

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
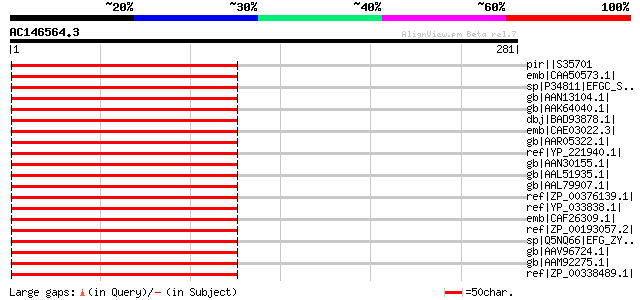
Score E
Sequences producing significant alignments: (bits) Value
pir||S35701 translation elongation factor EF-G, chloroplast - so... 186 8e-46
emb|CAA50573.1| translation elongation factor EF-G [Glycine max] 186 8e-46
sp|P34811|EFGC_SOYBN ELONGATION FACTOR G, CHLOROPLAST PRECURSOR ... 186 8e-46
gb|AAN13104.1| unknown protein [Arabidopsis thaliana] gi|1840765... 180 4e-44
gb|AAK64040.1| unknown protein [Arabidopsis thaliana] 180 4e-44
dbj|BAD93878.1| elongation factor G [Arabidopsis thaliana] 180 4e-44
emb|CAE03022.3| OSJNBa0091D06.15 [Oryza sativa (japonica cultiva... 176 7e-43
gb|AAR05322.1| predicted translation elongation factor G [uncult... 164 3e-39
ref|YP_221940.1| FusA, translation elongation factor G [Brucella... 158 2e-37
gb|AAN30155.1| translation elongation factor G [Brucella suis 13... 158 2e-37
gb|AAL51935.1| Protein Translation Elongation Factor G (EF-G) [B... 158 2e-37
gb|AAL79907.1| elongation factor EfG [Bartonella bacilliformis] 158 2e-37
ref|ZP_00376139.1| translation elongation factor [Erythrobacter ... 158 2e-37
ref|YP_033838.1| Elongation factor g (EF-g) [Bartonella henselae... 157 2e-37
emb|CAF26309.1| Elongation factor g (EF-g) [Bartonella quintana ... 156 5e-37
ref|ZP_00193057.2| COG0480: Translation elongation factors (GTPa... 156 7e-37
sp|Q5NQ66|EFG_ZYMMO Elongation factor G (EF-G) gi|56542985|gb|AA... 155 9e-37
gb|AAV96724.1| translation elongation factor G [Silicibacter pom... 155 1e-36
gb|AAM92275.1| elongation factor G [Rhodobacter capsulatus] 154 2e-36
ref|ZP_00338489.1| COG0480: Translation elongation factors (GTPa... 153 5e-36
>pir||S35701 translation elongation factor EF-G, chloroplast - soybean
Length = 787
Score = 186 bits (471), Expect = 8e-46
Identities = 91/126 (72%), Positives = 108/126 (85%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETLCDP P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 464 DIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVDKMATGLIKLAQ 523
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS+++EV
Sbjct: 524 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKISEV 583
Query: 121 RYVFNK 126
+YV K
Sbjct: 584 KYVHKK 589
Score = 37.4 bits (85), Expect = 0.48
Identities = 16/28 (57%), Positives = 22/28 (78%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L T
Sbjct: 753 MTKGRASYTMQLAMFDVVPQHIQNQLAT 780
>emb|CAA50573.1| translation elongation factor EF-G [Glycine max]
Length = 703
Score = 186 bits (471), Expect = 8e-46
Identities = 91/126 (72%), Positives = 108/126 (85%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETLCDP P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 380 DIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVDKMATGLIKLAQ 439
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS+++EV
Sbjct: 440 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKISEV 499
Query: 121 RYVFNK 126
+YV K
Sbjct: 500 KYVHKK 505
Score = 37.4 bits (85), Expect = 0.48
Identities = 16/28 (57%), Positives = 22/28 (78%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L T
Sbjct: 669 MTKGRASYTMQLAMFDVVPQHIQNQLAT 696
>sp|P34811|EFGC_SOYBN ELONGATION FACTOR G, CHLOROPLAST PRECURSOR (EF-G)
Length = 788
Score = 186 bits (471), Expect = 8e-46
Identities = 91/126 (72%), Positives = 108/126 (85%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETLCDP P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 465 DIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVDKMATGLIKLAQ 524
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS+++EV
Sbjct: 525 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKISEV 584
Query: 121 RYVFNK 126
+YV K
Sbjct: 585 KYVHKK 590
Score = 37.4 bits (85), Expect = 0.48
Identities = 16/28 (57%), Positives = 22/28 (78%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L T
Sbjct: 754 MTKGRASYTMQLAMFDVVPQHIQNQLAT 781
>gb|AAN13104.1| unknown protein [Arabidopsis thaliana] gi|18407650|ref|NP_564801.1|
elongation factor Tu family protein [Arabidopsis
thaliana] gi|6630460|gb|AAF19548.1| F23N19.11
[Arabidopsis thaliana] gi|25299484|pir||E96652 protein
F23N19.11 [imported] - Arabidopsis thaliana
