Arabidopsis thaliana]
Length = 1324
Score = 379 bits (972), Expect = e-103
Identities = 260/619 (42%), Positives = 352/619 (56%), Gaps = 86/619 (13%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
+++W FLE +LGVP R + + ED V K T + +SP A V S +
Sbjct: 734 LRLWENFLEAVLGVPPRAKGTDLVEDVVINPK------TLDV-NHSTSPNGEAAVSSGGD 786
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVN 120
T ++ + NG E S T + + + +N
Sbjct: 787 TARLASRKLKSAANG------------------DENSSSGTFKHGIGL----------LN 818
Query: 121 KQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGAD 180
K E L + ++ V S + +G A + G +P D
Sbjct: 819 KDSTGKENLEDVEIANRDGVACSAVKPQKEQETGNEAE--------KRFGKPIPM----D 866
Query: 181 STRPDTSTNGAIIEDTKAHRC--HKESV-GHFKSEREEGELSPNGDFEEDNFAVYANAGL 237
+ ++ +I + + C KE + G ++E+EEGELSPNGDFE DNF VY + G+
Sbjct: 867 ISERAAISSISIPSGAENNHCVVGKEVLPGPSRNEKEEGELSPNGDFE-DNFGVYKDHGV 925
Query: 238 EAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTES 297
++ K +N E ++ A EN+D++D DSENASE SGTES
Sbjct: 926 KSTSKP---------ENSAEAEVEADAEVENEDDADDV------DSENASE---ASGTES 967
Query: 298 ADGEECSREE--HEEDGDHDN---KVESEGEAEGMADANDVEGDGASLPYSERFLLTAKP 352
G+ CS++E EE+G+HD K ESEGEAEGM D + +EG+ LP SER LL+ +P
Sbjct: 968 G-GDVCSQDEDREEENGEHDEIDGKAESEGEAEGM-DPHLLEGESELLPQSERVLLSVRP 1025
Query: 353 LVKYVSPVF-HGKEENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDT 411
L K+V+ V + +++Q+FYGND FYVLFRLHQ LYERI AK N S E K + DT
Sbjct: 1026 LSKHVAAVLCDERTKDLQVFYGNDDFYVLFRLHQILYERILYAKRNCSGGELKSKNLKDT 1085
Query: 412 SSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVAS 471
++ D YARFM LY LLDGS++N+KFED+CRAIIG QSYVLFTLDKLIY+LVKQLQA+ +
Sbjct: 1086 NAGDPYARFMRVLYGLLDGSAENTKFEDECRAIIGNQSYVLFTLDKLIYRLVKQLQAIVA 1145
Query: 472 GTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECS--PTRLSIQLMD 529
DEMDNKLLQLY YE+SRK G D VY+EN RVL+H+ENIYR+ECS P+RLSIQLMD
Sbjct: 1146 --DEMDNKLLQLYEYEKSRKPGRVIDSVYYENVRVLVHEENIYRLECSSLPSRLSIQLMD 1203
Query: 530 YGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKS-GIFLKRNKLKCARSDELSSQVMD 588
+KPE AVS+DP F++Y+ + LSV KE+ I L+RN + M+
Sbjct: 1204 NIIEKPEAYAVSMDPTFASYMQTELLSVSSGKKEEGHDIVLQRNLTGLYD----LCKAME 1259
Query: 589 GIQVINGLECKIACGSSKV 607
G++V+NGLECK++C S K+
Sbjct: 1260 GVEVVNGLECKMSCSSYKI 1278
>ref|NP_176197.2| paired amphipathic helix repeat-containing protein [Arabidopsis
thaliana]
Length = 1159
Score = 375 bits (962), Expect = e-102
Identities = 224/448 (50%), Positives = 288/448 (64%), Gaps = 33/448 (7%)
Query: 175 SSEGADSTRPDTSTNGAIIEDTKAH-RCHKESVGHFKSEREEGELSPNGDFEEDNFAVYA 233
++E + ++ ++ N I+E + R S+G K EREEGELSP E++NF VY
Sbjct: 673 TNEDSQPSKLVSTRNDLIMEGVENRSRVSDVSMGGHKVEREEGELSPTESCEQENFEVYK 732
Query: 234 NAGLEAVHKGKDCNTSQHYQNRREEQICGV-----AGGENDDESDGSPHRSSDDSENASE 288
GLE V K D N + +E CG + +D+ + + S+ ENAS+
Sbjct: 733 ENGLEPVQKLPD-NEISNTDREPKEGACGTEAVTRSNALPEDDDNKITQKLSEGDENASK 791
Query: 289 NGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLL 348
+ G+ S EEH+ ESE EA GM ++N+ E DG+ +SER+L
Sbjct: 792 F--IVSASKFGGQVSSDEEHKG-------AESENEAGGMVNSNEGE-DGSFFTFSERYLQ 841
