

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146584.2 - phase: 0 /pseudo
(1452 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
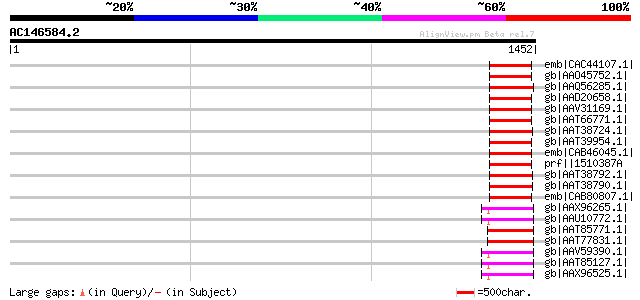
Score E
Sequences producing significant alignments: (bits) Value
emb|CAC44107.1| putative polyprotein [Cicer arietinum] 161 2e-37
gb|AAO45752.1| pol protein [Cucumis melo] 139 9e-31
gb|AAQ56285.1| putative gag-pol protein [Oryza sativa (japonica ... 120 3e-25
gb|AAD20658.1| putative retroelement pol polyprotein [Arabidopsi... 119 6e-25
gb|AAV31169.1| putative polyprotein [Solanum tuberosum] 119 6e-25
gb|AAT66771.1| putative polyprotein [Solanum demissum] 119 8e-25
gb|AAT38724.1| putative retrotransposon protein [Solanum demissum] 119 1e-24
gb|AAT39954.1| putative integrase [Solanum demissum] 117 2e-24
emb|CAB46045.1| retrotransposon like protein [Arabidopsis thalia... 115 8e-24
prf||1510387A retrotransposon del1-46 115 8e-24
gb|AAT38792.1| putative gag-pol polyprotein [Solanum demissum] g... 115 1e-23
gb|AAT38790.1| putative gag-pol polyprotein [Solanum demissum] 115 1e-23
emb|CAB80807.1| putative transposon protein [Arabidopsis thalian... 114 3e-23
gb|AAX96265.1| retrotransposon protein, putative, Ty3-gypsy sub-... 108 2e-21
gb|AAU10772.1| putative polyprotein [Oryza sativa (japonica cult... 105 1e-20
gb|AAT85771.1| hypothetical protein [Oryza sativa (japonica cult... 104 2e-20
gb|AAT77831.1| putative pol polyprotein [Oryza sativa (japonica ... 104 2e-20
gb|AAV59390.1| hypothetical protein [Oryza sativa (japonica cult... 104 3e-20
gb|AAT85127.1| putative polyprotein [Oryza sativa (japonica cult... 103 3e-20
gb|AAX96525.1| retrotransposon protein, putative, Ty3-gypsy sub-... 103 3e-20
>emb|CAC44107.1| putative polyprotein [Cicer arietinum]
Length = 141
Score = 161 bits (407), Expect = 2e-37
Identities = 75/117 (64%), Positives = 95/117 (81%), Gaps = 1/117 (0%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ PKFIGPYQI R+G VAYR+ LPP LSN+H+VFHVSQLRKY+ DPSHVI+ D +Q+
Sbjct: 23 KKLTPKFIGPYQISARIGPVAYRIALPPILSNIHDVFHVSQLRKYLADPSHVIEPDTIQL 82
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
+DNL+ E PVRIDD K+K LR K++ LV+V+W TG++ TWELESKM E +PELF
Sbjct: 83 KDNLSFEVPPVRIDDMKIKRLRTKDVSLVKVIWNPITGDA-TWELESKMREQHPELF 138
>gb|AAO45752.1| pol protein [Cucumis melo]
Length = 923
Score = 139 bits (349), Expect = 9e-31
Identities = 63/116 (54%), Positives = 84/116 (72%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P+F+GP++ILER+G VAYR+ LPP LS +H+VFHVS LRKYVPDPSHV+ + +++
Sbjct: 806 KLSPRFVGPFEILERIGPVAYRLALPPSLSTVHDVFHVSMLRKYVPDPSHVVDYEPLEID 865
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
+NL+ PV + R VKTLR K+IPLV+V+W E TWE E M YPELF
Sbjct: 866 ENLSYVEQPVEVLARGVKTLRNKQIPLVKVLWRNHRVEEATWEREDDMRSRYPELF 921
>gb|AAQ56285.1| putative gag-pol protein [Oryza sativa (japonica cultivar-group)]
Length = 552
Score = 120 bits (301), Expect = 3e-25
Identities = 55/121 (45%), Positives = 80/121 (65%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP+ I RVG++AYR+ LP ++ +H+VFHVS LRKY+ DP H I + + V
Sbjct: 430 KLSPRYIGPFVITARVGSLAYRLQLPESMNGVHDVFHVSMLRKYLRDPEHKIDLEPIMVE 489
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*GK 1446
+LT+E PVRI D + +R + I V+V+WT + TWELE +M + YPELF G
Sbjct: 490 QDLTIECRPVRIFDWSERVMRRRTIKYVKVLWTNQSEREATWELEEEMKKKYPELFVDGM 549
Query: 1447 F 1447
F
Sbjct: 550 F 550
>gb|AAD20658.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25411368|pir||G84493 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1611
Score = 119 bits (299), Expect = 6e-25
Identities = 55/117 (47%), Positives = 79/117 (67%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ P+++GPY+++ERVG VAY++ LPP L+ HNVFHVSQLRK + D ++ +
