

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146570.9 + phase: 0
(475 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
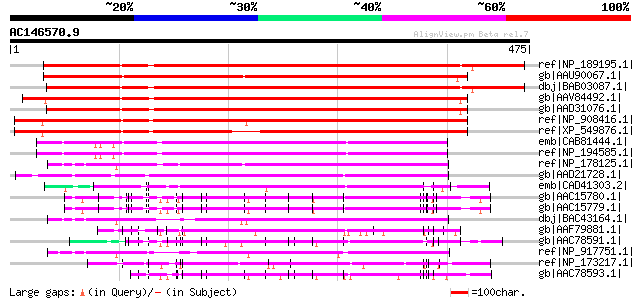
Sequences producing significant alignments: (bits) Value
ref|NP_189195.1| leucine-rich repeat family protein [Arabidopsis... 461 e-128
gb|AAU90067.1| At1g68780 [Arabidopsis thaliana] gi|13605581|gb|A... 461 e-128
dbj|BAB03087.1| unnamed protein product [Arabidopsis thaliana] 458 e-127
gb|AAV84492.1| At1g13230 [Arabidopsis thaliana] gi|42562036|ref|... 456 e-127
gb|AAD31076.1| Contains similarity to gb|U42445 Cf-2.2 from Lyco... 450 e-125
ref|NP_908416.1| P0439B06.12 [Oryza sativa (japonica cultivar-gr... 431 e-119
ref|XP_549876.1| putative disease resistance protein [Oryza sati... 402 e-110
emb|CAB81444.1| putative protein (fragment) [Arabidopsis thalian... 199 2e-49
ref|NP_194585.1| leucine-rich repeat family protein (fragment) [... 199 2e-49
ref|NP_178125.1| leucine-rich repeat family protein [Arabidopsis... 182 2e-44
gb|AAD21728.1| hypothetical protein [Arabidopsis thaliana] gi|66... 182 2e-44
emb|CAD41303.2| OSJNBa0020J04.8 [Oryza sativa (japonica cultivar... 167 9e-40
gb|AAC15780.1| Cf-2.2 [Lycopersicon pimpinellifolium] 165 3e-39
gb|AAC15779.1| Cf-2.1 [Lycopersicon pimpinellifolium] gi|7489083... 165 3e-39
dbj|BAC43164.1| GPI-anchored protein [Arabidopsis thaliana] gi|2... 160 1e-37
gb|AAF79881.1| Contains similarity to receptor protein kinase-li... 157 5e-37
gb|AAC78591.1| disease resistance protein [Lycopersicon esculentum] 155 3e-36
ref|NP_917751.1| P0501G01.13 [Oryza sativa (japonica cultivar-gr... 153 1e-35
ref|NP_173217.1| leucine-rich repeat transmembrane protein kinas... 152 2e-35
gb|AAC78593.1| Hcr2-0B [Lycopersicon esculentum] 152 2e-35
>ref|NP_189195.1| leucine-rich repeat family protein [Arabidopsis thaliana]
Length = 475
Score = 461 bits (1187), Expect = e-128
Identities = 242/444 (54%), Positives = 314/444 (70%), Gaps = 10/444 (2%)
Query: 32 SPMEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPI 91
+PMEK EQEALYS IQGFVG+SWNGSDLYPDPCG T I+GVSCD++ LWYVT + +G +
Sbjct: 31 APMEKTEQEALYSAIQGFVGDSWNGSDLYPDPCGWTPIQGVSCDLYGDLWYVTDLTLGLV 90
Query: 92 HENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEF 151
HENSL CA LE KP+LF+LKHLK+++FFNCF SP + IP +W LA +LES+EF
Sbjct: 91 HENSLSCATS-LEIKPQLFKLKHLKSLTFFNCFTSP----IRIPKEDWINLASNLESLEF 145
Query: 152 RSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
RSNPGLIG +P T G L L+SLV+LENG G +P I NL +LKRLVL+GN F+G IPD
Sbjct: 146 RSNPGLIGELPETIGSLTKLKSLVVLENGFNGKLPTRICNLTRLKRLVLAGNLFTGTIPD 205
Query: 212 IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMD 271
F G DLLILD+SRNS SG LP+++G ++S+LKLDLS+N LEG+L E G LKNLTL+D
Sbjct: 206 CFNGFKDLLILDMSRNSFSGILPLSVGEMVSLLKLDLSNNQLEGRLPQEIGFLKNLTLLD 265
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGG-DIRTLKWENLQNLVILELSNMELIGEI 330
LRNNR+ GL +++++ SL ++VLS NP+G D+ +KWEN+ NLVIL+LS M L GE+
Sbjct: 266 LRNNRISGGLFENIEKIPSLTDLVLSGNPMGSDDMMGIKWENMGNLVILDLSKMGLRGEV 325
Query: 331 PESLSQLKKLRFLGLSDNNITGNL-SPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLG 389
P L+ L++LRFLGL+DNN+TG + S +LETLP L ALY++GNNL GE++FS+ F+ K+G
Sbjct: 326 PLGLTSLRRLRFLGLNDNNLTGTVPSKELETLPCLGALYINGNNLSGELRFSRKFYEKMG 385
Query: 390 RRFGAWSNPKLCYPFELMSTNNVPYGVKPCHQE--EIHLVKSNAKTEVINGDINHNSNFI 447
RF A NP LC S V G+K C E E LV + + D + +S +
Sbjct: 386 TRFKASKNPNLCQDVVSESRQYV-VGLKSCMMEKAEDSLVIKQTWSNLKKEDESSSSMGV 444
Query: 448 TSMGFSSCATSCFWWIFMILGLVL 471
+ W + + L L+L
Sbjct: 445 MVTRHVLLSNGFMWDLLLELSLIL 468
>gb|AAU90067.1| At1g68780 [Arabidopsis thaliana] gi|13605581|gb|AAK32784.1|
At1g68780/F14K14_11 [Arabidopsis thaliana]
gi|18409119|ref|NP_564942.1| leucine-rich repeat family
protein [Arabidopsis thaliana]
gi|12324135|gb|AAG52036.1| unknown protein; 64153-65607
[Arabidopsis thaliana] gi|25404776|pir||E96712 unknown
protein, 64153-65607 [imported] - Arabidopsis thaliana
Length = 432
Score = 461 bits (1186), Expect = e-128
Identities = 230/391 (58%), Positives = 299/391 (75%), Gaps = 5/391 (1%)
Query: 32 SPMEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPI 91
SPMEK EQ ALYSTIQGFVG SWNGS LYPDPCG T I+GV+CDI++ LWYVT ++ G +
Sbjct: 36 SPMEKTEQAALYSTIQGFVGESWNGSYLYPDPCGWTPIQGVTCDIYDELWYVTALSFGTM 95
Query: 92 HENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEF 151
+NSL C+ + +P+LF+LKHLK++S FNCF +PN+ SI W L++SLE +E
Sbjct: 96 KDNSLACSESPV-IRPQLFELKHLKSLSLFNCFTTPNRYLASISDEKWLDLSKSLERLEI 154
Query: 152 RSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
RSNPGLIG +PS L NLQSLV+LEN LTG +P + L +L+RLVLSGN F+G IP+
Sbjct: 155 RSNPGLIGELPSVITNLTNLQSLVVLENKLTGPLPVNLAKLTRLRRLVLSGNRFTGRIPE 214
Query: 212 IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMD 271
++G L+ LLILD+SRN LSG LP+++G L S+LKLDLS+N+LEGKL E +LKNLTL+D
Sbjct: 215 VYG-LTGLLILDVSRNFLSGALPLSVGGLYSLLKLDLSNNYLEGKLPRELESLKNLTLLD 273
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
LRNNRL GL +QEM SL E+VLSNN L GD+ +KW NL+NLV+L+LSN L GEIP
Sbjct: 274 LRNNRLSGGLSKEIQEMTSLVELVLSNNRLAGDLTGIKWRNLKNLVVLDLSNTGLKGEIP 333
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLET-LPSLNALYLSGNNLKGEIQFSKGFFGKLGR 390
S+ +LKKLRFLGLS+NN+ G L P++ET +PSL+ALY++GNN+ GE++FS+ F+ ++GR
Sbjct: 334 GSILELKKLRFLGLSNNNLGGKLIPQMETEMPSLSALYVNGNNISGELEFSRYFYERMGR 393
Query: 391 RFGAWSNPKLCYPFELMS--TNNVPYGVKPC 419
R G W NP LCY + +++VP+GV C
Sbjct: 394 RLGVWGNPNLCYNGDETKNLSDHVPFGVNQC 424
>dbj|BAB03087.1| unnamed protein product [Arabidopsis thaliana]
Length = 443
Score = 458 bits (1179), Expect = e-127
Identities = 241/442 (54%), Positives = 312/442 (70%), Gaps = 10/442 (2%)
Query: 34 MEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHE 93
MEK EQEALYS IQGFVG+SWNGSDLYPDPCG T I+GVSCD++ LWYVT + +G +HE
Sbjct: 1 MEKTEQEALYSAIQGFVGDSWNGSDLYPDPCGWTPIQGVSCDLYGDLWYVTDLTLGLVHE 60
Query: 94 NSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRS 153
NSL CA LE KP+LF+LKHLK+++FFNCF SP + IP +W LA +LES+EFRS
Sbjct: 61 NSLSCATS-LEIKPQLFKLKHLKSLTFFNCFTSP----IRIPKEDWINLASNLESLEFRS 115
Query: 154 NPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF 213
NPGLIG +P T G L L+SLV+LENG G +P I NL +LKRLVL+GN F+G IPD F
Sbjct: 116 NPGLIGELPETIGSLTKLKSLVVLENGFNGKLPTRICNLTRLKRLVLAGNLFTGTIPDCF 175
Query: 214 GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLR 273
G DLLILD+SRNS SG LP+++G ++S+LKLDLS+N LEG+L E G LKNLTL+DLR
Sbjct: 176 NGFKDLLILDMSRNSFSGILPLSVGEMVSLLKLDLSNNQLEGRLPQEIGFLKNLTLLDLR 235
Query: 274 NNRLCCGLVLSLQEMNSLEEMVLSNNPLGG-DIRTLKWENLQNLVILELSNMELIGEIPE 332
NNR+ GL +++++ SL ++VLS NP+G D+ +KWEN+ NLVIL+LS M L GE+P
Sbjct: 236 NNRISGGLFENIEKIPSLTDLVLSGNPMGSDDMMGIKWENMGNLVILDLSKMGLRGEVPL 295
Query: 333 SLSQLKKLRFLGLSDNNITGNL-SPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRR 391
L+ L++LRFLGL+DNN+TG + S +LETLP L ALY++GNNL GE++FS+ F+ K+G R
Sbjct: 296 GLTSLRRLRFLGLNDNNLTGTVPSKELETLPCLGALYINGNNLSGELRFSRKFYEKMGTR 355
Query: 392 FGAWSNPKLCYPFELMSTNNVPYGVKPCHQE--EIHLVKSNAKTEVINGDINHNSNFITS 449
F A NP LC S V G+K C E E LV + + D + +S +
Sbjct: 356 FKASKNPNLCQDVVSESRQYV-VGLKSCMMEKAEDSLVIKQTWSNLKKEDESSSSMGVMV 414
Query: 450 MGFSSCATSCFWWIFMILGLVL 471
+ W + + L L+L
Sbjct: 415 TRHVLLSNGFMWDLLLELSLIL 436
>gb|AAV84492.1| At1g13230 [Arabidopsis thaliana] gi|42562036|ref|NP_172782.2|
leucine-rich repeat family protein [Arabidopsis
thaliana]
Length = 424
Score = 456 bits (1173), Expect = e-127
Identities = 239/418 (57%), Positives = 304/418 (72%), Gaps = 14/418 (3%)
Query: 12 MFILSMNEKCYGQEEDATVV----SPMEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGST 67
+ +LS+ C G E V +PM+K E+EALYS IQGFVG+SWNGS LYPDPCG T
Sbjct: 10 LLLLSLLFGCNGDESLPEVTDSEEAPMDKREREALYSAIQGFVGDSWNGSALYPDPCGWT 69
Query: 68 SIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSP 127
I+GVSCDI+N LWYVT +++G I+ENSLPC++ L+ +PELF+LKHL+++SFFNCF SP
Sbjct: 70 PIQGVSCDIYNDLWYVTDLSLGLIYENSLPCSSS-LQIRPELFELKHLRSLSFFNCFISP 128
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
V W A +LES+EFRSNPGLIG +P T G L L+SLV+LENG +G +P
Sbjct: 129 M---VIAKEELWTNFASNLESLEFRSNPGLIGELPETIGNLTKLKSLVVLENGFSGELPA 185
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD 247
I NL +LKRLV +GN+F+G IP+ F GL +LLILDLSRNS SGTLP + G L+S+LKLD
Sbjct: 186 SICNLKRLKRLVFAGNSFAGMIPNCFKGLKELLILDLSRNSFSGTLPTSFGDLVSLLKLD 245
Query: 248 LSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLG-GDIR 306
LS+N LEG L E G LKNLTL+DLRNNR GL +++ + SL E+VLSNNP+G D+
Sbjct: 246 LSNNLLEGNLPQELGFLKNLTLLDLRNNRFSGGLSKNIENIQSLTELVLSNNPMGEEDMV 305
Query: 307 TLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNL-SPKLETLPSLN 365
W + NLV+L+LS M L GEIP SL+ LK+LRFLGL++NN+TG + S KLE LP L
Sbjct: 306 GTNWGKMSNLVVLDLSKMGLRGEIPTSLTNLKRLRFLGLNNNNLTGFVPSKKLEALPCLG 365
Query: 366 ALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNN----VPYGVKPC 419
ALY++GNNL GE++FS F+ K+GRRF A NP LC P E++ + + P GVKPC
Sbjct: 366 ALYINGNNLTGELRFSTKFYEKMGRRFKASKNPNLCQPLEMVMSESHKHLSPLGVKPC 423
>gb|AAD31076.1| Contains similarity to gb|U42445 Cf-2.2 from Lycopersicon
pimpinellifolium and contains 5 PF|00560 Leucine rich
repeat domains. [Arabidopsis thaliana]
gi|25518745|pir||H86266 hypothetical protein F3F19.26 -
Arabidopsis thaliana
Length = 389
Score = 450 bits (1158), Expect = e-125
Identities = 232/392 (59%), Positives = 293/392 (74%), Gaps = 10/392 (2%)
Query: 34 MEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHE 93
M+K E+EALYS IQGFVG+SWNGS LYPDPCG T I+GVSCDI+N LWYVT +++G I+E
Sbjct: 1 MDKREREALYSAIQGFVGDSWNGSALYPDPCGWTPIQGVSCDIYNDLWYVTDLSLGLIYE 60
Query: 94 NSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRS 153
NSLPC++ L+ +PELF+LKHL+++SFFNCF SP V W A +LES+EFRS
Sbjct: 61 NSLPCSSS-LQIRPELFELKHLRSLSFFNCFISPM---VIAKEELWTNFASNLESLEFRS 116
Query: 154 NPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF 213
NPGLIG +P T G L L+SLV+LENG +G +P I NL +LKRLV +GN+F+G IP+ F
Sbjct: 117 NPGLIGELPETIGNLTKLKSLVVLENGFSGELPASICNLKRLKRLVFAGNSFAGMIPNCF 176
Query: 214 GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLR 273
GL +LLILDLSRNS SGTLP + G L+S+LKLDLS+N LEG L E G LKNLTL+DLR
Sbjct: 177 KGLKELLILDLSRNSFSGTLPTSFGDLVSLLKLDLSNNLLEGNLPQELGFLKNLTLLDLR 236
Query: 274 NNRLCCGLVLSLQEMNSLEEMVLSNNPLG-GDIRTLKWENLQNLVILELSNMELIGEIPE 332
NNR GL +++ + SL E+VLSNNP+G D+ W + NLV+L+LS M L GEIP
Sbjct: 237 NNRFSGGLSKNIENIQSLTELVLSNNPMGEEDMVGTNWGKMSNLVVLDLSKMGLRGEIPT 296
Query: 333 SLSQLKKLRFLGLSDNNITGNL-SPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRR 391
SL+ LK+LRFLGL++NN+TG + S KLE LP L ALY++GNNL GE++FS F+ K+GRR
Sbjct: 297 SLTNLKRLRFLGLNNNNLTGFVPSKKLEALPCLGALYINGNNLTGELRFSTKFYEKMGRR 356
Query: 392 FGAWSNPKLCYPFELMSTNN----VPYGVKPC 419
F A NP LC P E++ + + P GVKPC
Sbjct: 357 FKASKNPNLCQPLEMVMSESHKHLSPLGVKPC 388
>ref|NP_908416.1| P0439B06.12 [Oryza sativa (japonica cultivar-group)]
Length = 477
Score = 431 bits (1109), Expect = e-119
Identities = 218/423 (51%), Positives = 295/423 (69%), Gaps = 11/423 (2%)
Query: 5 ISSVIFVMFILSMNEKCYGQEEDA-----TVVSPMEKAEQEALYSTIQGFVGNSWNGSDL 59
+S ++F++ + S C GQ E A T +PME+ E+ ALY+ I+GFVG WNGS L
Sbjct: 15 LSLLLFIIALHSRLHGCSGQGEAADGSASTAAAPMEEKEKRALYAAIEGFVGKGWNGSAL 74
Query: 60 YPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAIS 119
YPDPCG + I+GVSCD+FNGLWY TV++IGP+ +NSL C+ + +F P+LF LK LK +S
Sbjct: 75 YPDPCGWSPIQGVSCDLFNGLWYPTVMSIGPVLDNSLRCSADA-KFSPQLFDLKRLKTLS 133
Query: 120 FFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLEN 179
F++CF + N P IP +W+KLA SLE++EFR+NPGL G IP++ G L +LQSLV +EN