Query: 349 TAKPLVKYVSPVFHGKE----ENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKK 404
KPL K+V E + ++FYGNDS YVLFRLHQ LYERI+SAKI+S E+K
Sbjct: 842 PVKPLAKHVPGTLQASECDTRNDSRVFYGNDSLYVLFRLHQMLYERIQSAKIHS---ERK 898
Query: 405 WRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVK 464
W+A D++STD Y RFM +LY+LLDGSSDN+KFED+CRAIIG QSYVLFTLDKL+ K VK
Sbjct: 899 WKAP-DSTSTDSYTRFMEALYNLLDGSSDNTKFEDECRAIIGAQSYVLFTLDKLVQKFVK 957
Query: 465 QLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECS--PTR 522
L AVA+ DE D KLLQLYAYE RK G FFD+VYHENAR LLHD+NIYRIE S TR
Sbjct: 958 HLHAVAA--DETDTKLLQLYAYENYRKPGRFFDIVYHENARALLHDQNIYRIEYSSAQTR 1015
Query: 523 LSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDEL 582
L IQLM+ +D+PEVTAV+++P F+ YL NDFLS V D +EK G+FLKRNK K + E
Sbjct: 1016 LGIQLMNSWNDQPEVTAVTVEPGFANYLQNDFLSFVSD-EEKPGLFLKRNKAKLSGPGEE 1074
Query: 583 S---SQVMDGIQVINGLECKIACGSSKV 607
S S+ ++G+ +IN +ECKIAC S KV
Sbjct: 1075 SLGMSRALEGLNIINEVECKIACSSFKV 1102
>gb|AAD39330.1| Hypothetical protein [Arabidopsis thaliana] gi|25404243|pir||A96623
hypothetical protein F23H11.20 [imported] - Arabidopsis
thaliana
Length = 1108
Score = 370 bits (951), Expect = e-101
Identities = 221/443 (49%), Positives = 285/443 (63%), Gaps = 33/443 (7%)
Query: 175 SSEGADSTRPDTSTNGAIIEDTKAH-RCHKESVGHFKSEREEGELSPNGDFEEDNFAVYA 233
++E + ++ ++ N I+E + R S+G K EREEGELSP E++NF VY
Sbjct: 634 TNEDSQPSKLVSTRNDLIMEGVENRSRVSDVSMGGHKVEREEGELSPTESCEQENFEVYK 693
Query: 234 NAGLEAVHKGKDCNTSQHYQNRREEQICGV-----AGGENDDESDGSPHRSSDDSENASE 288
GLE V K D N + +E CG + +D+ + + S+ ENAS+
Sbjct: 694 ENGLEPVQKLPD-NEISNTDREPKEGACGTEAVTRSNALPEDDDNKITQKLSEGDENASK 752
Query: 289 NGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLL 348
+ G+ S EEH+ ESE EA GM ++N+ E DG+ +SER+L
Sbjct: 753 F--IVSASKFGGQVSSDEEHKG-------AESENEAGGMVNSNEGE-DGSFFTFSERYLQ 802
Query: 349 TAKPLVKYVSPVFHGKE----ENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKK 404
KPL K+V E + ++FYGNDS YVLFRLHQ LYERI+SAKI+S E+K
Sbjct: 803 PVKPLAKHVPGTLQASECDTRNDSRVFYGNDSLYVLFRLHQMLYERIQSAKIHS---ERK 859
Query: 405 WRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVK 464
W+A D++STD Y RFM +LY+LLDGSSDN+KFED+CRAIIG QSYVLFTLDKL+ K VK
Sbjct: 860 WKAP-DSTSTDSYTRFMEALYNLLDGSSDNTKFEDECRAIIGAQSYVLFTLDKLVQKFVK 918
Query: 465 QLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECS--PTR 522
L AVA+ DE D KLLQLYAYE RK G FFD+VYHENAR LLHD+NIYRIE S TR
Sbjct: 919 HLHAVAA--DETDTKLLQLYAYENYRKPGRFFDIVYHENARALLHDQNIYRIEYSSAQTR 976
Query: 523 LSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDEL 582
L IQLM+ +D+PEVTAV+++P F+ YL NDFLS V D +EK G+FLKRNK K + E
Sbjct: 977 LGIQLMNSWNDQPEVTAVTVEPGFANYLQNDFLSFVSD-EEKPGLFLKRNKAKLSGPGEE 1035
Query: 583 S---SQVMDGIQVINGLECKIAC 602
S S+ ++G+ +IN +ECKIAC
Sbjct: 1036 SLGMSRALEGLNIINEVECKIAC 1058
>emb|CAC01821.1| transcriptional regulatory-like protein [Arabidopsis thaliana]
gi|15242192|ref|NP_197006.1| paired amphipathic helix
repeat-containing protein [Arabidopsis thaliana]
gi|11358896|pir||T51447 transcription regulator-like