Sbjct: 1466 KKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSDQEESVEDIPPGL 1525
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++N+TVE PVRI DR K RGK L++V+W E TWE E+KM ++PE F
Sbjct: 1526 KENMTVEAWPVRIMDRMTKGTRGKARDLLKVLWNCRGREEYTWETENKMKANFPEWF 1582
>gb|AAV31169.1| putative polyprotein [Solanum tuberosum]
Length = 394
Score = 119 bits (299), Expect = 6e-25
Identities = 58/116 (50%), Positives = 77/116 (66%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP++IL VG VAY + LPP S +H VFHVS LR+YVPD SHV+Q D V++
Sbjct: 273 KLSPRYIGPFEILRTVGEVAYELALPPVFSAIHPVFHVSMLRRYVPDESHVLQYDAVELD 332
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
D LT PV I R V+ LR + IP+V+V W + E TWE E +M E +P LF
Sbjct: 333 DRLTFVEEPVAILARDVRRLRSRAIPVVKVRWRHCSVEEATWETEQEMREQFPGLF 388
>gb|AAT66771.1| putative polyprotein [Solanum demissum]
Length = 1769
Score = 119 bits (298), Expect = 8e-25
Identities = 58/116 (50%), Positives = 76/116 (65%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP++IL VG VAY + LPP S +H VFHVS LR+YVPD SHV+Q D V++
Sbjct: 1648 KLSPRYIGPFEILRTVGEVAYELALPPVFSAIHPVFHVSMLRRYVPDESHVLQYDAVELD 1707
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
D LT PV I R V+ LR + IP+V+V W E TWE E +M E +P LF
Sbjct: 1708 DRLTFVEEPVAILARDVRRLRSRAIPVVKVRWRHRPVEEATWETEQEMREQFPSLF 1763
>gb|AAT38724.1| putative retrotransposon protein [Solanum demissum]
Length = 1602
Score = 119 bits (297), Expect = 1e-24
Identities = 53/119 (44%), Positives = 84/119 (70%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGPY+I++RVG+VAY + LP L+ +H VFH+S L+K + DPS ++ ++ V+++
Sbjct: 1480 KLSPRYIGPYRIVQRVGSVAYELELPQELAAVHPVFHISMLKKCIGDPSLILPTESVKIK 1539
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*G 1445
DNL+ E +PV+I DR+V+ LR K++ V+V+W E TWE E M + YP LF G
Sbjct: 1540 DNLSYEEVPVQILDRQVRRLRTKDVASVKVLWRNQFVEEATWEAEEDMKKRYPHLFESG 1598
>gb|AAT39954.1| putative integrase [Solanum demissum]
Length = 1609
Score = 117 bits (294), Expect = 2e-24
Identities = 57/116 (49%), Positives = 75/116 (64%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGP++IL VG VAY + LPP S +H VFHV LR+YVPD SHV+Q D V++
Sbjct: 1347 KLSPRYIGPFEILRTVGEVAYELALPPAFSAIHPVFHVPMLRRYVPDESHVLQYDAVELD 1406
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
D LT P+ I R VK LR + IP+V+V W E TWE E +M E +P LF
Sbjct: 1407 DRLTFVEEPIAILARDVKRLRSRAIPVVKVHWRHRPVEEATWETEQEMREQFPVLF 1462
>emb|CAB46045.1| retrotransposon like protein [Arabidopsis thaliana]
gi|7268441|emb|CAB80961.1| retrotransposon like protein
[Arabidopsis thaliana] gi|25407549|pir||F85188
retrotransposon like protein [imported] - Arabidopsis
thaliana
Length = 687
Score = 115 bits (289), Expect = 8e-24
Identities = 52/117 (44%), Positives = 79/117 (67%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ P+++GPY+++ERVG VAY++ LPP L HNVFHVSQLRK + + ++ +
Sbjct: 550 KKLRPRYVGPYKVIERVGAVAYKLDLPPKLDAFHNVFHVSQLRKCLSEQEESMEDVPPGL 609
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++N+TVE PVRI D+ K RGK + L++++W E TWE E+KM ++PE F
Sbjct: 610 KENMTVEAWPVRIMDQMKKGTRGKSMDLLKILWNCGGREEYTWETETKMKANFPEWF 666
>prf||1510387A retrotransposon del1-46
Length = 1443
Score = 115 bits (289), Expect = 8e-24
Identities = 53/117 (45%), Positives = 83/117 (70%), Gaps = 1/117 (0%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++ GP++ILE + VAYR+ LPP LS++HNVFH+S LRKY PDPSH++ +D+++
Sbjct: 1326 KLSPRYTGPFEILEIIWPVAYRLALPPMLSSIHNVFHISMLRKYEPDPSHILDWEDLRLN 1385
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA 1443
+++ E PV++ + K LR K I +V+V+W + E TWELE+ M E +P LF+
Sbjct: 1386 PDISYEEKPVQVLASESKVLRNKIILMVKVLWQHHSEEEATWELEADMQE-FPNLFS 1441
>gb|AAT38792.1| putative gag-pol polyprotein [Solanum demissum]