Sbjct: 134 FYSCFPATN--PTPIPATSWDKLAGSLETLEFRTNPGLTGPIPASLGRLSSLQSLVFVEN 191
Query: 180 GLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGG---LSDLLILDLSRNSLSGTLPVT 236
LTG +P E+G+LV+L+RLVLSGN SG IP G LLI+D+S NSL+G+LP +
Sbjct: 192 NLTGAVPAELGSLVRLRRLVLSGNGLSGQIPASLGNGHFAEQLLIMDVSNNSLTGSLPSS 251
Query: 237 LGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVL 296
LG L +LK+DLS+N L+G L E L +LTL+DLRNN GL LQ M SL++++L
Sbjct: 252 LGGLKGLLKMDLSNNLLQGSLPPELAGLGSLTLLDLRNNSFTGGLPSFLQGMASLQDLLL 311
Query: 297 SNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSP 356
SNNPLGG + L WE L+ L L+LSN+ L+G IPES++ L +LRFL L N +TG++
Sbjct: 312 SNNPLGGSLGQLGWERLRGLATLDLSNLGLVGAIPESMAALTRLRFLALDHNRLTGDVPA 371
Query: 357 KLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNVPYGV 416
+L LP++ ALYL+GNNL G +QFS F+ ++GRRF +W NP LCY + + P GV
Sbjct: 372 RLAELPNIGALYLNGNNLTGTLQFSPAFYQRMGRRFASWDNPGLCYSNAAVDAAHAPPGV 431
Query: 417 KPC 419
C
Sbjct: 432 TVC 434
>ref|XP_549876.1| putative disease resistance protein [Oryza sativa (japonica
cultivar-group)] gi|52075719|dbj|BAD44939.1| putative
disease resistance protein [Oryza sativa (japonica
cultivar-group)]
Length = 450
Score = 402 bits (1032), Expect = e-110
Identities = 206/420 (49%), Positives = 280/420 (66%), Gaps = 32/420 (7%)
Query: 5 ISSVIFVMFILSMNEKCYGQEEDA-----TVVSPMEKAEQEALYSTIQGFVGNSWNGSDL 59
+S ++F++ + S C GQ E A T +PME+ E+ ALY+ I+GFVG WNGS L
Sbjct: 15 LSLLLFIIALHSRLHGCSGQGEAADGSASTAAAPMEEKEKRALYAAIEGFVGKGWNGSAL 74
Query: 60 YPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAIS 119
YPDPCG + I+GVSCD+FNGLWY TV++IGP+ +NSL C+ + +F P+LF LK LK +S
Sbjct: 75 YPDPCGWSPIQGVSCDLFNGLWYPTVMSIGPVLDNSLRCSADA-KFSPQLFDLKRLKTLS 133
Query: 120 FFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLEN 179
F++CF + N P IP +W+KLA SLE++EFR+NPGL G IP++ G L +LQSLV +EN
Sbjct: 134 FYSCFPATN--PTPIPATSWDKLAGSLETLEFRTNPGLTGPIPASLGRLSSLQSLVFVEN 191
Query: 180 GLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGR 239
LTG +P E+G+LV+L+RLVLSGN LSG +P +LG
Sbjct: 192 NLTGAVPAELGSLVRLRRLVLSGNG------------------------LSGQIPASLGG 227
Query: 240 LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNN 299
L +LK+DLS+N L+G L E L +LTL+DLRNN GL LQ M SL++++LSNN
Sbjct: 228 LKGLLKMDLSNNLLQGSLPPELAGLGSLTLLDLRNNSFTGGLPSFLQGMASLQDLLLSNN 287
Query: 300 PLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLE 359
PLGG + L WE L+ L L+LSN+ L+G IPES++ L +LRFL L N +TG++ +L
Sbjct: 288 PLGGSLGQLGWERLRGLATLDLSNLGLVGAIPESMAALTRLRFLALDHNRLTGDVPARLA 347
Query: 360 TLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNVPYGVKPC 419
LP++ ALYL+GNNL G +QFS F+ ++GRRF +W NP LCY + + P GV C
Sbjct: 348 ELPNIGALYLNGNNLTGTLQFSPAFYQRMGRRFASWDNPGLCYSNAAVDAAHAPPGVTVC 407
>emb|CAB81444.1| putative protein (fragment) [Arabidopsis thaliana]
gi|25407606|pir||G85332 hypothetical protein AT4g28560
[imported] - Arabidopsis thaliana gi|7487680|pir||T10650
hypothetical protein T5F17.10 - Arabidopsis thaliana
(fragment)
Length = 449
Score = 199 bits (505), Expect = 2e-49
Identities = 137/390 (35%), Positives = 210/390 (53%), Gaps = 21/390 (5%)
Query: 25 EEDATVVSPMEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIF----NGL 80
+E+A+ ++ +EQEA+Y + V ++ + ++PD ++ +GV CD NG+
Sbjct: 28 DENASPQLALDPSEQEAVYRVLDS-VNSAISWRTIFPDDICASPPDGVVCDNLYASQNGV 86
Query: 81 W---YVTVINIGPIHE--NSLPCANEKLEFKPELFQ-LKHLKAISFFNCFQSPN-KLPVS 133
+VT ++G + + + PC++ P LF KHL+ + F+ CF LP++
Sbjct: 87 ATSVHVTEFHLGYLSDYTQNPPCSSNAT-LDPLLFTAFKHLRKLFFYKCFTDARASLPLT 145
Query: 134 IPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLV 193
+P E LE + F NP L+G I + G L+ LVL NG G+IP +IG+LV
Sbjct: 146 VP----EDFGSVLEELVFIENPSLVGEIGAMIGNFTKLRRLVLTGNGFHGSIPGQIGDLV 201
Query: 194 KLKRLVLSGNNFSGNIP-DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNF 252
L+ + LS N+ +G P + L +L +LD S N ++G P ++G L +LKLDLS N
Sbjct: 202 SLEEITLSRNSLTGGFPANATSRLKNLKVLDFSHNFINGNAPDSIGDLTELLKLDLSFNE 261
Query: 253 LEGKLLNEFGNLKNLTLMDLRNNRL-CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWE 311
G++ + GNLK L +DL NR G+ L L EM+SL E+ LS N LGG I + W+
Sbjct: 262 FTGEVPSGVGNLKKLVFLDLSYNRFGNFGVPLFLAEMSSLREVHLSGNKLGGRIPAI-WK 320
Query: 312 NLQNLVILELSNMELIGEIPESL-SQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLS 370
NL+ + + S M L G IP S+ S LK L FL L +NN+ G + + L S + L
Sbjct: 321 NLEGISGIGFSRMGLEGNIPASMGSSLKNLCFLALDNNNLDGQIPEEFGFLDSAREINLE 380
Query: 371 GNNLKGEIQFSKGFFGKLGRRFGAWSNPKL 400
NNL G+ FS F ++G++ N L
Sbjct: 381 NNNLTGKAPFSDSFRDRIGKKLKLSGNVNL 410
>ref|NP_194585.1| leucine-rich repeat family protein (fragment) [Arabidopsis
thaliana]
Length = 450
Score = 199 bits (505), Expect = 2e-49
Identities = 137/390 (35%), Positives = 210/390 (53%), Gaps = 21/390 (5%)
Query: 25 EEDATVVSPMEKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIF----NGL 80
+E+A+ ++ +EQEA+Y + V ++ + ++PD ++ +GV CD NG+
Sbjct: 28 DENASPQLALDPSEQEAVYRVLDS-VNSAISWRTIFPDDICASPPDGVVCDNLYASQNGV 86
Query: 81 W---YVTVINIGPIHE--NSLPCANEKLEFKPELFQ-LKHLKAISFFNCFQSPN-KLPVS 133
+VT ++G + + + PC++ P LF KHL+ + F+ CF LP++
Sbjct: 87 ATSVHVTEFHLGYLSDYTQNPPCSSNAT-LDPLLFTAFKHLRKLFFYKCFTDARASLPLT 145
Query: 134 IPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLV 193
+P E LE + F NP L+G I + G L+ LVL NG G+IP +IG+LV
Sbjct: 146 VP----EDFGSVLEELVFIENPSLVGEIGAMIGNFTKLRRLVLTGNGFHGSIPGQIGDLV 201
Query: 194 KLKRLVLSGNNFSGNIP-DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNF 252
L+ + LS N+ +G P + L +L +LD S N ++G P ++G L +LKLDLS N
Sbjct: 202 SLEEITLSRNSLTGGFPANATSRLKNLKVLDFSHNFINGNAPDSIGDLTELLKLDLSFNE 261
Query: 253 LEGKLLNEFGNLKNLTLMDLRNNRL-CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWE 311
G++ + GNLK L +DL NR G+ L L EM+SL E+ LS N LGG I + W+
Sbjct: 262 FTGEVPSGVGNLKKLVFLDLSYNRFGNFGVPLFLAEMSSLREVHLSGNKLGGRIPAI-WK 320
Query: 312 NLQNLVILELSNMELIGEIPESL-SQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLS 370
NL+ + + S M L G IP S+ S LK L FL L +NN+ G + + L S + L
Sbjct: 321 NLEGISGIGFSRMGLEGNIPASMGSSLKNLCFLALDNNNLDGQIPEEFGFLDSAREINLE 380
Query: 371 GNNLKGEIQFSKGFFGKLGRRFGAWSNPKL 400
NNL G+ FS F ++G++ N L
Sbjct: 381 NNNLTGKAPFSDSFRDRIGKKLKLSGNVNL 410
>ref|NP_178125.1| leucine-rich repeat family protein [Arabidopsis thaliana]
gi|5902366|gb|AAD55468.1| Hypothetical protein
[Arabidopsis thaliana] gi|22096215|sp|Q9SSD1|TMM_ARATH
TOO MANY MOUTHS protein precursor (TMM)
Length = 496
Score = 182 bits (463), Expect = 2e-44
Identities = 124/372 (33%), Positives = 198/372 (52%), Gaps = 17/372 (4%)
Query: 35 EKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSC-DIFNGLWYVTVINIGPIHE 93
E EQ+A+Y ++ GN W + PD C G+ C + +++V ++ G + +
Sbjct: 55 EPDEQDAVYDIMRA-TGNDWAAA--IPDVCRGRW-HGIECMPDQDNVYHVVSLSFGALSD 110
Query: 94 NSL--PCANEKLEFKPELFQLKHLKAISFFNCF-QSPNKLPVSIPTGNWEKLAESLESIE 150
++ C ++ L +LKHLKA+ F+ C ++P ++P + +L SL+++
Sbjct: 111 DTAFPTCDPQRSYVSESLTRLKHLKALFFYRCLGRAPQRIPAFLG-----RLGSSLQTLV 165
Query: 151 FRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
R N G +G IP G L NL+ L L +N L G+IP L+ L LSGN +G+IP
Sbjct: 166 LREN-GFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGNRLTGSIP 224
Query: 211 DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLM 270
L L +LDL++N L+G +P TL S++K+DLS N + G + L L L+
Sbjct: 225 GFV--LPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTGPIPESINRLNQLVLL 282
Query: 271 DLRNNRLCCGLVLSLQEMNSLEEMVLSNNP-LGGDIRTLKWENLQNLVILELSNMELIGE 329
DL NRL SLQ +NSL+ ++L N I ++ L+NL+IL LSN + G
Sbjct: 283 DLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNLMILVLSNTNIQGS 342
Query: 330 IPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLG 389
IP+SL++L LR L L NN+TG + + + L+ L L+ N+L G + F + ++
Sbjct: 343 IPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNSLTGPVPFERDTVWRMR 402
Query: 390 RRFGAWSNPKLC 401
R+ ++N LC
Sbjct: 403 RKLRLYNNAGLC 414
>gb|AAD21728.1| hypothetical protein [Arabidopsis thaliana]
gi|66792706|gb|AAY56455.1| At2g42800 [Arabidopsis
thaliana] gi|25408813|pir||D84858 hypothetical protein
At2g42800 [imported] - Arabidopsis thaliana
gi|15228004|ref|NP_181808.1| leucine-rich repeat family
protein [Arabidopsis thaliana]
Length = 462
Score = 182 bits (462), Expect = 2e-44
Identities = 132/399 (33%), Positives = 201/399 (50%), Gaps = 16/399 (4%)
Query: 6 SSVIFVMFILSMNEKCYGQEEDATVVSPMEKAEQEALYSTIQGFVGNS-WNGSDLYPDPC 64
SS++F +++ C ++ V + M +E E L+ ++ + W S +P+PC
Sbjct: 11 SSLLFFFLLITPLFLC----QENRVSASMPPSESETLFKIMESMSSDQQWRQS--HPNPC 64
Query: 65 G-STSIEGVSCDIF-NGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAISFFN 122
+S G+ C + L +V+ ++ G P F +F L L+++ FFN
Sbjct: 65 APGSSWPGIECKTGPDHLSHVSRLDFGSAPN---PSCKSSASFPSSIFTLPFLQSVFFFN 121
Query: 123 CFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLT 182
CF P +I SL+ + RSNP L G IP LK+LQ L L +N LT
Sbjct: 122 CF---THFPTTIMFPIKLIPNSSLQQLSLRSNPSLSGQIPPRISSLKSLQILTLSQNRLT 178
Query: 183 GNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLIS 242
G+IP I +L L L LS N +G IP G L++L+ LDLS NSL+GT+P T+ +L
Sbjct: 179 GDIPPAIFSLKSLVHLDLSYNKLTGKIPLQLGNLNNLVGLDLSYNSLTGTIPPTISQLGM 238
Query: 243 VLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLG 302
+ KLDLS N L G++ L++L+ M L NN+L + + SL+ ++ NNP+
Sbjct: 239 LQKLDLSSNSLFGRIPEGVEKLRSLSFMALSNNKLKGAFPKGISNLQSLQYFIMDNNPMF 298
Query: 303 GDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLP 362
+ ++ L L L+L N G IPES ++L L L L++N +TG + E+LP
Sbjct: 299 VAL-PVELGFLPKLQELQLENSGYSGVIPESYTKLTNLSSLSLANNRLTGEIPSGFESLP 357
Query: 363 SLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLC 401
+ L LS N L G + F F +LG+ N LC
Sbjct: 358 HVFHLNLSRNLLIGVVPFDSSFLRRLGKNLDLSGNRGLC 396
>emb|CAD41303.2| OSJNBa0020J04.8 [Oryza sativa (japonica cultivar-group)]
gi|50928147|ref|XP_473601.1| OSJNBa0020J04.8 [Oryza
sativa (japonica cultivar-group)]
Length = 1104
Score = 167 bits (422), Expect = 9e-40
Identities = 125/332 (37%), Positives = 175/332 (52%), Gaps = 20/332 (6%)
Query: 77 FNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPT 136
F+G ++ N+ + S P + ELF L +L + S NKL IP
Sbjct: 375 FSGQIPASLGNLSWLEALSTPGNRLTGDLPSELFVLGNLTFLDL-----SDNKLAGEIPP 429
Query: 137 GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLL-ENGLTGNIPQEIGNLVKL 195
A L+S+ N G IPS G L NL+ L L + L+GN+P E+ L +L
Sbjct: 430 SIGNLAA--LQSLNLSGN-SFSGRIPSNIGNLLNLRVLDLSGQKNLSGNLPAELFGLPQL 486
Query: 196 KRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEG 255
+ + L+GN+FSG++P+ F L L L+LS NS +G++P T G L S+ L SHN + G
Sbjct: 487 QYVSLAGNSFSGDVPEGFSSLWSLRHLNLSVNSFTGSMPATYGYLPSLQVLSASHNRICG 546
Query: 256 KLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQN 315
+L E N NLT++DLR+N+L + + LEE+ LS+N L I + N +
Sbjct: 547 ELPVELANCSNLTVLDLRSNQLTGPIPGDFARLGELEELDLSHNQLSRKIPP-EISNCSS 605
Query: 316 LVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLK 375
LV L+L + L GEIP SLS L KL+ L LS NN+TG++ L +P + +L +S N L
Sbjct: 606 LVTLKLDDNHLGGEIPASLSNLSKLQTLDLSSNNLTGSIPASLAQIPGMLSLNVSQNELS 665
Query: 376 GEIQFSKGFFGKLGRRFGA----WSNPKLCYP 403
GEI LG RFG SNP LC P
Sbjct: 666 GEIP------AMLGSRFGTPSVFASNPNLCGP 691
Score = 126 bits (316), Expect = 2e-27
Identities = 101/313 (32%), Positives = 151/313 (47%), Gaps = 15/313 (4%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL P+ W A L ++ N G +P G L LQ L L N TG +P
Sbjct: 277 NKLAGPFPS--WLAGAGGLTVLDLSGN-AFTGEVPPAVGQLTALQELRLGGNAFTGTVPA 333
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD 247
EIG L+ L L N FSG +P GGL L + L NS SG +P +LG L + L
Sbjct: 334 EIGRCGALQVLDLEDNRFSGEVPAALGGLRRLREVYLGGNSFSGQIPASLGNLSWLEALS 393