protein - Arabidopsis thaliana
Length = 1377
Score = 360 bits (924), Expect = 8e-98
Identities = 255/634 (40%), Positives = 350/634 (54%), Gaps = 85/634 (13%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
+K+W FLE ML V R + + ED V+ + A T + D+ V + N
Sbjct: 759 LKLWANFLELMLDVAPRAKGSDSVEDVVETQHQRAFTSGEANESSDAISLVSRQLKFATN 818
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVN 120
++S GV E L D S K +V A P + +
Sbjct: 819 GDVHASS-------GVSKHGETGLLNRDSS-GKENLKDGDLANKDVATCAEKPQKDQEIG 870
Query: 121 ----KQDHSI-EQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPS 175
K+ + E++ ++ S S VE +NG+ ++SG LS+P + +
Sbjct: 871 NGAAKRSGDVDERVATSSSSFPSGVENNNGKVGSRDSSGSRGILSKPSEAIDKVD-SIQH 929
Query: 176 SEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANA 235
++G D R NG + +KA+ + ES G K E+EEGELSP GD EDNF VY +
Sbjct: 930 TQGVDIGRIIVLGNGLQSDTSKANSNYDESGGPSKIEKEEGELSPVGD-SEDNFVVYEDR 988
Query: 236 GLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENG-DVSG 294
L+A K ++H A GEND+++D +D ++ASE G D SG
Sbjct: 989 ELKATAK------TEHSVE---------AEGENDEDAD------DEDGDDASEAGEDASG 1027
Query: 295 TESADGEECSREEH--EEDGDHDN---KVESEGEAEGMADANDVEGDGASLPYSERFLLT 349
TES G+ECS++++ EE+G+HD K ESEGEAEGM +++ +E G P SER LL+
Sbjct: 1028 TESI-GDECSQDDNGVEEEGEHDEIDGKAESEGEAEGM-ESHLIEDKGL-FPSSERVLLS 1084
Query: 350 AKPLVKYVSP--VFHGKEENVQIFYGNDSFYVLFRLHQT------------LYERIRSAK 395
KPL K+++ + K+++ ++FYGND FYVLFRLH+ LYERI SAK
Sbjct: 1085 VKPLSKHIAAAALVDEKKKDSRVFYGNDDFYVLFRLHRVSAIDSYDLLSHILYERILSAK 1144
Query: 396 INSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTL 455
S +E K R + DT S D YARFMN+L+SLL+GS++NSKFED+CRAIIG QSYVLFTL
Sbjct: 1145 TYCSGSEMKLRNTKDTCSPDPYARFMNALFSLLNGSAENSKFEDECRAIIGNQSYVLFTL 1204
Query: 456 DKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYR 515
+KLIYKLVKQLQAV + D+MDNKLLQLY YE SR+ G
Sbjct: 1205 EKLIYKLVKQLQAVVA--DDMDNKLLQLYEYENSRRPGR--------------------- 1241
Query: 516 IECSPTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLK 575
SP+RLSIQLMD +KP+ AVS++P F++YL N+FLS KE I L+RN
Sbjct: 1242 -SSSPSRLSIQLMDNIIEKPDAYAVSMEPTFTSYLQNEFLSNSSGKKELQDIVLQRNMRG 1300
Query: 576 CARSDEL--SSQVMDGIQVINGLECKIACGSSKV 607
D+L + + M+G+QVINGLECK++C S K+
Sbjct: 1301 YNGLDDLAVACKAMEGVQVINGLECKMSCSSYKI 1334
>ref|NP_172515.1| paired amphipathic helix repeat-containing protein [Arabidopsis
thaliana]
Length = 1173
Score = 323 bits (827), Expect = 1e-86
Identities = 232/622 (37%), Positives = 322/622 (51%), Gaps = 120/622 (19%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MK+W TFLEP+ G+ SR + ED K KN
Sbjct: 610 MKLWITFLEPIFGILSRSQDNLALEDVSKL----------------------------KN 641
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSST---LQGNVHINASIPDEVS 117
E ++C + G ++ S T+ GSS+ + N+ + + PD++
Sbjct: 642 NRELQDACLAVKETASGSNRKHPISPKRLSKDNTKMQGSSSREDVSANIKVKTAQPDKL- 700
Query: 118 RVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSE 177
QD + +M++ V QS+ +
Sbjct: 701 ----QD---------DAAMTNEVIQSSKFVS----------------------------- 718