gi|47825020|gb|AAT38791.1| putative gag-pol polyprotein
[Solanum demissum]
Length = 1351
Score = 115 bits (287), Expect = 1e-23
Identities = 51/119 (42%), Positives = 81/119 (67%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGPY+I +R+G VAY + LP L+ +H VFH+S L+K + DPS ++ ++ +++
Sbjct: 1229 KLSPRYIGPYRIAKRIGNVAYELELPQELAAVHPVFHISMLKKCIGDPSLILPTESIKIN 1288
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*G 1445
DNL+ E +PV+I DR+V+ LR K++ V+V+W E TWE E M + YP LF G
Sbjct: 1289 DNLSYEEVPVQILDRQVRRLRTKDVASVKVLWRDQFVEEATWEAEEDMKKKYPYLFESG 1347
>gb|AAT38790.1| putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 115 bits (287), Expect = 1e-23
Identities = 51/119 (42%), Positives = 81/119 (67%)
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P++IGPY+I +R+G VAY + LP L+ +H VFH+S L+K + DPS ++ ++ +++
Sbjct: 1229 KLSPRYIGPYRIAKRIGNVAYELELPQELAAVHPVFHISMLKKCIGDPSLILPTESIKIN 1288
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELFA*G 1445
DNL+ E +PV+I DR+V+ LR K++ V+V+W E TWE E M + YP LF G
Sbjct: 1289 DNLSYEEVPVQILDRQVRRLRTKDVASVKVLWRDQFVEEATWEAEEDMKKKYPYLFESG 1347
>emb|CAB80807.1| putative transposon protein [Arabidopsis thaliana]
gi|3319363|gb|AAC26251.1| contains similarity to reverse
transcriptase (Pfam: rvt.hmm, score: 33.26) [Arabidopsis
thaliana] gi|7487790|pir||T01862 hypothetical protein
T7M24.4 - Arabidopsis thaliana
Length = 973
Score = 114 bits (284), Expect = 3e-23
Identities = 52/117 (44%), Positives = 78/117 (66%)
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
K++ P+++GPY+++ERVG VAY++ LPP L+ HNVFHVSQLRK + + ++ +
Sbjct: 828 KKLSPRYVGPYKVIERVGAVAYKLDLPPKLNAFHNVFHVSQLRKCLSNQEESVEDVPPGL 887
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++N+TVE PV+I DR K RGK L++V+W E TWE E+KM ++ E F
Sbjct: 888 KENMTVEAWPVQIMDRMTKGTRGKSRDLLKVLWNCGGREQYTWETENKMKANFSEWF 944
>gb|AAX96265.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 1333
Score = 108 bits (269), Expect = 2e-21
Identities = 58/154 (37%), Positives = 89/154 (57%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRT-------CFEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR VF++ Y R CF+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1175 NRRRELVFQAGDYVYLRVTPLRGVHCFQTKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1234
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1235 LGIHNVFHVSQLKKCLRVPEEQASPDQIEIQEDLTYAEKPTRILETSERRTRNKVIRFCK 1294
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1295 VQWSHHSEEEATWEREDELKATHPHLFASSSESR 1328
>gb|AAU10772.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50511449|gb|AAT77372.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1501
Score = 105 bits (261), Expect = 1e-20
Identities = 57/154 (37%), Positives = 89/154 (57%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LPP++
Sbjct: 1343 NRRRELTFEAGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPPNM 1402
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+H+VFHVSQL+K + P S+ + ++++LT P+RI + + R K I +
Sbjct: 1403 VGIHDVFHVSQLKKCLRVPEEQASSEHIDIQEDLTYVEKPIRILETSERRTRNKVIRFCK 1462
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1463 VQWSHHSEEEATWEREDELKAAHPHLFANSSESR 1496
>gb|AAT85771.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 1169
Score = 104 bits (260), Expect = 2e-20
Identities = 52/128 (40%), Positives = 81/128 (62%), Gaps = 1/128 (0%)
Query: 1323 FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSD 1381
F+ K ++ P+F+GP++I+ R G VAY++ LP L N+H+VFHVSQL+K + PS S+
Sbjct: 1037 FQTKGKLAPRFVGPFRIIARRGEVAYQLELPASLGNVHDVFHVSQLKKCLRVPSEQADSE 1096
Query: 1382 DVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPEL 1441
++VR++LT E PV+I D + R + I +V W+ E TWE + ++ +YP+L
Sbjct: 1097 QIEVREDLTYEERPVKILDTMERRTRNRVIRFCKVQWSNHAEEEATWERQDELKAAYPDL 1156