Query: 248 LSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT 307
N L G L +E L NLT +DL +N+L + S+ + +L+ + LS N G I +
Sbjct: 394 TPGNRLTGDLPSELFVLGNLTFLDLSDNKLAGEIPPSIGNLAALQSLNLSGNSFSGRIPS 453
Query: 308 LKWENLQNLVILELSNME-LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNA 366
NL NL +L+LS + L G +P L L +L+++ L+ N+ +G++ +L SL
Sbjct: 454 -NIGNLLNLRVLDLSGQKNLSGNLPAELFGLPQLQYVSLAGNSFSGDVPEGFSSLWSLRH 512
Query: 367 LYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNVPYGVKPCHQEEIHL 426
L LS N+ G + + G+ L + + S+ ++C +P + C +
Sbjct: 513 LNLSVNSFTGSMPATYGYLPSL--QVLSASHNRIC--------GELPVELANCSNLTVLD 562
Query: 427 VKSNAKTEVINGD 439
++SN T I GD
Sbjct: 563 LRSNQLTGPIPGD 575
Score = 108 bits (270), Expect = 4e-22
Identities = 84/255 (32%), Positives = 127/255 (48%), Gaps = 5/255 (1%)
Query: 126 SPNKLPVSIPTGNWEKLAES-LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGN 184
S N+L +IP + + S L ++ N ++P + G K+LQ + L N L G
Sbjct: 225 SRNRLTGAIPAAAFGGVGNSSLRIVQVGGNAFSQVDVPVSLG--KDLQVVDLRANKLAGP 282
Query: 185 IPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVL 244
P + L L LSGN F+G +P G L+ L L L N+ +GT+P +GR ++
Sbjct: 283 FPSWLAGAGGLTVLDLSGNAFTGEVPPAVGQLTALQELRLGGNAFTGTVPAEIGRCGALQ 342
Query: 245 KLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGD 304
LDL N G++ G L+ L + L N + SL ++ LE + N L GD
Sbjct: 343 VLDLEDNRFSGEVPAALGGLRRLREVYLGGNSFSGQIPASLGNLSWLEALSTPGNRLTGD 402
Query: 305 IRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSL 364
+ + + L NL L+LS+ +L GEIP S+ L L+ L LS N+ +G + + L +L
Sbjct: 403 LPSELFV-LGNLTFLDLSDNKLAGEIPPSIGNLAALQSLNLSGNSFSGRIPSNIGNLLNL 461
Query: 365 NALYLSG-NNLKGEI 378
L LSG NL G +
Sbjct: 462 RVLDLSGQKNLSGNL 476
Score = 95.5 bits (236), Expect = 3e-18
Identities = 82/291 (28%), Positives = 126/291 (43%), Gaps = 33/291 (11%)
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S N +IP N A SL+ + N L G +P++ G L++L L L N L G I
Sbjct: 128 SSNAFSGTIPA-NVSASATSLQFLNLAVNR-LRGTVPASLGTLQDLHYLWLDGNLLEGTI 185
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTL------------ 233
P + N L L L GN G +P + L IL +SRN L+G +
Sbjct: 186 PSALSNCSALLHLSLQGNALRGILPPAVAAIPSLQILSVSRNRLTGAIPAAAFGGVGNSS 245
Query: 234 ----------------PVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
PV+LG+ + V +DL N L G + LT++DL N
Sbjct: 246 LRIVQVGGNAFSQVDVPVSLGKDLQV--VDLRANKLAGPFPSWLAGAGGLTVLDLSGNAF 303
Query: 278 CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQL 337
+ ++ ++ +L+E+ L N G + + L +L+L + GE+P +L L
Sbjct: 304 TGEVPPAVGQLTALQELRLGGNAFTGTV-PAEIGRCGALQVLDLEDNRFSGEVPAALGGL 362
Query: 338 KKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
++LR + L N+ +G + L L L AL GN L G++ G L
Sbjct: 363 RRLREVYLGGNSFSGQIPASLGNLSWLEALSTPGNRLTGDLPSELFVLGNL 413
Score = 79.3 bits (194), Expect = 2e-13
Identities = 102/364 (28%), Positives = 137/364 (37%), Gaps = 61/364 (16%)
Query: 33 PMEKAEQEALYSTIQGF-----VGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVIN 87
P KAE +AL G + WN S P S GV+C
Sbjct: 31 PEVKAEIDALLMFRSGLRDPYAAMSGWNASS----PSAPCSWRGVAC----------AAG 76
Query: 88 IGPIHENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLE 147
G + E +LP P L L + N P +PVS P SL+
Sbjct: 77 TGRVVELALPKLRLSGAISPALSSLTFDVS---GNLLSGP--VPVSFPP--------SLK 123
Query: 148 SIEFRSNPGLIGNIPSTFGV-LKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFS 206
+E SN G IP+ +LQ L L N L G +P +G L L L L GN
Sbjct: 124 YLELSSN-AFSGTIPANVSASATSLQFLNLAVNRLRGTVPASLGTLQDLHYLWLDGNLLE 182
Query: 207 GNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKL-LNEFGNLK 265
G IP S LL L L N+L G LP + + S+ L +S N L G + FG +
Sbjct: 183 GTIPSALSNCSALLHLSLQGNALRGILPPAVAAIPSLQILSVSRNRLTGAIPAAAFGGVG 242
Query: 266 NLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGG-DIRTLKWENLQNLVILELSNM 324
N +SL + + N D+ ++LQ +++L
Sbjct: 243 N----------------------SSLRIVQVGGNAFSQVDVPVSLGKDLQ---VVDLRAN 277
Query: 325 ELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGF 384
+L G P L+ L L LS N TG + P + L +L L L GN G + G
Sbjct: 278 KLAGPFPSWLAGAGGLTVLDLSGNAFTGEVPPAVGQLTALQELRLGGNAFTGTVPAEIGR 337
Query: 385 FGKL 388
G L
Sbjct: 338 CGAL 341
>gb|AAC15780.1| Cf-2.2 [Lycopersicon pimpinellifolium]
Length = 1112
Score = 165 bits (418), Expect = 3e-39
Identities = 123/343 (35%), Positives = 175/343 (50%), Gaps = 18/343 (5%)
Query: 51 GNSWNGSDLYPDPCG---STSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKP 107
GN +GS P+ G S ++ G+S + NG ++ N+ + +L
Sbjct: 296 GNQLSGS--IPEEIGYLRSLNVLGLSENALNGSIPASLGNLKNLSRLNLVNNQLSGSIPA 353
Query: 108 ELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTF 165
L L +L + +N N+L SIP GN L S+ + N L G+IP++
Sbjct: 354 SLGNLNNLSMLYLYN-----NQLSGSIPASLGNLNNL-----SMLYLYNNQLSGSIPASL 403
Query: 166 GVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLS 225
G L NL L L N L+G+IP+EIG L L L LS N+ +G IP FG +S+L L L
Sbjct: 404 GNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFGNMSNLAFLFLY 463
Query: 226 RNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL 285
N L+ ++P +G L S+ LDLS N L G + FGNL NL+ ++L NN+L + +
Sbjct: 464 ENQLASSVPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVNNQLSGSIPEEI 523
Query: 286 QEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGL 345
+ SL + LS N L G I + NL NL L L N +L G IPE + L+ L LGL
Sbjct: 524 GYLRSLNVLDLSENALNGSI-PASFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNDLGL 582
Query: 346 SDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
S+N + G++ L L +L+ LYL N L G I G+ L
Sbjct: 583 SENALNGSIPASLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSL 625
Score = 155 bits (393), Expect = 2e-36
Identities = 107/275 (38%), Positives = 152/275 (54%), Gaps = 13/275 (4%)
Query: 109 LFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLA--ESLESIEFRSNPGLIGNIPSTFG 166
L L +L + +N N+L SIP E++ SL ++ SN + G IP++FG
Sbjct: 403 LGNLNNLSRLYLYN-----NQLSGSIP----EEIGYLSSLTYLDL-SNNSINGFIPASFG 452
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
+ NL L L EN L ++P+EIG L L L LS N +G+IP FG L++L L+L
Sbjct: 453 NMSNLAFLFLYENQLASSVPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVN 512
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N LSG++P +G L S+ LDLS N L G + FGNL NL+ ++L NN+L + +
Sbjct: 513 NQLSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVNNQLSGSIPEEIG 572
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
+ SL ++ LS N L G I NL NL +L L N +L G IPE + L L +L L
Sbjct: 573 YLRSLNDLGLSENALNGSI-PASLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSLTYLSLG 631
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFS 381
+N++ G + + +L AL L+ NNL GEI S
Sbjct: 632 NNSLNGLIPASFGNMRNLQALILNDNNLIGEIPSS 666
Score = 150 bits (378), Expect = 1e-34
Identities = 101/265 (38%), Positives = 139/265 (52%), Gaps = 8/265 (3%)
Query: 126 SPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTG 183
S N L SIP GN L S F L G+IP L++L L L EN L G
Sbjct: 223 SDNALNGSIPASLGNMNNL-----SFLFLYGNQLSGSIPEEICYLRSLTYLDLSENALNG 277
Query: 184 NIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISV 243
+IP +GNL L L L GN SG+IP+ G L L +L LS N+L+G++P +LG L ++
Sbjct: 278 SIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLGNLKNL 337
Query: 244 LKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGG 303
+L+L +N L G + GNL NL+++ L NN+L + SL +N+L + L NN L G
Sbjct: 338 SRLNLVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYNNQLSG 397
Query: 304 DIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPS 363
I NL NL L L N +L G IPE + L L +L LS+N+I G + + +
Sbjct: 398 SI-PASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFGNMSN 456
Query: 364 LNALYLSGNNLKGEIQFSKGFFGKL 388
L L+L N L + G+ L
Sbjct: 457 LAFLFLYENQLASSVPEEIGYLRSL 481
Score = 145 bits (366), Expect = 3e-33
Identities = 116/351 (33%), Positives = 166/351 (47%), Gaps = 35/351 (9%)
Query: 108 ELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGV 167
E+ L+ L +S F S +P S+ GN L S + N L G+IP
Sbjct: 162 EIGYLRSLTKLSLGINFLS-GSIPASV--GNLNNL-----SFLYLYNNQLSGSIPEEISY 213
Query: 168 LKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRN 227
L++L L L +N L G+IP +GN+ L L L GN SG+IP+ L L LDLS N
Sbjct: 214 LRSLTELDLSDNALNGSIPASLGNMNNLSFLFLYGNQLSGSIPEEICYLRSLTYLDLSEN 273
Query: 228 SLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQE 287
+L+G++P +LG L ++ L L N L G + E G L++L ++ L N L + SL
Sbjct: 274 ALNGSIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLGN 333
Query: 288 MNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSD 347
+ +L + L NN L G I NL NL +L L N +L G IP SL L L L L +
Sbjct: 334 LKNLSRLNLVNNQLSGSI-PASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYN 392
Query: 348 NNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFF--------------GKLGRRFG 393
N ++G++ L L +L+ LYL N L G I G+ G + FG
Sbjct: 393 NQLSGSIPASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFG 452
Query: 394 AWSNPKLCYPFELMSTNNVPYGVKPCHQEEIHLVKS----NAKTEVINGDI 440
SN + +E ++VP EEI ++S + +NG I
Sbjct: 453 NMSNLAFLFLYENQLASSVP--------EEIGYLRSLNVLDLSENALNGSI 495
Score = 144 bits (362), Expect = 8e-33
Identities = 105/281 (37%), Positives = 150/281 (53%), Gaps = 10/281 (3%)
Query: 112 LKHLKAISFFNCFQSPNKLPVSIPTGNWEKLA--ESLESIEFRSNPGLIGNIPSTFGVLK 169
L +L +SF F N+L SIP E++ SL + N L G+IP++ G LK
Sbjct: 283 LGNLNNLSFL--FLYGNQLSGSIP----EEIGYLRSLNVLGLSEN-ALNGSIPASLGNLK 335
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
NL L L+ N L+G+IP +GNL L L L N SG+IP G L++L +L L N L
Sbjct: 336 NLSRLNLVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYNNQL 395
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMN 289
SG++P +LG L ++ +L L +N L G + E G L +LT +DL NN + + S M+
Sbjct: 396 SGSIPASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFGNMS 455
Query: 290 SLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNN 349
+L + L N L + + L++L +L+LS L G IP S L L L L +N
Sbjct: 456 NLAFLFLYENQLASSVPE-EIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVNNQ 514
Query: 350 ITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGR 390
++G++ ++ L SLN L LS N L G I S G L R
Sbjct: 515 LSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSR 555
Score = 143 bits (360), Expect = 1e-32
Identities = 90/237 (37%), Positives = 129/237 (53%), Gaps = 2/237 (0%)
Query: 153 SNPGLIGNIPS-TFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
+N +IG + + F L +L++L L +N + G IP EIGNL L L L+ N SG IP
Sbjct: 78 TNASVIGTLYAFPFSSLPSLENLDLSKNNIYGTIPPEIGNLTNLVYLDLNNNQISGTIPP 137
Query: 212 IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMD 271
G L+ L I+ + N L+G +P +G L S+ KL L NFL G + GNL NL+ +
Sbjct: 138 QIGLLAKLQIIRIFHNQLNGFIPKEIGYLRSLTKLSLGINFLSGSIPASVGNLNNLSFLY 197
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
L NN+L + + + SL E+ LS+N L G I N+ NL L L +L G IP
Sbjct: 198 LYNNQLSGSIPEEISYLRSLTELDLSDNALNGSI-PASLGNMNNLSFLFLYGNQLSGSIP 256
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
E + L+ L +L LS+N + G++ L L +L+ L+L GN L G I G+ L
Sbjct: 257 EEICYLRSLTYLDLSENALNGSIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSL 313
Score = 139 bits (351), Expect = 1e-31
Identities = 102/300 (34%), Positives = 156/300 (52%), Gaps = 20/300 (6%)
Query: 82 YVTVINIGPIHENSLPCANEKLEFKPELF-QLKHLKAISFFNCFQSPNKLPVSIPTGNWE 140
Y+ +N+ + EN+L N + P F L +L ++ N N+L SIP E
Sbjct: 477 YLRSLNVLDLSENAL---NGSI---PASFGNLNNLSRLNLVN-----NQLSGSIP----E 521
Query: 141 KLA--ESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRL 198
++ SL ++ N L G+IP++FG L NL L L+ N L+G+IP+EIG L L L