Query: 178 GADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGL 237
N I+ED H + SV K E EEGELSP E+ NF V
Sbjct: 719 ---------PKNDQIMEDEGNHMVNAASVE--KHELEEGELSPTASREQSNFEVNGQNAF 767
Query: 238 EAVHKGKDCNTSQHYQNRREEQICGVAGGEN----DDESDGSPHRSSDDSENASENGDVS 293
+ + K D + ++ +++Q C G +N +D+ + H+ S++++ ASE VS
Sbjct: 768 KPLQKVTD-----NVRSNKDKQSCDKKGAKNKTRAEDDKQENCHKLSENNKTASEML-VS 821
Query: 294 GTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPL 353
GT+ + EE +R + G G AN +G+ S +SERFL T KP+
Sbjct: 822 GTKVSCHEENNRVMN---------CNGRGSVAGEM-ANGNQGEDGSFAFSERFLQTVKPV 871
Query: 354 VKYVSPVFHGKE----ENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASN 409
K++S E + Q+FYGNDS+YVLFRLHQ LYERI++AK +S EKKW+A++
Sbjct: 872 AKHLSWPLQASETCSQNDSQVFYGNDSYYVLFRLHQMLYERIQTAKKHS---EKKWKAAD 928
Query: 410 DTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAV 469
+T+ D Y RFM++LY+LLDGS DN+KFED+CRAI G QSYVLFTLDKL+ K VK L +V
Sbjct: 929 NTTP-DSYPRFMDALYNLLDGSIDNTKFEDECRAIFGAQSYVLFTLDKLVQKFVKHLHSV 987
Query: 470 ASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECSP--TRLSIQL 527
AS DE D KLLQL+AYE RK G FFD+VYHENA LLH+ NIYRI S TRLSIQL
Sbjct: 988 AS--DETDTKLLQLHAYENYRKPGKFFDLVYHENACALLHEANIYRIRYSSEGTRLSIQL 1045
Query: 528 MDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQ-- 585
M+ G+++ EV V+++P F+ YL N L V D +E G+FL RNK K DE
Sbjct: 1046 MNSGNNQLEVMGVAMEPAFADYLQNKCLKSVND-EENHGLFLNRNKKKFTSLDESRGMPV 1104
Query: 586 VMDGIQVINGLECKIACGSSKV 607
M+ + +IN +EC++AC SSKV
Sbjct: 1105 AMERLNIINEMECRMACSSSKV 1126
>gb|AAD39565.1| T10O24.5 [Arabidopsis thaliana] gi|25402600|pir||C86238 protein
T10O24.5 [imported] - Arabidopsis thaliana
Length = 1164
Score = 322 bits (824), Expect = 3e-86
Identities = 231/621 (37%), Positives = 321/621 (51%), Gaps = 119/621 (19%)
Query: 1 MKIWTTFLEPMLGVPSRLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKN 60
MK+W TFLEP+ G+ SR + ED K KN
Sbjct: 630 MKLWITFLEPIFGILSRSQDNLALEDVSKL----------------------------KN 661
Query: 61 TFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSST---LQGNVHINASIPDEVS 117
E ++C + G ++ S T+ GSS+ + N+ + + PD++
Sbjct: 662 NRELQDACLAVKETASGSNRKHPISPKRLSKDNTKMQGSSSREDVSANIKVKTAQPDKL- 720
Query: 118 RVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSE 177
QD + +M++ V QS+ +
Sbjct: 721 ----QD---------DAAMTNEVIQSSKFVS----------------------------- 738
Query: 178 GADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGL 237
N I+ED H + SV K E EEGELSP E+ NF V
Sbjct: 739 ---------PKNDQIMEDEGNHMVNAASVE--KHELEEGELSPTASREQSNFEVNGQNAF 787
Query: 238 EAVHKGKDCNTSQHYQNRREEQICGVAGGEN----DDESDGSPHRSSDDSENASENGDVS 293
+ + K D + ++ +++Q C G +N +D+ + H+ S++++ ASE VS
Sbjct: 788 KPLQKVTD-----NVRSNKDKQSCDKKGAKNKTRAEDDKQENCHKLSENNKTASEML-VS 841
Query: 294 GTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPL 353
GT+ + EE +R + G G AN +G+ S +SERFL T KP+
Sbjct: 842 GTKVSCHEENNRVMN---------CNGRGSVAGEM-ANGNQGEDGSFAFSERFLQTVKPV 891
Query: 354 VKYVSPVFHGKE----ENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASN 409
K++S E + Q+FYGNDS+YVLFRLHQ LYERI++AK +S EKKW+A++