Query: 1442 FA*GKFSR 1449
FA SR
Sbjct: 1157 FASSSESR 1164
>gb|AAT77831.1| putative pol polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 922
Score = 104 bits (260), Expect = 2e-20
Identities = 52/128 (40%), Positives = 81/128 (62%), Gaps = 1/128 (0%)
Query: 1323 FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSD 1381
F+ K ++ P+F+GP++I+ R G VAY++ LP L N+H+VFHVSQL+K + PS S+
Sbjct: 790 FQTKGKLAPRFVGPFRIIARRGEVAYQLELPASLGNVHDVFHVSQLKKCLRVPSEQADSE 849
Query: 1382 DVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPEL 1441
++VR++LT E PV+I D + R + I +V W+ E TWE + ++ +YP+L
Sbjct: 850 QIEVREDLTYEERPVKILDTMERRTRNRVIRFCKVQWSNHAEEEATWERQDELKAAYPDL 909
Query: 1442 FA*GKFSR 1449
FA SR
Sbjct: 910 FASSSESR 917
>gb|AAV59390.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|50933023|ref|XP_476039.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
gi|57900672|gb|AAW57797.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 1253
Score = 104 bits (259), Expect = 3e-20
Identities = 57/154 (37%), Positives = 88/154 (57%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR VF++ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1095 NRRRELVFQAGDYVYLRVTPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1154
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1155 LGIHNVFHVSQLKKCLRVPEEQASPDQIEIQEDLTYVEKPTRILETSERRTRNKVIRFCK 1214
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ + E TWE E ++ ++P LFA SR
Sbjct: 1215 VQWSHLSKEEATWEREDELKAAHPHLFASSSQSR 1248
>gb|AAT85127.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 103 bits (258), Expect = 3e-20
Identities = 58/154 (37%), Positives = 87/154 (55%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1366 NRRRELAFETGDYVYLRVTPLRGVHRFQKKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1425
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P D ++++++LT P RI + + R K I +
Sbjct: 1426 LGIHNVFHVSQLKKCLRVPEEQASPDHIEIQEDLTYVEKPTRILETNERRTRNKVIRFCK 1485
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
V W+ T E TWE E ++ ++P LFA SR
Sbjct: 1486 VQWSHHTEEEATWEREDELKATHPHLFASSSESR 1519
>gb|AAX96525.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 1506
Score = 103 bits (258), Expect = 3e-20
Identities = 56/154 (36%), Positives = 88/154 (56%), Gaps = 8/154 (5%)
Query: 1304 SRRRPRVFESHSYDRCRTC-------FEVK-EVDPKFIGPYQILERVGTVAYRVGLPPHL 1355
+RRR FE+ Y R F+ K ++ P+F+GPY+ILER G VAY++ LP ++
Sbjct: 1348 NRRRELAFETGDYVYLRITPLRGVHRFQTKGKLAPRFVGPYRILERRGEVAYQLELPSNM 1407
Query: 1356 SNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVR 1415
+HNVFHVSQL+K + P + D ++++++LT P RI + + R K I L +
Sbjct: 1408 LGIHNVFHVSQLKKCLRVPEEQVSPDHIEIQEDLTYVEKPTRILETSERRTRNKVIRLCK 1467
Query: 1416 VVWTGATGESLTWELESKMLESYPELFA*GKFSR 1449
+ W+ + E TWE E ++ ++P LF SR
Sbjct: 1468 IQWSHHSEEEATWEREDELKAAHPHLFTSSSESR 1501
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.365 0.164 0.591
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,860,576,722
Number of Sequences: 2540612
Number of extensions: 62583916
Number of successful extensions: 313711
Number of sequences better than 10.0: 583
Number of HSP's better than 10.0 without gapping: 478
Number of HSP's successfully gapped in prelim test: 106
Number of HSP's that attempted gapping in prelim test: 312787
Number of HSP's gapped (non-prelim): 738
length of query: 1452
length of database: 863,360,394
effective HSP length: 141
effective length of query: 1311
effective length of database: 505,134,102
effective search space: 662230807722
effective search space used: 662230807722
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 82 (36.2 bits)
Medicago: description of AC146584.2