Sbjct: 522 EIGYLRSLNVLDLSEN-ALNGSIPASFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNDL 580
Query: 199 VLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLL 258
LS N +G+IP G L++L +L L N LSG++P +G L S+ L L +N L G +
Sbjct: 581 GLSENALNGSIPASLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSLTYLSLGNNSLNGLIP 640
Query: 259 NEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVI 318
FGN++NL + L +N L + S+ + SLE + + N L G + N+ NL +
Sbjct: 641 ASFGNMRNLQALILNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGKVPQC-LGNISNLQV 699
Query: 319 LELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L +S+ GE+P S+S L L+ L NN+ G + + SL + N L G +
Sbjct: 700 LSMSSNSFSGELPSSISNLTSLQILDFGRNNLEGAIPQCFGNISSLEVFDMQNNKLSGTL 759
Score = 136 bits (342), Expect = 2e-30
Identities = 97/276 (35%), Positives = 139/276 (50%), Gaps = 37/276 (13%)
Query: 109 LFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRS--NPGLIGNIPSTFG 166
L L +L + +N N+L SIP E++ L S+ + S N L G IP++FG
Sbjct: 595 LGNLNNLSMLYLYN-----NQLSGSIP----EEIGY-LSSLTYLSLGNNSLNGLIPASFG 644
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
++NLQ+L+L +N L G IP + NL L+ L + NN G +P G +S+L +L +S
Sbjct: 645 NMRNLQALILNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGKVPQCLGNISNLQVLSMSS 704
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
NS SG LP ++ L S+ LD N LEG + FGN+ +L + D++NN+L L
Sbjct: 705 NSFSGELPSSISNLTSLQILDFGRNNLEGAIPQCFGNISSLEVFDMQNNKLSGTLP---- 760
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
+N +G +L+ L L EL EIP SL KKL+ L L
Sbjct: 761 ----------TNFSIG-----------CSLISLNLHGNELEDEIPRSLDNCKKLQVLDLG 799
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSK 382
DN + L TLP L L L+ N L G I+ S+
Sbjct: 800 DNQLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSR 835
Score = 70.9 bits (172), Expect = 8e-11
Identities = 80/265 (30%), Positives = 115/265 (43%), Gaps = 43/265 (16%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL ++PT + SL S+ N L IP + K LQ L L +N L P
Sbjct: 753 NKLSGTLPTNF--SIGCSLISLNLHGNE-LEDEIPRSLDNCKKLQVLDLGDNQLNDTFPM 809
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLS--DLLILDLSRNSLSGTLPVTLGRLISVLK 245
+G L +L+ L L+ N G I + DL I+DLSRN+ S LP +L
Sbjct: 810 WLGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLSRNAFSQDLPTSL-------- 861
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR-------------LCCGLVLSLQEMNSLE 292
F +LK + +D + GL L + + SL
Sbjct: 862 ---------------FEHLKGMRTVDKTMEEPSYESYYDDSVVVVTKGLELEIVRILSLY 906
Query: 293 EMV-LSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNIT 351
++ LS+N G I ++ + L + IL +S+ L G IP SL L L L LS N ++
Sbjct: 907 TVIDLSSNKFEGHIPSVLGD-LIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLS 965
Query: 352 GNLSPKLETLPSLNALYLSGNNLKG 376
G + +L +L L L LS N L+G
Sbjct: 966 GEIPQQLASLTFLEFLNLSHNYLQG 990
Score = 62.0 bits (149), Expect = 4e-08
Identities = 35/80 (43%), Positives = 47/80 (58%)
Query: 176 LLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPV 235
L N G+IP +G+L+ ++ L +S N G IP G LS L LDLS N LSG +P
Sbjct: 911 LSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEIPQ 970
Query: 236 TLGRLISVLKLDLSHNFLEG 255
L L + L+LSHN+L+G
Sbjct: 971 QLASLTFLEFLNLSHNYLQG 990
Score = 60.8 bits (146), Expect = 9e-08
Identities = 35/98 (35%), Positives = 57/98 (57%), Gaps = 1/98 (1%)
Query: 181 LTGNIPQEIGNLVKLKRLV-LSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGR 239
+T + EI ++ L ++ LS N F G+IP + G L + IL++S N+L G +P +LG
Sbjct: 891 VTKGLELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGS 950
Query: 240 LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L + LDLS N L G++ + +L L ++L +N L
Sbjct: 951 LSILESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYL 988
Score = 55.8 bits (133), Expect = 3e-06
Identities = 34/90 (37%), Positives = 51/90 (55%)
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
LS ++DLS N G +P LG LI++ L++SHN L+G + + G+L L +DL N
Sbjct: 903 LSLYTVIDLSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFN 962
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+L + L + LE + LS+N L G I
Sbjct: 963 QLSGEIPQQLASLTFLEFLNLSHNYLQGCI 992
Score = 54.3 bits (129), Expect = 8e-06
Identities = 32/76 (42%), Positives = 42/76 (55%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G+IPS G L ++ L + N L G IP +G+L L+ L LS N SG IP L+
Sbjct: 918 GHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEIPQQLASLTF 977
Query: 219 LLILDLSRNSLSGTLP 234
L L+LS N L G +P
Sbjct: 978 LEFLNLSHNYLQGCIP 993
Score = 46.6 bits (109), Expect = 0.002
Identities = 35/101 (34%), Positives = 55/101 (53%), Gaps = 4/101 (3%)
Query: 111 QLKHLKAISFFNCFQ-SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLK 169
+L+ ++ +S + S NK IP+ + +A + ++ S+ L G IPS+ G L
Sbjct: 896 ELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIAIRILNV---SHNALQGYIPSSLGSLS 952
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
L+SL L N L+G IPQ++ +L L+ L LS N G IP
Sbjct: 953 ILESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYLQGCIP 993
>gb|AAC15779.1| Cf-2.1 [Lycopersicon pimpinellifolium] gi|7489083|pir||T10504
disease resistance protein Cf-2.1 - currant tomato
gi|1587673|prf||2207203A Cf-2 gene
Length = 1112
Score = 165 bits (418), Expect = 3e-39
Identities = 123/343 (35%), Positives = 175/343 (50%), Gaps = 18/343 (5%)
Query: 51 GNSWNGSDLYPDPCG---STSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKP 107
GN +GS P+ G S ++ G+S + NG ++ N+ + +L
Sbjct: 296 GNQLSGS--IPEEIGYLRSLNVLGLSENALNGSIPASLGNLKNLSRLNLVNNQLSGSIPA 353
Query: 108 ELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTF 165
L L +L + +N N+L SIP GN L S+ + N L G+IP++
Sbjct: 354 SLGNLNNLSMLYLYN-----NQLSGSIPASLGNLNNL-----SMLYLYNNQLSGSIPASL 403
Query: 166 GVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLS 225
G L NL L L N L+G+IP+EIG L L L LS N+ +G IP FG +S+L L L
Sbjct: 404 GNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFGNMSNLAFLFLY 463
Query: 226 RNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL 285
N L+ ++P +G L S+ LDLS N L G + FGNL NL+ ++L NN+L + +
Sbjct: 464 ENQLASSVPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVNNQLSGSIPEEI 523
Query: 286 QEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGL 345
+ SL + LS N L G I + NL NL L L N +L G IPE + L+ L LGL
Sbjct: 524 GYLRSLNVLDLSENALNGSI-PASFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNDLGL 582
Query: 346 SDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
S+N + G++ L L +L+ LYL N L G I G+ L
Sbjct: 583 SENALNGSIPASLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSL 625
Score = 155 bits (393), Expect = 2e-36
Identities = 107/275 (38%), Positives = 152/275 (54%), Gaps = 13/275 (4%)
Query: 109 LFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLA--ESLESIEFRSNPGLIGNIPSTFG 166
L L +L + +N N+L SIP E++ SL ++ SN + G IP++FG
Sbjct: 403 LGNLNNLSRLYLYN-----NQLSGSIP----EEIGYLSSLTYLDL-SNNSINGFIPASFG 452
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
+ NL L L EN L ++P+EIG L L L LS N +G+IP FG L++L L+L
Sbjct: 453 NMSNLAFLFLYENQLASSVPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVN 512
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N LSG++P +G L S+ LDLS N L G + FGNL NL+ ++L NN+L + +
Sbjct: 513 NQLSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVNNQLSGSIPEEIG 572
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
+ SL ++ LS N L G I NL NL +L L N +L G IPE + L L +L L
Sbjct: 573 YLRSLNDLGLSENALNGSI-PASLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSLTYLSLG 631
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFS 381
+N++ G + + +L AL L+ NNL GEI S
Sbjct: 632 NNSLNGLIPASFGNMRNLQALILNDNNLIGEIPSS 666
Score = 150 bits (378), Expect = 1e-34
Identities = 101/265 (38%), Positives = 139/265 (52%), Gaps = 8/265 (3%)
Query: 126 SPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTG 183
S N L SIP GN L S F L G+IP L++L L L EN L G
Sbjct: 223 SDNALNGSIPASLGNMNNL-----SFLFLYGNQLSGSIPEEICYLRSLTYLDLSENALNG 277
Query: 184 NIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISV 243
+IP +GNL L L L GN SG+IP+ G L L +L LS N+L+G++P +LG L ++
Sbjct: 278 SIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLGNLKNL 337
Query: 244 LKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGG 303
+L+L +N L G + GNL NL+++ L NN+L + SL +N+L + L NN L G
Sbjct: 338 SRLNLVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYNNQLSG 397
Query: 304 DIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPS 363
I NL NL L L N +L G IPE + L L +L LS+N+I G + + +
Sbjct: 398 SI-PASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFGNMSN 456
Query: 364 LNALYLSGNNLKGEIQFSKGFFGKL 388
L L+L N L + G+ L
Sbjct: 457 LAFLFLYENQLASSVPEEIGYLRSL 481
Score = 145 bits (366), Expect = 3e-33
Identities = 116/351 (33%), Positives = 166/351 (47%), Gaps = 35/351 (9%)
Query: 108 ELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGV 167
E+ L+ L +S F S +P S+ GN L S + N L G+IP
Sbjct: 162 EIGYLRSLTKLSLGINFLS-GSIPASV--GNLNNL-----SFLYLYNNQLSGSIPEEISY 213
Query: 168 LKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRN 227
L++L L L +N L G+IP +GN+ L L L GN SG+IP+ L L LDLS N
Sbjct: 214 LRSLTELDLSDNALNGSIPASLGNMNNLSFLFLYGNQLSGSIPEEICYLRSLTYLDLSEN 273
Query: 228 SLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQE 287
+L+G++P +LG L ++ L L N L G + E G L++L ++ L N L + SL
Sbjct: 274 ALNGSIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSLNVLGLSENALNGSIPASLGN 333
Query: 288 MNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSD 347
+ +L + L NN L G I NL NL +L L N +L G IP SL L L L L +
Sbjct: 334 LKNLSRLNLVNNQLSGSI-PASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYN 392
Query: 348 NNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFF--------------GKLGRRFG 393
N ++G++ L L +L+ LYL N L G I G+ G + FG
Sbjct: 393 NQLSGSIPASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFG 452
Query: 394 AWSNPKLCYPFELMSTNNVPYGVKPCHQEEIHLVKS----NAKTEVINGDI 440
SN + +E ++VP EEI ++S + +NG I
Sbjct: 453 NMSNLAFLFLYENQLASSVP--------EEIGYLRSLNVLDLSENALNGSI 495
Score = 144 bits (362), Expect = 8e-33
Identities = 105/281 (37%), Positives = 150/281 (53%), Gaps = 10/281 (3%)
Query: 112 LKHLKAISFFNCFQSPNKLPVSIPTGNWEKLA--ESLESIEFRSNPGLIGNIPSTFGVLK 169
L +L +SF F N+L SIP E++ SL + N L G+IP++ G LK
Sbjct: 283 LGNLNNLSFL--FLYGNQLSGSIP----EEIGYLRSLNVLGLSEN-ALNGSIPASLGNLK 335
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
NL L L+ N L+G+IP +GNL L L L N SG+IP G L++L +L L N L
Sbjct: 336 NLSRLNLVNNQLSGSIPASLGNLNNLSMLYLYNNQLSGSIPASLGNLNNLSMLYLYNNQL 395
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMN 289
SG++P +LG L ++ +L L +N L G + E G L +LT +DL NN + + S M+
Sbjct: 396 SGSIPASLGNLNNLSRLYLYNNQLSGSIPEEIGYLSSLTYLDLSNNSINGFIPASFGNMS 455
Query: 290 SLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNN 349
+L + L N L + + L++L +L+LS L G IP S L L L L +N
Sbjct: 456 NLAFLFLYENQLASSVPE-EIGYLRSLNVLDLSENALNGSIPASFGNLNNLSRLNLVNNQ 514
Query: 350 ITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGR 390
++G++ ++ L SLN L LS N L G I S G L R
Sbjct: 515 LSGSIPEEIGYLRSLNVLDLSENALNGSIPASFGNLNNLSR 555
Score = 143 bits (360), Expect = 1e-32
Identities = 90/237 (37%), Positives = 129/237 (53%), Gaps = 2/237 (0%)
Query: 153 SNPGLIGNIPS-TFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
+N +IG + + F L +L++L L +N + G IP EIGNL L L L+ N SG IP
Sbjct: 78 TNASVIGTLYAFPFSSLPSLENLDLSKNNIYGTIPPEIGNLTNLVYLDLNNNQISGTIPP 137
Query: 212 IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMD 271