Sbjct: 892 AKHLSWPLQASETCSQNDSQVFYGNDSYYVLFRLHQMLYERIQTAKKHS---EKKWKAAD 948
Query: 410 DTSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAV 469
+T+ D Y RFM++LY+LLDGS DN+KFED+CRAI G QSYVLFTLDKL+ K VK L +V
Sbjct: 949 NTTP-DSYPRFMDALYNLLDGSIDNTKFEDECRAIFGAQSYVLFTLDKLVQKFVKHLHSV 1007
Query: 470 ASGTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIE-CSPTRLSIQLM 528
AS DE D KLLQL+AYE RK G FFD+VYHENA LLH+ NIYRI TRLSIQLM
Sbjct: 1008 AS--DETDTKLLQLHAYENYRKPGKFFDLVYHENACALLHEANIYRIRYVEGTRLSIQLM 1065
Query: 529 DYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQ--V 586
+ G+++ EV V+++P F+ YL N L V D +E G+FL RNK K DE
Sbjct: 1066 NSGNNQLEVMGVAMEPAFADYLQNKCLKSVND-EENHGLFLNRNKKKFTSLDESRGMPVA 1124
Query: 587 MDGIQVINGLECKIACGSSKV 607
M+ + +IN +EC++AC SSKV
Sbjct: 1125 MERLNIINEMECRMACSSSKV 1145
>ref|XP_596697.1| PREDICTED: similar to mSin3A, partial [Bos taurus]
Length = 1198
Score = 102 bits (255), Expect = 3e-20
Identities = 99/375 (26%), Positives = 162/375 (42%), Gaps = 83/375 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G A ++ G G S P S+
Sbjct: 756 AQRGDLSDVE---------EEEEEEMDVD-------EATGAAKKHN--GVGGSPPKSKLL 797
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 798 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 851
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 852 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 911
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E +
Sbjct: 912 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEICVQVTDLYLAENNNGATGGQLNT 969
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRIECSPTRLSIQLMDYGHDKPE--VTAVSIDPNFS 547
+ S S + Y A L+ DEN ++ + +L+I+L+D + + V A
Sbjct: 970 QTSRSLLESTYQRKAEQLMSDENCFKSQ-GQVQLTIELLDTEEENSDDPVEAERWSDYVE 1028
Query: 548 TYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMDGIQV 592
Y+++D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1029 RYMNSDTTS--PELREHLARKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTMENADS 1086
Query: 593 INGLECKIACGSSKV 607
++ LEC+ S K+
Sbjct: 1087 LDKLECRFKLNSYKM 1101
>ref|XP_343396.1| PREDICTED: similar to mSin3A [Rattus norvegicus]
Length = 1274
Score = 101 bits (252), Expect = 6e-20
Identities = 99/379 (26%), Positives = 162/379 (42%), Gaps = 86/379 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 827 AQRGDLSDVE---------EEEEEEMDVD-------EATGAPKKHN--GVGGSPPKSKLL 868
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 869 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 922
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 923 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 982
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E S
Sbjct: 983 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEVCVQVTDLYLAENNNGATGGQLSS 1040
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHDKPE--VTAVSID 543
+ S S + Y A L+ DEN +++ +L+I+L+D + + V A
Sbjct: 1041 QTSRSLLESTYQRKAEQLMSDENCFKLMFIQSQGQVQLTIELLDTEEENSDDPVEAERWS 1100
Query: 544 PNFSTYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMD 588