G L+ L I+ + N L+G +P +G L S+ KL L NFL G + GNL NL+ +
Sbjct: 138 QIGLLAKLQIIRIFHNQLNGFIPKEIGYLRSLTKLSLGINFLSGSIPASVGNLNNLSFLY 197
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
L NN+L + + + SL E+ LS+N L G I N+ NL L L +L G IP
Sbjct: 198 LYNNQLSGSIPEEISYLRSLTELDLSDNALNGSI-PASLGNMNNLSFLFLYGNQLSGSIP 256
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
E + L+ L +L LS+N + G++ L L +L+ L+L GN L G I G+ L
Sbjct: 257 EEICYLRSLTYLDLSENALNGSIPASLGNLNNLSFLFLYGNQLSGSIPEEIGYLRSL 313
Score = 139 bits (351), Expect = 1e-31
Identities = 102/300 (34%), Positives = 156/300 (52%), Gaps = 20/300 (6%)
Query: 82 YVTVINIGPIHENSLPCANEKLEFKPELF-QLKHLKAISFFNCFQSPNKLPVSIPTGNWE 140
Y+ +N+ + EN+L N + P F L +L ++ N N+L SIP E
Sbjct: 477 YLRSLNVLDLSENAL---NGSI---PASFGNLNNLSRLNLVN-----NQLSGSIP----E 521
Query: 141 KLA--ESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRL 198
++ SL ++ N L G+IP++FG L NL L L+ N L+G+IP+EIG L L L
Sbjct: 522 EIGYLRSLNVLDLSEN-ALNGSIPASFGNLNNLSRLNLVNNQLSGSIPEEIGYLRSLNDL 580
Query: 199 VLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLL 258
LS N +G+IP G L++L +L L N LSG++P +G L S+ L L +N L G +
Sbjct: 581 GLSENALNGSIPASLGNLNNLSMLYLYNNQLSGSIPEEIGYLSSLTYLSLGNNSLNGLIP 640
Query: 259 NEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVI 318
FGN++NL + L +N L + S+ + SLE + + N L G + N+ NL +
Sbjct: 641 ASFGNMRNLQALILNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGKVPQC-LGNISNLQV 699
Query: 319 LELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L +S+ GE+P S+S L L+ L NN+ G + + SL + N L G +
Sbjct: 700 LSMSSNSFSGELPSSISNLTSLQILDFGRNNLEGAIPQCFGNISSLEVFDMQNNKLSGTL 759
Score = 136 bits (342), Expect = 2e-30
Identities = 97/276 (35%), Positives = 139/276 (50%), Gaps = 37/276 (13%)
Query: 109 LFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRS--NPGLIGNIPSTFG 166
L L +L + +N N+L SIP E++ L S+ + S N L G IP++FG
Sbjct: 595 LGNLNNLSMLYLYN-----NQLSGSIP----EEIGY-LSSLTYLSLGNNSLNGLIPASFG 644
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
++NLQ+L+L +N L G IP + NL L+ L + NN G +P G +S+L +L +S
Sbjct: 645 NMRNLQALILNDNNLIGEIPSSVCNLTSLEVLYMPRNNLKGKVPQCLGNISNLQVLSMSS 704
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
NS SG LP ++ L S+ LD N LEG + FGN+ +L + D++NN+L L
Sbjct: 705 NSFSGELPSSISNLTSLQILDFGRNNLEGAIPQCFGNISSLEVFDMQNNKLSGTLP---- 760
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
+N +G +L+ L L EL EIP SL KKL+ L L
Sbjct: 761 ----------TNFSIG-----------CSLISLNLHGNELEDEIPRSLDNCKKLQVLDLG 799
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSK 382
DN + L TLP L L L+ N L G I+ S+
Sbjct: 800 DNQLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSR 835
Score = 70.9 bits (172), Expect = 8e-11
Identities = 80/265 (30%), Positives = 115/265 (43%), Gaps = 43/265 (16%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL ++PT + SL S+ N L IP + K LQ L L +N L P
Sbjct: 753 NKLSGTLPTNF--SIGCSLISLNLHGNE-LEDEIPRSLDNCKKLQVLDLGDNQLNDTFPM 809
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLS--DLLILDLSRNSLSGTLPVTLGRLISVLK 245
+G L +L+ L L+ N G I + DL I+DLSRN+ S LP +L
Sbjct: 810 WLGTLPELRVLRLTSNKLHGPIRSSRAEIMFPDLRIIDLSRNAFSQDLPTSL-------- 861
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR-------------LCCGLVLSLQEMNSLE 292
F +LK + +D + GL L + + SL
Sbjct: 862 ---------------FEHLKGMRTVDKTMEEPSYESYYDDSVVVVTKGLELEIVRILSLY 906
Query: 293 EMV-LSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNIT 351
++ LS+N G I ++ + L + IL +S+ L G IP SL L L L LS N ++
Sbjct: 907 TVIDLSSNKFEGHIPSVLGD-LIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLS 965
Query: 352 GNLSPKLETLPSLNALYLSGNNLKG 376
G + +L +L L L LS N L+G
Sbjct: 966 GEIPQQLASLTFLEFLNLSHNYLQG 990
Score = 62.0 bits (149), Expect = 4e-08
Identities = 35/80 (43%), Positives = 47/80 (58%)
Query: 176 LLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPV 235
L N G+IP +G+L+ ++ L +S N G IP G LS L LDLS N LSG +P
Sbjct: 911 LSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEIPQ 970
Query: 236 TLGRLISVLKLDLSHNFLEG 255
L L + L+LSHN+L+G
Sbjct: 971 QLASLTFLEFLNLSHNYLQG 990
Score = 60.8 bits (146), Expect = 9e-08
Identities = 35/98 (35%), Positives = 57/98 (57%), Gaps = 1/98 (1%)
Query: 181 LTGNIPQEIGNLVKLKRLV-LSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGR 239
+T + EI ++ L ++ LS N F G+IP + G L + IL++S N+L G +P +LG
Sbjct: 891 VTKGLELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGS 950
Query: 240 LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L + LDLS N L G++ + +L L ++L +N L
Sbjct: 951 LSILESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYL 988
Score = 55.8 bits (133), Expect = 3e-06
Identities = 34/90 (37%), Positives = 51/90 (55%)
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
LS ++DLS N G +P LG LI++ L++SHN L+G + + G+L L +DL N
Sbjct: 903 LSLYTVIDLSSNKFEGHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFN 962
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+L + L + LE + LS+N L G I
Sbjct: 963 QLSGEIPQQLASLTFLEFLNLSHNYLQGCI 992
Score = 54.3 bits (129), Expect = 8e-06
Identities = 32/76 (42%), Positives = 42/76 (55%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G+IPS G L ++ L + N L G IP +G+L L+ L LS N SG IP L+
Sbjct: 918 GHIPSVLGDLIAIRILNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEIPQQLASLTF 977
Query: 219 LLILDLSRNSLSGTLP 234
L L+LS N L G +P
Sbjct: 978 LEFLNLSHNYLQGCIP 993
Score = 46.6 bits (109), Expect = 0.002
Identities = 35/101 (34%), Positives = 55/101 (53%), Gaps = 4/101 (3%)
Query: 111 QLKHLKAISFFNCFQ-SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLK 169
+L+ ++ +S + S NK IP+ + +A + ++ S+ L G IPS+ G L
Sbjct: 896 ELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIAIRILNV---SHNALQGYIPSSLGSLS 952
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
L+SL L N L+G IPQ++ +L L+ L LS N G IP
Sbjct: 953 ILESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYLQGCIP 993
>dbj|BAC43164.1| GPI-anchored protein [Arabidopsis thaliana]
gi|29028896|gb|AAO64827.1| At1g80080 [Arabidopsis
thaliana]
Length = 496
Score = 160 bits (404), Expect = 1e-37
Identities = 120/394 (30%), Positives = 193/394 (48%), Gaps = 61/394 (15%)
Query: 35 EKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSC-DIFNGLWYVTVINIGPIHE 93
E EQ+A+Y ++ GN W + PD C G+ C + +++V ++ G + +
Sbjct: 55 EPDEQDAVYDIMRA-TGNDWAAA--IPDVCRGRW-HGIECMPDQDNVYHVVSLSFGALSD 110
Query: 94 NSL--PCANEKLEFKPELFQLKHLKAISFFNCF-QSPNKLPVSIPTGNWEKLAESLESIE 150
++ C ++ L +LKHLKA+ F+ C ++P ++P +
Sbjct: 111 DTAFPTCDPQRSYVSESLTRLKHLKALFFYRCLGRAPQRIPAFL---------------- 154
Query: 151 FRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
G +G+ +LQ+LVL ENG G IP E+GNL LK L L N+ +G+IP
Sbjct: 155 -----GRLGS---------SLQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIP 200
Query: 211 ------------DIFGG----------LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDL 248
D+ G L L +LDL++N L+G +P TL S++K+DL
Sbjct: 201 LSFNRFSGLRSLDLSGNRLTGSIPGFVLPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDL 260
Query: 249 SHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVL-SNNPLGGDIRT 307
S N + G + L L+DL NRL SLQ +NSL+ ++L N I
Sbjct: 261 SRNRVTGPIPESINRFNQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPE 320
Query: 308 LKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNAL 367
++ L+NL+IL LSN + G IP+SL++L LR L L NN+TG + + + L+ L
Sbjct: 321 NAFKGLKNLMILVLSNTNIQGSIPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSEL 380
Query: 368 YLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLC 401
L+ ++L G + F + ++ R+ ++N LC
Sbjct: 381 RLNDSSLTGPVPFERDTVWRMRRKLRLYNNAGLC 414
>gb|AAF79881.1| Contains similarity to receptor protein kinase-like protein from
Arabidopsis thaliana gb|AL161513. It contains a
eukaryotic protein kinase domain PF|00069. EST
gb|AI997574 comes from this gene
gi|15219699|ref|NP_174809.1| leucine-rich repeat
transmembrane protein kinase, putative [Arabidopsis
thaliana] gi|25518391|pir||B86479 hypothetical protein
F14D7.1 - Arabidopsis thaliana
Length = 1120
Score = 157 bits (398), Expect = 5e-37
Identities = 118/311 (37%), Positives = 156/311 (49%), Gaps = 37/311 (11%)
Query: 104 EFKPELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNI 161
E P L LK+L + + N L IP+ GN E + + S L G+I
Sbjct: 141 EISPSLGNLKNLTVL-----YLHQNYLTSVIPSELGNMESMTDLA-----LSQNKLTGSI 190
Query: 162 PSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLI 221
PS+ G LKNL L L EN LTG IP E+GN+ + L LS N +G+IP G L +L++
Sbjct: 191 PSSLGNLKNLMVLYLYENYLTGVIPPELGNMESMTDLALSQNKLTGSIPSTLGNLKNLMV 250
Query: 222 LDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGL 281
L L N L+G +P +G + S+ L LS N L G + + GNLKNLTL+ L N L G+
Sbjct: 251 LYLYENYLTGVIPPEIGNMESMTNLALSQNKLTGSIPSSLGNLKNLTLLSLFQNYLTGGI 310
Query: 282 VLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVIL-------------ELSNME--- 325
L + S+ ++ LSNN L G I + NL+NL IL EL NME
Sbjct: 311 PPKLGNIESMIDLELSNNKLTGSIPS-SLGNLKNLTILYLYENYLTGVIPPELGNMESMI 369
Query: 326 --------LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGE 377
L G IP S LK L +L L N +TG + +L + S+ L LS N L G
Sbjct: 370 DLQLNNNKLTGSIPSSFGNLKNLTYLYLYLNYLTGVIPQELGNMESMINLDLSQNKLTGS 429
Query: 378 IQFSKGFFGKL 388
+ S G F KL
Sbjct: 430 VPDSFGNFTKL 440
Score = 156 bits (394), Expect = 2e-36
Identities = 96/235 (40%), Positives = 134/235 (56%), Gaps = 2/235 (0%)
Query: 144 ESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGN 203
ES+ + N L G+IPST G LKNL L L EN LTG IP EIGN+ + L LS N
Sbjct: 222 ESMTDLALSQNK-LTGSIPSTLGNLKNLMVLYLYENYLTGVIPPEIGNMESMTNLALSQN 280
Query: 204 NFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGN 263
+G+IP G L +L +L L +N L+G +P LG + S++ L+LS+N L G + + GN
Sbjct: 281 KLTGSIPSSLGNLKNLTLLSLFQNYLTGGIPPKLGNIESMIDLELSNNKLTGSIPSSLGN 340
Query: 264 LKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSN 323
LKNLT++ L N L + L M S+ ++ L+NN L G I + + NL+NL L L
Sbjct: 341 LKNLTILYLYENYLTGVIPPELGNMESMIDLQLNNNKLTGSIPS-SFGNLKNLTYLYLYL 399
Query: 324 MELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L G IP+ L ++ + L LS N +TG++ L +LYL N+L G I
Sbjct: 400 NYLTGVIPQELGNMESMINLDLSQNKLTGSVPDSFGNFTKLESLYLRVNHLSGAI 454
Score = 142 bits (357), Expect = 3e-32
Identities = 115/330 (34%), Positives = 161/330 (47%), Gaps = 37/330 (11%)
Query: 81 WY-VTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPT--G 137
WY V+ + G I E +L N +E + F L +++ + S N L +IP G
Sbjct: 68 WYGVSCNSRGSIEELNL--TNTGIEGTFQDFPFISLSNLAYVDL--SMNLLSGTIPPQFG 123
Query: 138 NWEKLAESLESIEFR-SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLK 196
N KL I F S L G I + G LKNL L L +N LT IP E+GN+ +
Sbjct: 124 NLSKL------IYFDLSTNHLTGEISPSLGNLKNLTVLYLHQNYLTSVIPSELGNMESMT 177
Query: 197 RLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGK 256
L LS N +G+IP G L +L++L L N L+G +P LG + S+ L LS N L G
Sbjct: 178 DLALSQNKLTGSIPSSLGNLKNLMVLYLYENYLTGVIPPELGNMESMTDLALSQNKLTGS 237
Query: 257 LLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTL-------- 308
+ + GNLKNL ++ L N L + + M S+ + LS N L G I +
Sbjct: 238 IPSTLGNLKNLMVLYLYENYLTGVIPPEIGNMESMTNLALSQNKLTGSIPSSLGNLKNLT 297
Query: 309 ---------------KWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGN 353
K N+++++ LELSN +L G IP SL LK L L L +N +TG
Sbjct: 298 LLSLFQNYLTGGIPPKLGNIESMIDLELSNNKLTGSIPSSLGNLKNLTILYLYENYLTGV 357
Query: 354 LSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
+ P+L + S+ L L+ N L G I S G
Sbjct: 358 IPPELGNMESMIDLQLNNNKLTGSIPSSFG 387
Score = 136 bits (343), Expect = 1e-30
Identities = 114/318 (35%), Positives = 157/318 (48%), Gaps = 30/318 (9%)
Query: 112 LKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLK 169
L +LK ++ + FQ N L IP GN ES+ +E SN L G+IPS+ G LK
Sbjct: 290 LGNLKNLTLLSLFQ--NYLTGGIPPKLGN----IESMIDLEL-SNNKLTGSIPSSLGNLK 342
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
NL L L EN LTG IP E+GN+ + L L+ N +G+IP FG L +L L L N L
Sbjct: 343 NLTILYLYENYLTGVIPPELGNMESMIDLQLNNNKLTGSIPSSFGNLKNLTYLYLYLNYL 402
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMN 289
+G +P LG + S++ LDLS N L G + + FGN L + LR N L + + +
Sbjct: 403 TGVIPQELGNMESMINLDLSQNKLTGSVPDSFGNFTKLESLYLRVNHLSGAIPPGVANSS 462
Query: 290 SLEEMVLSNNPLGGDI--RTLKWENLQNLVILELSNMELIGEIPESLSQLKKL---RFLG 344
L ++L N G K LQN + L+ +++E G IP+SL K L RFLG
Sbjct: 463 HLTTLILDTNNFTGFFPETVCKGRKLQN-ISLDYNHLE--GPIPKSLRDCKSLIRARFLG 519
Query: 345 LSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPF 404
N TG++ P LN + S N GEI + KLG + +N P
Sbjct: 520 ---NKFTGDIFEAFGIYPDLNFIDFSHNKFHGEISSNWEKSPKLGALIMSNNNITGAIPT 576
Query: 405 EL----------MSTNNV 412
E+ +STNN+
Sbjct: 577 EIWNMTQLVELDLSTNNL 594
Score = 112 bits (281), Expect = 2e-23
Identities = 80/243 (32%), Positives = 116/243 (46%), Gaps = 6/243 (2%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G+I FG+ +L + N G I KL L++S NN +G IP ++
Sbjct: 524 GDIFEAFGIYPDLNFIDFSHNKFHGEISSNWEKSPKLGALIMSNNNITGAIPTEIWNMTQ 583
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L+ LDLS N+L G LP +G L ++ +L L+ N L G++ L NL +DL +N
Sbjct: 584 LVELDLSTNNLFGELPEAIGNLTNLSRLRLNGNQLSGRVPAGLSFLTNLESLDLSSNNFS 643
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLK 338
+ + L +M LS N G I L L L L+LS+ +L GEIP LS L+
Sbjct: 644 SEIPQTFDSFLKLHDMNLSRNKFDGSIPRL--SKLTQLTQLDLSHNQLDGEIPSQLSSLQ 701
Query: 339 KLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI----QFSKGFFGKLGRRFGA 394
L L LS NN++G + E + +L + +S N L+G + F K L G
Sbjct: 702 SLDKLDLSHNNLSGLIPTTFEGMIALTNVDISNNKLEGPLPDTPTFRKATADALEENIGL 761
Query: 395 WSN 397
SN
Sbjct: 762 CSN 764
Score = 104 bits (260), Expect = 5e-21
Identities = 95/309 (30%), Positives = 141/309 (44%), Gaps = 55/309 (17%)
Query: 119 SFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSN-------PGLI----------- 158
S N S NKL S+P GN+ KL ES+ R N PG+
Sbjct: 415 SMINLDLSQNKLTGSVPDSFGNFTKL----ESLYLRVNHLSGAIPPGVANSSHLTTLILD 470
Query: 159 -----GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF 213
G P T + LQ++ L N L G IP+ + + L R GN F+G+I + F
Sbjct: 471 TNNFTGFFPETVCKGRKLQNISLDYNHLEGPIPKSLRDCKSLIRARFLGNKFTGDIFEAF 530
Query: 214 GGLSDLLILD------------------------LSRNSLSGTLPVTLGRLISVLKLDLS 249
G DL +D +S N+++G +P + + +++LDLS
Sbjct: 531 GIYPDLNFIDFSHNKFHGEISSNWEKSPKLGALIMSNNNITGAIPTEIWNMTQLVELDLS 590
Query: 250 HNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLK 309
N L G+L GNL NL+ + L N+L + L + +LE + LS+N +I
Sbjct: 591 TNNLFGELPEAIGNLTNLSRLRLNGNQLSGRVPAGLSFLTNLESLDLSSNNFSSEI-PQT 649
Query: 310 WENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYL 369
+++ L + LS + G IP LS+L +L L LS N + G + +L +L SL+ L L
Sbjct: 650 FDSFLKLHDMNLSRNKFDGSIPR-LSKLTQLTQLDLSHNQLDGEIPSQLSSLQSLDKLDL 708
Query: 370 SGNNLKGEI 378
S NNL G I
Sbjct: 709 SHNNLSGLI 717
Score = 102 bits (253), Expect = 3e-20
Identities = 68/198 (34%), Positives = 107/198 (53%), Gaps = 4/198 (2%)
Query: 136 TGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKL 195
+ NWEK + L ++ SN + G IP+ + L L L N L G +P+ IGNL L
Sbjct: 551 SSNWEK-SPKLGAL-IMSNNNITGAIPTEIWNMTQLVELDLSTNNLFGELPEAIGNLTNL 608
Query: 196 KRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEG 255
RL L+GN SG +P L++L LDLS N+ S +P T + + ++LS N +G
Sbjct: 609 SRLRLNGNQLSGRVPAGLSFLTNLESLDLSSNNFSSEIPQTFDSFLKLHDMNLSRNKFDG 668
Query: 256 KLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQN 315
+ L LT +DL +N+L + L + SL+++ LS+N L G I T +E +
Sbjct: 669 S-IPRLSKLTQLTQLDLSHNQLDGEIPSQLSSLQSLDKLDLSHNNLSGLIPT-TFEGMIA 726
Query: 316 LVILELSNMELIGEIPES 333
L +++SN +L G +P++
Sbjct: 727 LTNVDISNNKLEGPLPDT 744
>gb|AAC78591.1| disease resistance protein [Lycopersicon esculentum]
Length = 968
Score = 155 bits (392), Expect = 3e-36
Identities = 112/308 (36%), Positives = 155/308 (49%), Gaps = 13/308 (4%)
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPST 164
P++ L L+ I FN N L IP G L + I F S G+IP++
Sbjct: 137 PQIGSLAKLQIIRIFN-----NHLNGFIPEEIGYLRSLTKLSLGINFLS-----GSIPAS 186
Query: 165 FGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDL 224
G + NL L L EN L+G IP+EIG L L +L L N SG+IP G L++L L L
Sbjct: 187 LGNMTNLSFLFLYENQLSGFIPEEIGYLRSLTKLSLDINFLSGSIPASLGNLNNLSFLYL 246
Query: 225 SRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLS 284
N LSG++P +G L S+ KL L NFL G + GNL NL+ +DL NN+L +
Sbjct: 247 YNNQLSGSIPEEIGYLRSLTKLSLGINFLSGSIPASLGNLNNLSRLDLYNNKLSGSIPEE 306
Query: 285 LQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLG 344
+ + SL + L N L G I + NL NL L+L N +L G IPE + L+ L +L
Sbjct: 307 IGYLRSLTYLDLGENALNGSIPS-SLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTYLD 365
Query: 345 LSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPF 404
L +N + G++ L L +L LYL N L G I G+ L + ++ P
Sbjct: 366 LGENALNGSIPASLGNLNNLFMLYLYNNQLSGSIPEEIGYLSSLTELYLGNNSLNGSIPA 425
Query: 405 ELMSTNNV 412
L + NN+
Sbjct: 426 SLGNLNNL 433
Score = 153 bits (387), Expect = 1e-35
Identities = 99/256 (38%), Positives = 136/256 (52%), Gaps = 1/256 (0%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G+IP++ G L NL L L N L+G+IP+EIG L L +L L N SG+IP G L
Sbjct: 227 LSGSIPASLGNLNNLSFLYLYNNQLSGSIPEEIGYLRSLTKLSLGINFLSGSIPASLGNL 286
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
++L LDL N LSG++P +G L S+ LDL N L G + + GNL NL+ +DL NN+
Sbjct: 287 NNLSRLDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPSSLGNLNNLSRLDLYNNK 346
Query: 277 LCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQ 336
L + + + SL + L N L G I NL NL +L L N +L G IPE +
Sbjct: 347 LSGSIPEEIGYLRSLTYLDLGENALNGSIPA-SLGNLNNLFMLYLYNNQLSGSIPEEIGY 405
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWS 396
L L L L +N++ G++ L L +L LYL N L G I G+ L F +
Sbjct: 406 LSSLTELYLGNNSLNGSIPASLGNLNNLFMLYLYNNQLSGSIPEEIGYLSSLTELFLGNN 465
Query: 397 NPKLCYPFELMSTNNV 412
+ P L + NN+
Sbjct: 466 SLNGSIPASLGNLNNL 481
Score = 150 bits (380), Expect = 6e-35
Identities = 104/267 (38%), Positives = 142/267 (52%), Gaps = 8/267 (2%)
Query: 128 NKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
N L SIP GN L S + N L G+IP G L++L L L N L+G+I
Sbjct: 225 NFLSGSIPASLGNLNNL-----SFLYLYNNQLSGSIPEEIGYLRSLTKLSLGINFLSGSI 279
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P +GNL L RL L N SG+IP+ G L L LDL N+L+G++P +LG L ++ +
Sbjct: 280 PASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPSSLGNLNNLSR 339
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
LDL +N L G + E G L++LT +DL N L + SL +N+L + L NN L G I
Sbjct: 340 LDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLFMLYLYNNQLSGSI 399
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLN 365
+ L +L L L N L G IP SL L L L L +N ++G++ ++ L SL
Sbjct: 400 PE-EIGYLSSLTELYLGNNSLNGSIPASLGNLNNLFMLYLYNNQLSGSIPEEIGYLSSLT 458
Query: 366 ALYLSGNNLKGEIQFSKGFFGKLGRRF 392
L+L N+L G I S G L R +
Sbjct: 459 ELFLGNNSLNGSIPASLGNLNNLSRLY 485
Score = 150 bits (378), Expect = 1e-34
Identities = 103/282 (36%), Positives = 151/282 (53%), Gaps = 16/282 (5%)
Query: 109 LFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLA--ESLESIEFRSNPGLIGNIPSTFG 166
L L +L + +N NKL SIP E++ SL ++ N L G+IP++ G
Sbjct: 331 LGNLNNLSRLDLYN-----NKLSGSIP----EEIGYLRSLTYLDLGEN-ALNGSIPASLG 380
Query: 167 VLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
L NL L L N L+G+IP+EIG L L L L N+ +G+IP G L++L +L L
Sbjct: 381 NLNNLFMLYLYNNQLSGSIPEEIGYLSSLTELYLGNNSLNGSIPASLGNLNNLFMLYLYN 440
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N LSG++P +G L S+ +L L +N L G + GNL NL+ + L NN+L + S
Sbjct: 441 NQLSGSIPEEIGYLSSLTELFLGNNSLNGSIPASLGNLNNLSRLYLYNNQLSGSIPASFG 500
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
M +L+ + LS+N L G+I + NL +L +L +S L G++P+ L + L L +S
Sbjct: 501 NMRNLQTLFLSDNDLIGEIPSFVC-NLTSLEVLYMSRNNLKGKVPQCLGNISDLHILSMS 559
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
N+ G L + L SL L NNL+G I FFG +
Sbjct: 560 SNSFRGELPSSISNLTSLKILDFGRNNLEGAI---PQFFGNI 598
Score = 137 bits (346), Expect = 6e-31
Identities = 96/265 (36%), Positives = 129/265 (48%), Gaps = 32/265 (12%)
Query: 119 SFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVL 176
S F N L SIP GN L+ + N L G+IP++FG ++NLQ+L L
Sbjct: 456 SLTELFLGNNSLNGSIPASLGNLNNLSRL-----YLYNNQLSGSIPASFGNMRNLQTLFL 510
Query: 177 LENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVT 236
+N L G IP + NL L+ L +S NN G +P G +SDL IL +S NS G LP +
Sbjct: 511 SDNDLIGEIPSFVCNLTSLEVLYMSRNNLKGKVPQCLGNISDLHILSMSSNSFRGELPSS 570
Query: 237 LGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVL 296
+ L S+ LD N LEG + FGN+ +L + D++NN+L L
Sbjct: 571 ISNLTSLKILDFGRNNLEGAIPQFFGNISSLQVFDMQNNKLSGTLP-------------- 616
Query: 297 SNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSP 356
+N +G +L+ L L EL EIP SL KKL+ L L DN +
Sbjct: 617 TNFSIG-----------CSLISLNLHGNELADEIPRSLDNCKKLQVLDLGDNQLNDTFPM 665
Query: 357 KLETLPSLNALYLSGNNLKGEIQFS 381
L TLP L L L+ N L G I+ S
Sbjct: 666 WLGTLPELRVLRLTSNKLHGPIRSS 690
Score = 137 bits (345), Expect = 7e-31
Identities = 102/307 (33%), Positives = 147/307 (47%), Gaps = 25/307 (8%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G IP G L LQ + + N L G IP+EIG L L +L L N SG+IP G +++
Sbjct: 133 GTIPPQIGSLAKLQIIRIFNNHLNGFIPEEIGYLRSLTKLSLGINFLSGSIPASLGNMTN 192
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L L L N LSG +P +G L S+ KL L NFL G + GNL NL+ + L NN+L
Sbjct: 193 LSFLFLYENQLSGFIPEEIGYLRSLTKLSLDINFLSGSIPASLGNLNNLSFLYLYNNQLS 252
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLK 338
+ + + SL ++ L N L G I NL NL L+L N +L G IPE + L+
Sbjct: 253 GSIPEEIGYLRSLTKLSLGINFLSGSI-PASLGNLNNLSRLDLYNNKLSGSIPEEIGYLR 311
Query: 339 KLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFF------------- 385
L +L L +N + G++ L L +L+ L L N L G I G+
Sbjct: 312 SLTYLDLGENALNGSIPSSLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTYLDLGENAL 371
Query: 386 -GKLGRRFGAWSNPKLCYPFELMSTNNVPYGVKPCHQEEIHLVKSNAKTEVINGDINHNS 444
G + G +N + Y + + ++P EEI + S TE+ G+ + N
Sbjct: 372 NGSIPASLGNLNNLFMLYLYNNQLSGSIP--------EEIGYLSS--LTELYLGNNSLNG 421
Query: 445 NFITSMG 451
+ S+G
Sbjct: 422 SIPASLG 428
Score = 126 bits (317), Expect = 1e-27
Identities = 86/237 (36%), Positives = 122/237 (51%), Gaps = 2/237 (0%)
Query: 153 SNPGLIGNIPS-TFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
+N +IG + + F L L++L L N ++G IP EIGNL L L L+ N SG IP
Sbjct: 78 TNASVIGTLYAFPFSSLPFLENLDLSNNNISGTIPPEIGNLTNLVYLDLNTNQISGTIPP 137
Query: 212 IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMD 271
G L+ L I+ + N L+G +P +G L S+ KL L NFL G + GN+ NL+ +
Sbjct: 138 QIGSLAKLQIIRIFNNHLNGFIPEEIGYLRSLTKLSLGINFLSGSIPASLGNMTNLSFLF 197
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
L N+L + + + SL ++ L N L G I NL NL L L N +L G IP
Sbjct: 198 LYENQLSGFIPEEIGYLRSLTKLSLDINFLSGSI-PASLGNLNNLSFLYLYNNQLSGSIP 256
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
E + L+ L L L N ++G++ L L +L+ L L N L G I G+ L
Sbjct: 257 EEIGYLRSLTKLSLGINFLSGSIPASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSL 313
Score = 62.4 bits (150), Expect = 3e-08
Identities = 35/80 (43%), Positives = 47/80 (58%)