Y+ +D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1101 DYVERYMSSDTTS--PELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTME 1158
Query: 589 GIQVINGLECKIACGSSKV 607
++ ++ LEC+ S K+
Sbjct: 1159 NVESLDKLECRFKLNSYKM 1177
>ref|NP_056292.1| transcriptional co-repressor Sin3A [Homo sapiens]
gi|37999759|sp|Q96ST3|SIN3A_HUMAN Paired amphipathic
helix protein Sin3a (Transcriptional corepressor Sin3a)
(Histone deacetylase complex subunit Sin3a)
Length = 1273
Score = 101 bits (251), Expect = 8e-20
Identities = 98/379 (25%), Positives = 162/379 (41%), Gaps = 86/379 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 826 AQRGDLSDVE---------EEEEEEMDVD-------EATGAVKKHN--GVGGSPPKSKLL 867
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 868 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 921
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 922 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 981
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E +
Sbjct: 982 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEICVQVTDLYLAENNNGATGGQLNT 1039
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHDKPE--VTAVSID 543
+ S S + Y A L+ DEN +++ +L+I+L+D + + V A
Sbjct: 1040 QNSRSLLESTYQRKAEQLMSDENCFKLMFIQSQGQVQLTIELLDTEEENSDDPVEAERWS 1099
Query: 544 PNFSTYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMD 588
Y+++D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1100 DYVERYMNSDTTS--PELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTME 1157
Query: 589 GIQVINGLECKIACGSSKV 607
+ ++ LEC+ S K+
Sbjct: 1158 NVDSLDKLECRFKLNSYKM 1176
>pir||I61713 co-repressor protein - mouse gi|642619|gb|AAA69773.1| mSin3A gene
product
Length = 1219
Score = 101 bits (251), Expect = 8e-20
Identities = 98/379 (25%), Positives = 162/379 (41%), Gaps = 86/379 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 827 AQRGDLSDVE---------EEEEEEMDVD-------EATGAPKKHN--GVGGSPPKSKLL 868
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 869 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 922
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 923 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 982
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E S
Sbjct: 983 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEVCVQVTDLYLAENNNGATGGQLNS 1040
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHDKPE--VTAVSID 543
+ S S + Y A L+ DEN +++ +L+++L+D + + V A
Sbjct: 1041 QTSRSLLESAYQRKAEQLMSDENCFKLMFIQSQGQVQLTVELLDTEEENSDDPVEAERWS 1100
Query: 544 PNFSTYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMD 588
Y+ +D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1101 DYVERYMSSDTTS--PELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTME 1158
Query: 589 GIQVINGLECKIACGSSKV 607
++ ++ LEC+ S K+
Sbjct: 1159 NVESLDKLECRFKLNSYKM 1177
>gb|AAP97288.1| MSIN3A [Homo sapiens]
Length = 1273
Score = 101 bits (251), Expect = 8e-20
Identities = 98/379 (25%), Positives = 162/379 (41%), Gaps = 86/379 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 826 AQRGDLSDVE---------EEEEEEMDVD-------EATGAVKKHN--GVGGSPPKSKLL 867
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 868 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 921
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 922 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 981
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E +
Sbjct: 982 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEICVQVTDLYLAENNNGATGGQLNT 1039
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHDKPE--VTAVSID 543
+ S S + Y A L+ DEN +++ +L+I+L+D + + V A
Sbjct: 1040 QNSRSLLESTYQRKAEQLMSDENCFKLMFIQSQGQVQLTIELLDTEEENSDDPVEAERWS 1099
Query: 544 PNFSTYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMD 588
Y+++D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1100 DYVERYMNSDTTS--PELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTME 1157
Query: 589 GIQVINGLECKIACGSSKV 607
+ ++ LEC+ S K+
Sbjct: 1158 NVDSLDKLECRFKLNSYKM 1176
>ref|XP_510682.1| PREDICTED: similar to transcriptional co-repressor Sin3A;
transcriptional regulator, SIN3A (yeast) [Pan
troglodytes]
Length = 1296
Score = 101 bits (251), Expect = 8e-20
Identities = 98/379 (25%), Positives = 162/379 (41%), Gaps = 86/379 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 849 AQRGDLSDVE---------EEEEEEMDVD-------EATGAVKKHN--GVGGSPPKSKLL 890
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 891 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 944
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 945 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 1004
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E +
Sbjct: 1005 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEICVQVTDLYLAENNNGATGGQLNT 1062
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHDKPE--VTAVSID 543
+ S S + Y A L+ DEN +++ +L+I+L+D + + V A
Sbjct: 1063 QNSRSLLESTYQRKAEQLMSDENCFKLMFIQSQGQVQLTIELLDTEEENSDDPVEAERWS 1122
Query: 544 PNFSTYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMD 588
Y+++D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1123 DYVERYMNSDTTS--PELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTME 1180
Query: 589 GIQVINGLECKIACGSSKV 607
+ ++ LEC+ S K+
Sbjct: 1181 NVDSLDKLECRFKLNSYKM 1199
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.311 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,108,842,512
Number of Sequences: 2540612
Number of extensions: 48980432
Number of successful extensions: 197018
Number of sequences better than 10.0: 2369
Number of HSP's better than 10.0 without gapping: 686
Number of HSP's successfully gapped in prelim test: 1843
Number of HSP's that attempted gapping in prelim test: 166044
Number of HSP's gapped (non-prelim): 13377
length of query: 637
length of database: 863,360,394
effective HSP length: 134
effective length of query: 503
effective length of database: 522,918,386
effective search space: 263027948158
effective search space used: 263027948158
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC146588.5