Query: 176 LLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPV 235
L N G+IP +G+L+ ++ L +S N G IP G LS L LDLS N LSG +P
Sbjct: 767 LSSNKFEGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEIPQ 826
Query: 236 TLGRLISVLKLDLSHNFLEG 255
L L + L+LSHN+L+G
Sbjct: 827 QLASLTFLEVLNLSHNYLQG 846
Score = 61.2 bits (147), Expect = 7e-08
Identities = 34/98 (34%), Positives = 58/98 (58%), Gaps = 1/98 (1%)
Query: 181 LTGNIPQEIGNLVKLKRLV-LSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGR 239
+T + EI ++ L ++ LS N F G+IP + G L + +L++S N+L G +P +LG
Sbjct: 747 VTKGLELEIVRILSLYTIIDLSSNKFEGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGS 806
Query: 240 LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L + LDLS N L G++ + +L L +++L +N L
Sbjct: 807 LSILESLDLSFNQLSGEIPQQLASLTFLEVLNLSHNYL 844
Score = 58.2 bits (139), Expect = 6e-07
Identities = 81/316 (25%), Positives = 124/316 (38%), Gaps = 81/316 (25%)
Query: 55 NGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKH 114
N SDL+ S S G + L + +++ G N+L A P+ F +
Sbjct: 549 NISDLHILSMSSNSFRGELPSSISNLTSLKILDFG---RNNLEGAI------PQFFG--N 597
Query: 115 LKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSL 174
+ ++ F+ NKL ++PT + SL S+ N L IP + K LQ L
Sbjct: 598 ISSLQVFD--MQNNKLSGTLPTNF--SIGCSLISLNLHGNE-LADEIPRSLDNCKKLQVL 652
Query: 175 VLLENGLTGNIPQEIGNLVKLK-------------------------RLV-LSGNNFSGN 208
L +N L P +G L +L+ R++ LS N FS +
Sbjct: 653 DLGDNQLNDTFPMWLGTLPELRVLRLTSNKLHGPIRSSGAEIMFPDLRIIDLSRNAFSQD 712
Query: 209 IP---------------------------------------DIFGGLSDLLILDLSRNSL 229
+P +I LS I+DLS N
Sbjct: 713 LPTSLFEHLKGMRTVDKTMEEPSYESYYDDSVVVVTKGLELEIVRILSLYTIIDLSSNKF 772
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMN 289
G +P LG LI++ L++SHN L+G + + G+L L +DL N+L + L +
Sbjct: 773 EGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGSLSILESLDLSFNQLSGEIPQQLASLT 832
Query: 290 SLEEMVLSNNPLGGDI 305
LE + LS+N L G I
Sbjct: 833 FLEVLNLSHNYLQGCI 848
Score = 55.8 bits (133), Expect = 3e-06
Identities = 36/90 (40%), Positives = 49/90 (54%)
Query: 145 SLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNN 204
SL +I S+ G+IPS G L ++ L + N L G IP +G+L L+ L LS N
Sbjct: 760 SLYTIIDLSSNKFEGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGSLSILESLDLSFNQ 819
Query: 205 FSGNIPDIFGGLSDLLILDLSRNSLSGTLP 234
SG IP L+ L +L+LS N L G +P
Sbjct: 820 LSGEIPQQLASLTFLEVLNLSHNYLQGCIP 849
Score = 48.5 bits (114), Expect = 4e-04
Identities = 36/132 (27%), Positives = 60/132 (45%), Gaps = 7/132 (5%)
Query: 261 FGNLKNLTLMDLRNNRLCC----GLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNL 316
F N N L + C G+V +N+L ++N + G + + +L L
Sbjct: 41 FKNQNNSFLASWTTSSNACKDWYGVVCLNGRVNTLN---ITNASVIGTLYAFPFSSLPFL 97
Query: 317 VILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKG 376
L+LSN + G IP + L L +L L+ N I+G + P++ +L L + + N+L G
Sbjct: 98 ENLDLSNNNISGTIPPEIGNLTNLVYLDLNTNQISGTIPPQIGSLAKLQIIRIFNNHLNG 157
Query: 377 EIQFSKGFFGKL 388
I G+ L
Sbjct: 158 FIPEEIGYLRSL 169
>ref|NP_917751.1| P0501G01.13 [Oryza sativa (japonica cultivar-group)]
Length = 494
Score = 153 bits (386), Expect = 1e-35
Identities = 121/398 (30%), Positives = 179/398 (44%), Gaps = 67/398 (16%)
Query: 35 EKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNG-LWYVTVINIGPIHE 93
+ AEQ A+ + GN W D C G+ C G +++V ++ G + +
Sbjct: 58 DPAEQRAVQE-VMAATGNGWASG--IADVCRGRW-HGIECVPDRGEVYHVVSLSFGALSD 113
Query: 94 NSL--PCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEF 151
++ C + P + L HL+++ F+ CF + N PV
Sbjct: 114 DTAFPACDAARATLSPAVLALPHLRSLFFYRCFTA-NPQPV------------------- 153
Query: 152 RSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
P +G + F +SLVL ENG G IP E+GNL L+ L L GNN + IP
Sbjct: 154 ---PAFLGRLGPAF------RSLVLRENGHVGAIPPELGNLTALRVLDLHGNNLTSAIPA 204
Query: 212 IFGGLS----------------------DLLILDLSRNSLSGTLPVTLGRLISVLKLDLS 249
L+ L ILDLS N+L G +P +LG+ S+LK DLS
Sbjct: 205 TVQSLAHLQLLDLSYNQLAGEVPPFKFQHLSILDLSHNALQGGVPASLGQCRSLLKFDLS 264
Query: 250 HNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNN-----PLGGD 304
N G + + G+L +L L+DL +N L + +L ++SL ++L +N + GD
Sbjct: 265 QNRFAGTIPDALGDLSDLILLDLSHNALSGPIPAALGRLSSLRSLILGDNRMQFTTVPGD 324
Query: 305 IRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSL 364
I + L+ L L LS M L G +PES+ +L LR L L +N TG + L
Sbjct: 325 I----FAGLRALTTLVLSGMGLEGSLPESIGELGHLRVLRLDNNEFTGVIPASFRRLERA 380
Query: 365 NALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCY 402
+ L + GN L G I F K +LG++ N LCY
Sbjct: 381 SELRVDGNRLVGPIPFGKQMMWRLGKKLRVGGNEGLCY 418
>ref|NP_173217.1| leucine-rich repeat transmembrane protein kinase, putative
[Arabidopsis thaliana] gi|25513592|pir||E86312 F11A6.9
protein - Arabidopsis thaliana gi|9802748|gb|AAF99817.1|
Unknown protein [Arabidopsis thaliana]
Length = 1088
Score = 152 bits (385), Expect = 2e-35
Identities = 112/325 (34%), Positives = 156/325 (47%), Gaps = 41/325 (12%)
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
S GL G + S G LK+L +L L N +G +P +GN L+ L LS N+FSG +PDI
Sbjct: 84 SASGLSGQLGSEIGELKSLVTLDLSLNSFSGLLPSTLGNCTSLEYLDLSNNDFSGEVPDI 143
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDL 272
FG L +L L L RN+LSG +P ++G LI ++ L +S+N L G + GN L + L
Sbjct: 144 FGSLQNLTFLYLDRNNLSGLIPASVGGLIELVDLRMSYNNLSGTIPELLGNCSKLEYLAL 203
Query: 273 RNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELS---------- 322
NN+L L SL + +L E+ +SNN LGG + N + LV L+LS
Sbjct: 204 NNNKLNGSLPASLYLLENLGELFVSNNSLGGRLH-FGSSNCKKLVSLDLSFNDFQGGVPP 262
Query: 323 --------------NMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALY 368
L G IP S+ L+K+ + LSDN ++GN+ +L SL L
Sbjct: 263 EIGNCSSLHSLVMVKCNLTGTIPSSMGMLRKVSVIDLSDNRLSGNIPQELGNCSSLETLK 322
Query: 369 LSGNNLKGEI----------QFSKGFFGKLGRR--FGAWSNPKLCYPFELMSTNNVPYGV 416
L+ N L+GEI Q + FF KL G W L +++ NN G
Sbjct: 323 LNDNQLQGEIPPALSKLKKLQSLELFFNKLSGEIPIGIWKIQSLT---QMLVYNNTLTGE 379
Query: 417 KPCHQEEI-HLVKSNAKTEVINGDI 440
P ++ HL K GDI
Sbjct: 380 LPVEVTQLKHLKKLTLFNNGFYGDI 404
Score = 123 bits (308), Expect = 1e-26
Identities = 104/331 (31%), Positives = 156/331 (46%), Gaps = 38/331 (11%)
Query: 72 VSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQLKHLKAISFFNCFQSPNKLP 131
+S +I G+W + + ++ N+L E E+ QLKHLK ++ FN +P
Sbjct: 352 LSGEIPIGIWKIQSLTQMLVYNNTLTG-----ELPVEVTQLKHLKKLTLFNN-GFYGDIP 405
Query: 132 VSIPTGNWEKLAESLESIEFRSNP-----------------------GLIGNIPSTFGVL 168
+S+ L SLE ++ N L G IP++
Sbjct: 406 MSLG------LNRSLEEVDLLGNRFTGEIPPHLCHGQKLRLFILGSNQLHGKIPASIRQC 459
Query: 169 KNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNS 228
K L+ + L +N L+G +P E + L + L N+F G+IP G +LL +DLS+N
Sbjct: 460 KTLERVRLEDNKLSGVLP-EFPESLSLSYVNLGSNSFEGSIPRSLGSCKNLLTIDLSQNK 518
Query: 229 LSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEM 288
L+G +P LG L S+ L+LSHN+LEG L ++ L D+ +N L + S +
Sbjct: 519 LTGLIPPELGNLQSLGLLNLSHNYLEGPLPSQLSGCARLLYFDVGSNSLNGSIPSSFRSW 578
Query: 289 NSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRF-LGLSD 347
SL +VLS+N G I E L L L ++ G+IP S+ LK LR+ L LS
Sbjct: 579 KSLSTLVLSDNNFLGAIPQFLAE-LDRLSDLRIARNAFGGKIPSSVGLLKSLRYGLDLSA 637
Query: 348 NNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
N TG + L L +L L +S N L G +
Sbjct: 638 NVFTGEIPTTLGALINLERLNISNNKLTGPL 668
Score = 118 bits (296), Expect = 4e-25
Identities = 82/227 (36%), Positives = 123/227 (54%), Gaps = 2/227 (0%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IPS+ G+L+ + + L +N L+GNIPQE+GN L+ L L+ N G IP L
Sbjct: 280 LTGTIPSSMGMLRKVSVIDLSDNRLSGNIPQELGNCSSLETLKLNDNQLQGEIPPALSKL 339
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
L L+L N LSG +P+ + ++ S+ ++ + +N L G+L E LK+L + L NN
Sbjct: 340 KKLQSLELFFNKLSGEIPIGIWKIQSLTQMLVYNNTLTGELPVEVTQLKHLKKLTLFNNG 399
Query: 277 LCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQ 336
+ +SL SLEE+ L N G+I Q L + L + +L G+IP S+ Q
Sbjct: 400 FYGDIPMSLGLNRSLEEVDLLGNRFTGEIPPHLCHG-QKLRLFILGSNQLHGKIPASIRQ 458
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
K L + L DN ++G L E+L SL+ + L N+ +G I S G
Sbjct: 459 CKTLERVRLEDNKLSGVLPEFPESL-SLSYVNLGSNSFEGSIPRSLG 504
Score = 115 bits (288), Expect = 3e-24
Identities = 87/254 (34%), Positives = 126/254 (49%), Gaps = 4/254 (1%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL S+P + L E+L + F SN L G + K L SL L N G +P
Sbjct: 206 NKLNGSLPASLY--LLENLGEL-FVSNNSLGGRLHFGSSNCKKLVSLDLSFNDFQGGVPP 262
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD 247
EIGN L LV+ N +G IP G L + ++DLS N LSG +P LG S+ L
Sbjct: 263 EIGNCSSLHSLVMVKCNLTGTIPSSMGMLRKVSVIDLSDNRLSGNIPQELGNCSSLETLK 322
Query: 248 LSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT 307
L+ N L+G++ LK L ++L N+L + + + ++ SL +M++ NN L G++
Sbjct: 323 LNDNQLQGEIPPALSKLKKLQSLELFFNKLSGEIPIGIWKIQSLTQMLVYNNTLTGEL-P 381
Query: 308 LKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNAL 367
++ L++L L L N G+IP SL + L + L N TG + P L L
Sbjct: 382 VEVTQLKHLKKLTLFNNGFYGDIPMSLGLNRSLEEVDLLGNRFTGEIPPHLCHGQKLRLF 441
Query: 368 YLSGNNLKGEIQFS 381
L N L G+I S
Sbjct: 442 ILGSNQLHGKIPAS 455
Score = 108 bits (270), Expect = 4e-22
Identities = 96/313 (30%), Positives = 140/313 (44%), Gaps = 34/313 (10%)
Query: 104 EFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPS 163
E P L +LK L+++ F NKL IP G W+ +SL + +N L G +P
Sbjct: 331 EIPPALSKLKKLQSLELFF-----NKLSGEIPIGIWK--IQSLTQMLVYNNT-LTGELPV 382
Query: 164 TFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILD 223
LK+L+ L L NG G+IP +G L+ + L GN F+G IP L +
Sbjct: 383 EVTQLKHLKKLTLFNNGFYGDIPMSLGLNRSLEEVDLLGNRFTGEIPPHLCHGQKLRLFI 442
Query: 224 LSRNSLSGTLPV------TLGRL-----------------ISVLKLDLSHNFLEGKLLNE 260
L N L G +P TL R+ +S+ ++L N EG +
Sbjct: 443 LGSNQLHGKIPASIRQCKTLERVRLEDNKLSGVLPEFPESLSLSYVNLGSNSFEGSIPRS 502
Query: 261 FGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILE 320
G+ KNL +DL N+L + L + SL + LS+N L G + + + L+ +
Sbjct: 503 LGSCKNLLTIDLSQNKLTGLIPPELGNLQSLGLLNLSHNYLEGPLPS-QLSGCARLLYFD 561
Query: 321 LSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQF 380
+ + L G IP S K L L LSDNN G + L L L+ L ++ N G+I
Sbjct: 562 VGSNSLNGSIPSSFRSWKSLSTLVLSDNNFLGAIPQFLAELDRLSDLRIARNAFGGKIPS 621
Query: 381 SKGFFGKLGRRFG 393
S G L R+G
Sbjct: 622 SVGLLKSL--RYG 632
Score = 59.7 bits (143), Expect = 2e-07
Identities = 46/133 (34%), Positives = 67/133 (49%), Gaps = 9/133 (6%)
Query: 128 NKLPVSIPTG--NWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
N L SIP+ +W+ L+ + S+ +G IP L L L + N G I
Sbjct: 565 NSLNGSIPSSFRSWKSLSTLV-----LSDNNFLGAIPQFLAELDRLSDLRIARNAFGGKI 619
Query: 186 PQEIGNLVKLKR-LVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVL 244
P +G L L+ L LS N F+G IP G L +L L++S N L+G L V L L S+
Sbjct: 620 PSSVGLLKSLRYGLDLSANVFTGEIPTTLGALINLERLNISNNKLTGPLSV-LQSLKSLN 678
Query: 245 KLDLSHNFLEGKL 257
++D+S+N G +
Sbjct: 679 QVDVSYNQFTGPI 691
Score = 54.3 bits (129), Expect = 8e-06
Identities = 32/80 (40%), Positives = 47/80 (58%), Gaps = 2/80 (2%)
Query: 159 GNIPSTFGVLKNLQ-SLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLS 217
G IPS+ G+LK+L+ L L N TG IP +G L+ L+RL +S N +G + + L
Sbjct: 617 GKIPSSVGLLKSLRYGLDLSANVFTGEIPTTLGALINLERLNISNNKLTGPL-SVLQSLK 675
Query: 218 DLLILDLSRNSLSGTLPVTL 237
L +D+S N +G +PV L
Sbjct: 676 SLNQVDVSYNQFTGPIPVNL 695
>gb|AAC78593.1| Hcr2-0B [Lycopersicon esculentum]
Length = 944
Score = 152 bits (385), Expect = 2e-35
Identities = 104/258 (40%), Positives = 140/258 (53%), Gaps = 8/258 (3%)
Query: 128 NKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
N L SIP GN L S + N L G+IP G L++L L L EN L G+I
Sbjct: 225 NFLSGSIPASLGNLNNL-----SFLYLYNNQLSGSIPEEIGYLRSLTYLDLGENALNGSI 279
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P +GNL L RL L N SG+IP+ G L L LDL N+L+G++P +LG L ++ +
Sbjct: 280 PASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSR 339
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
LDL +N L G + E G L++LT +DL N L + SL +N+L + L NN L G I
Sbjct: 340 LDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSRLDLYNNKLSGSI 399
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLN 365
+ L++L L L N L G IP SL L L L L +N ++G++ ++ L SL
Sbjct: 400 PE-EIGYLRSLTKLSLGNNFLSGSIPASLGNLNNLFMLYLYNNQLSGSIPEEIGYLSSLT 458
Query: 366 ALYLSGNNLKGEIQFSKG 383
LYL N+L G I S G
Sbjct: 459 NLYLGNNSLNGLIPASFG 476
Score = 144 bits (363), Expect = 6e-33
Identities = 105/286 (36%), Positives = 142/286 (48%), Gaps = 30/286 (10%)
Query: 128 NKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
N L SIP GN L S F L G IP G L++L L L N L+G+I
Sbjct: 177 NFLSGSIPASLGNMTNL-----SFLFLYENQLSGFIPEEIGYLRSLTKLSLDINFLSGSI 231
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P +GNL L L L N SG+IP+ G L L LDL N+L+G++P +LG L ++ +
Sbjct: 232 PASLGNLNNLSFLYLYNNQLSGSIPEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSR 291
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
LDL +N L G + E G L++LT +DL N L + SL +N+L + L NN L G I
Sbjct: 292 LDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSRLDLYNNKLSGSI 351
Query: 306 -------RTLKW----------------ENLQNLVILELSNMELIGEIPESLSQLKKLRF 342
R+L + NL NL L+L N +L G IPE + L+ L
Sbjct: 352 PEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTK 411
Query: 343 LGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
L L +N ++G++ L L +L LYL N L G I G+ L
Sbjct: 412 LSLGNNFLSGSIPASLGNLNNLFMLYLYNNQLSGSIPEEIGYLSSL 457
Score = 137 bits (345), Expect = 7e-31
Identities = 88/230 (38%), Positives = 119/230 (51%), Gaps = 1/230 (0%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G IP G L LQ + + N L G IP+EIG L L +L L N SG+IP G +++
Sbjct: 133 GTIPPQIGSLAKLQIIRIFNNHLNGFIPEEIGYLRSLTKLSLGINFLSGSIPASLGNMTN 192
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L L L N LSG +P +G L S+ KL L NFL G + GNL NL+ + L NN+L
Sbjct: 193 LSFLFLYENQLSGFIPEEIGYLRSLTKLSLDINFLSGSIPASLGNLNNLSFLYLYNNQLS 252
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLK 338
+ + + SL + L N L G I NL NL L+L N +L G IPE + L+
Sbjct: 253 GSIPEEIGYLRSLTYLDLGENALNGSI-PASLGNLNNLSRLDLYNNKLSGSIPEEIGYLR 311
Query: 339 KLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
L +L L +N + G++ L L +L+ L L N L G I G+ L
Sbjct: 312 SLTYLDLGENALNGSIPASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSL 361
Score = 136 bits (342), Expect = 2e-30
Identities = 95/270 (35%), Positives = 145/270 (53%), Gaps = 10/270 (3%)
Query: 111 QLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVL 168
++ +L+++++ + + N L SIP GN L+ ++ +N L G+IP G L
Sbjct: 306 EIGYLRSLTYLDLGE--NALNGSIPASLGNLNNLSR----LDLYNNK-LSGSIPEEIGYL 358
Query: 169 KNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNS 228
++L L L EN L G+IP +GNL L RL L N SG+IP+ G L L L L N
Sbjct: 359 RSLTYLDLGENALNGSIPASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTKLSLGNNF 418
Query: 229 LSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEM 288
LSG++P +LG L ++ L L +N L G + E G L +LT + L NN L + S M
Sbjct: 419 LSGSIPASLGNLNNLFMLYLYNNQLSGSIPEEIGYLSSLTNLYLGNNSLNGLIPASFGNM 478
Query: 289 NSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDN 348
+L+ + L++N L G+I + NL +L +L + L G++P+ L + L L +S N
Sbjct: 479 RNLQALFLNDNNLIGEIPSFVC-NLTSLELLYMPRNNLKGKVPQCLGNISDLLVLSMSSN 537
Query: 349 NITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+ +G L + L SL L NNL+G I
Sbjct: 538 SFSGELPSSISNLTSLKILDFGRNNLEGAI 567
Score = 135 bits (340), Expect = 3e-30
Identities = 95/265 (35%), Positives = 134/265 (49%), Gaps = 32/265 (12%)
Query: 119 SFFNCFQSPNKLPVSIPTGNWEKLA--ESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVL 176
+ F + N+L SIP E++ SL ++ + N L G IP++FG ++NLQ+L L
Sbjct: 432 NLFMLYLYNNQLSGSIP----EEIGYLSSLTNL-YLGNNSLNGLIPASFGNMRNLQALFL 486
Query: 177 LENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVT 236
+N L G IP + NL L+ L + NN G +P G +SDLL+L +S NS SG LP +
Sbjct: 487 NDNNLIGEIPSFVCNLTSLELLYMPRNNLKGKVPQCLGNISDLLVLSMSSNSFSGELPSS 546
Query: 237 LGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVL 296
+ L S+ LD N LEG + FGN+ +L + D++NN+L L
Sbjct: 547 ISNLTSLKILDFGRNNLEGAIPQCFGNISSLQVFDMQNNKLSGTLP-------------- 592
Query: 297 SNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSP 356
+N +G +L+ L L EL EIP SL KKL+ L L DN +
Sbjct: 593 TNFSIG-----------CSLISLNLHGNELEDEIPWSLDNCKKLQVLDLGDNQLNDTFPM 641
Query: 357 KLETLPSLNALYLSGNNLKGEIQFS 381
L TLP L L L+ N L G I+ S
Sbjct: 642 WLGTLPELRVLRLTSNKLHGPIRSS 666
Score = 132 bits (333), Expect = 2e-29
Identities = 110/320 (34%), Positives = 149/320 (46%), Gaps = 35/320 (10%)
Query: 154 NPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF 213
N L G IP G L++L L L N L+G+IP +GN+ L L L N SG IP+
Sbjct: 152 NNHLNGFIPEEIGYLRSLTKLSLGINFLSGSIPASLGNMTNLSFLFLYENQLSGFIPEEI 211
Query: 214 GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLR 273
G L L L L N LSG++P +LG L ++ L L +N L G + E G L++LT +DL
Sbjct: 212 GYLRSLTKLSLDINFLSGSIPASLGNLNNLSFLYLYNNQLSGSIPEEIGYLRSLTYLDLG 271
Query: 274 NNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI-------RTLKW---------------- 310
N L + SL +N+L + L NN L G I R+L +
Sbjct: 272 ENALNGSIPASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPASL 331
Query: 311 ENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLS 370
NL NL L+L N +L G IPE + L+ L +L L +N + G++ L L +L+ L L
Sbjct: 332 GNLNNLSRLDLYNNKLSGSIPEEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSRLDLY 391
Query: 371 GNNLKGEIQFSKGFFG-----KLGRRFGAWSNP----KLCYPFELMSTNNVPYGVKPCHQ 421
N L G I G+ LG F + S P L F L NN G P
Sbjct: 392 NNKLSGSIPEEIGYLRSLTKLSLGNNFLSGSIPASLGNLNNLFMLYLYNNQLSGSIP--- 448
Query: 422 EEIHLVKSNAKTEVINGDIN 441
EEI + S + N +N
Sbjct: 449 EEIGYLSSLTNLYLGNNSLN 468
Score = 130 bits (327), Expect = 9e-29
Identities = 86/237 (36%), Positives = 123/237 (51%), Gaps = 2/237 (0%)
Query: 153 SNPGLIGNIPS-TFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPD 211
+N +IG + + F L L++L L N ++G IP EIGNL L L L+ N SG IP
Sbjct: 78 TNASVIGTLYAFPFSSLPFLENLDLSNNNISGTIPPEIGNLTNLVYLDLNTNQISGTIPP 137
Query: 212 IFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMD 271
G L+ L I+ + N L+G +P +G L S+ KL L NFL G + GN+ NL+ +
Sbjct: 138 QIGSLAKLQIIRIFNNHLNGFIPEEIGYLRSLTKLSLGINFLSGSIPASLGNMTNLSFLF 197
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
L N+L + + + SL ++ L N L G I NL NL L L N +L G IP
Sbjct: 198 LYENQLSGFIPEEIGYLRSLTKLSLDINFLSGSI-PASLGNLNNLSFLYLYNNQLSGSIP 256
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
E + L+ L +L L +N + G++ L L +L+ L L N L G I G+ L
Sbjct: 257 EEIGYLRSLTYLDLGENALNGSIPASLGNLNNLSRLDLYNNKLSGSIPEEIGYLRSL 313
Score = 70.1 bits (170), Expect = 1e-10
Identities = 78/268 (29%), Positives = 121/268 (45%), Gaps = 25/268 (9%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
NKL ++PT + SL S+ N L IP + K LQ L L +N L P
Sbjct: 585 NKLSGTLPTNF--SIGCSLISLNLHGNE-LEDEIPWSLDNCKKLQVLDLGDNQLNDTFPM 641
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLS--DLLILDLSRNSLSGTLPVTLGRLISVLK 245
+G L +L+ L L+ N G I + DL I+DLSRN+ S LP +L + ++
Sbjct: 642 WLGTLPELRVLRLTSNKLHGPIRSSGAEIMFPDLRIIDLSRNAFSQDLPTSLFEHLKGMR 701
Query: 246 L---------------DLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNS 290
D +G L L T++DL +N+ + L ++ +
Sbjct: 702 TVDKTMEVPSYERYYDDSVVVVTKGLELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIA 761
Query: 291 LEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNI 350
+ + +S+N L G I + +L + L+LS +L GEIP+ L+ L L FL LS N +
Sbjct: 762 IRVLNVSHNALQGYIPS-SLGSLSRVESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYL 820
Query: 351 TGNL--SPKLETLPSLNALYLSGNNLKG 376
G + P+ T S + Y + L+G
Sbjct: 821 QGCIPQGPQFRTFESNS--YEGNDGLRG 846
Score = 62.0 bits (149), Expect = 4e-08
Identities = 35/98 (35%), Positives = 57/98 (57%), Gaps = 1/98 (1%)
Query: 181 LTGNIPQEIGNLVKLKRLV-LSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGR 239
+T + EI ++ L ++ LS N F G+IP + G L + +L++S N+L G +P +LG
Sbjct: 723 VTKGLELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGS 782
Query: 240 LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L V LDLS N L G++ + +L L ++L +N L
Sbjct: 783 LSRVESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYL 820
Score = 61.2 bits (147), Expect = 7e-08
Identities = 34/80 (42%), Positives = 47/80 (58%)
Query: 176 LLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPV 235
L N G+IP +G+L+ ++ L +S N G IP G LS + LDLS N LSG +P
Sbjct: 743 LSSNKFEGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGSLSRVESLDLSFNQLSGEIPQ 802
Query: 236 TLGRLISVLKLDLSHNFLEG 255
L L + L+LSHN+L+G
Sbjct: 803 QLASLTFLEFLNLSHNYLQG 822
Score = 56.2 bits (134), Expect = 2e-06
Identities = 33/90 (36%), Positives = 51/90 (56%)
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
LS ++DLS N G +P LG LI++ L++SHN L+G + + G+L + +DL N
Sbjct: 735 LSLYTVIDLSSNKFEGHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGSLSRVESLDLSFN 794
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+L + L + LE + LS+N L G I
Sbjct: 795 QLSGEIPQQLASLTFLEFLNLSHNYLQGCI 824
Score = 55.1 bits (131), Expect = 5e-06
Identities = 31/76 (40%), Positives = 43/76 (55%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G+IPS G L ++ L + N L G IP +G+L +++ L LS N SG IP L+
Sbjct: 750 GHIPSVLGDLIAIRVLNVSHNALQGYIPSSLGSLSRVESLDLSFNQLSGEIPQQLASLTF 809
Query: 219 LLILDLSRNSLSGTLP 234
L L+LS N L G +P
Sbjct: 810 LEFLNLSHNYLQGCIP 825
Score = 48.5 bits (114), Expect = 4e-04
Identities = 36/132 (27%), Positives = 60/132 (45%), Gaps = 7/132 (5%)
Query: 261 FGNLKNLTLMDLRNNRLCC----GLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNL 316
F N N L + C G+V +N+L ++N + G + + +L L
Sbjct: 41 FKNQNNSFLASWTTSSNACKDWYGVVCLNGRVNTLN---ITNASVIGTLYAFPFSSLPFL 97
Query: 317 VILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKG 376
L+LSN + G IP + L L +L L+ N I+G + P++ +L L + + N+L G
Sbjct: 98 ENLDLSNNNISGTIPPEIGNLTNLVYLDLNTNQISGTIPPQIGSLAKLQIIRIFNNHLNG 157
Query: 377 EIQFSKGFFGKL 388
I G+ L
Sbjct: 158 FIPEEIGYLRSL 169
Score = 46.2 bits (108), Expect = 0.002
Identities = 34/101 (33%), Positives = 55/101 (53%), Gaps = 4/101 (3%)
Query: 111 QLKHLKAISFFNCFQ-SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLK 169
+L+ ++ +S + S NK IP+ + +A + ++ S+ L G IPS+ G L
Sbjct: 728 ELEIVRILSLYTVIDLSSNKFEGHIPSVLGDLIAIRVLNV---SHNALQGYIPSSLGSLS 784
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
++SL L N L+G IPQ++ +L L+ L LS N G IP
Sbjct: 785 RVESLDLSFNQLSGEIPQQLASLTFLEFLNLSHNYLQGCIP 825
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 822,994,895
Number of Sequences: 2540612
Number of extensions: 36272024
Number of successful extensions: 150259
Number of sequences better than 10.0: 4980
Number of HSP's better than 10.0 without gapping: 2713
Number of HSP's successfully gapped in prelim test: 2315
Number of HSP's that attempted gapping in prelim test: 90855
Number of HSP's gapped (non-prelim): 23462
length of query: 475
length of database: 863,360,394
effective HSP length: 132
effective length of query: 343
effective length of database: 527,999,610
effective search space: 181103866230
effective search space used: 181103866230
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146570.9