Length = 783
Score = 180 bits (457), Expect = 4e-44
Identities = 89/126 (70%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETL DP+ P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 460 DIIALAGLKDTITGETLSDPENPVVLERMDFPDPVIKVAIEPKTKADIDKMATGLIKLAQ 519
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++ EV
Sbjct: 520 EDPSFHFSRDEEMNQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKIAEV 579
Query: 121 RYVFNK 126
+Y K
Sbjct: 580 KYTHKK 585
Score = 37.4 bits (85), Expect = 0.48
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ +FD V ++IQN+L++
Sbjct: 749 MTKGRASYTMQLAKFDVVPQHIQNQLSS 776
>gb|AAK64040.1| unknown protein [Arabidopsis thaliana]
Length = 783
Score = 180 bits (457), Expect = 4e-44
Identities = 89/126 (70%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETL DP+ P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 460 DIIALAGLKDTITGETLSDPENPVVLERMDFPDPVIKVAIEPKTKADIDKMATGLIKLAQ 519
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++ EV
Sbjct: 520 EDPSFHFSRDEEMNQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKIAEV 579
Query: 121 RYVFNK 126
+Y K
Sbjct: 580 KYTHKK 585
Score = 37.4 bits (85), Expect = 0.48
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ +FD V ++IQN+L++
Sbjct: 749 MTKGRASYTMQLAKFDVVPQHIQNQLSS 776
>dbj|BAD93878.1| elongation factor G [Arabidopsis thaliana]
Length = 410
Score = 180 bits (457), Expect = 4e-44
Identities = 89/126 (70%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETL DP+ P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 87 DIIALAGLKDTITGETLSDPENPVVLERMDFPDPVIKVAIEPKTKADIDKMATGLIKLAQ 146
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++ EV
Sbjct: 147 EDPSFHFSRDEEMNQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKIAEV 206
Query: 121 RYVFNK 126
+Y K
Sbjct: 207 KYTHKK 212
Score = 37.4 bits (85), Expect = 0.48
Identities = 15/28 (53%), Positives = 24/28 (85%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ +FD V ++IQN+L++
Sbjct: 376 MTKGRASYTMQLAKFDVVPQHIQNQLSS 403
>emb|CAE03022.3| OSJNBa0091D06.15 [Oryza sativa (japonica cultivar-group)]
gi|50926886|ref|XP_473337.1| OSJNBa0091D06.15 [Oryza
sativa (japonica cultivar-group)]
Length = 749
Score = 176 bits (446), Expect = 7e-43
Identities = 88/126 (69%), Positives = 106/126 (83%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDTI GETL DP P VL++M+FP VIK+AIEPKTKAD +KM GLIKLA+
Sbjct: 424 DIVALAGLKDTITGETLSDPDKPVVLERMEFPDPVIKVAIEPKTKADADKMATGLIKLAQ 483
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHL+ IVDRLKREF+VEANVGAPQ NYRESIS+++EV
Sbjct: 484 EDPSFHFSRDEETNQTVIEGMGELHLDIIVDRLKREFRVEANVGAPQVNYRESISKISEV 543
Query: 121 RYVFNK 126
+YV K
Sbjct: 544 QYVHKK 549
Score = 38.1 bits (87), Expect = 0.28
Identities = 16/27 (59%), Positives = 23/27 (84%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELT 279
+T+GRASY+MQ +FD V ++IQNEL+
Sbjct: 713 MTKGRASYTMQLAKFDVVPQHIQNELS 739
>gb|AAR05322.1| predicted translation elongation factor G [uncultured marine alpha
proteobacterium HOT2C01]
Length = 691
Score = 164 bits (414), Expect = 3e-39
Identities = 82/126 (65%), Positives = 98/126 (77%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV + GLKDTI GETLCD + P +L+ MDFP VI+IA+EPKTKAD EKM L +LA+
Sbjct: 373 DIVAIAGLKDTITGETLCDSQKPIILESMDFPDPVIEIAVEPKTKADQEKMGEALARLAK 432
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +T+I+G GELHL+ IVDR+KREFKVEANVGAPQ YRE+IS+ EV
Sbjct: 433 EDPSFRVTSDDESGQTVIKGMGELHLDIIVDRMKREFKVEANVGAPQVAYRETISKEVEV 492
Query: 121 RYVFNK 126
Y K
Sbjct: 493 TYTHKK 498
Score = 34.7 bits (78), Expect = 3.1
Identities = 13/26 (50%), Positives = 21/26 (80%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNEL 278
+T+GRA YSM FD ++ V +N+Q+E+
Sbjct: 661 MTQGRAQYSMFFDHYEQVPQNVQDEV 686
>ref|YP_221940.1| FusA, translation elongation factor G [Brucella abortus biovar 1
str. 9-941] gi|62196279|gb|AAX74579.1| FusA, translation
elongation factor G [Brucella abortus biovar 1 str.
9-941]
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>gb|AAN30155.1| translation elongation factor G [Brucella suis 1330]
gi|23502113|ref|NP_698240.1| translation elongation
factor G [Brucella suis 1330]
gi|34395605|sp|Q8G075|EFG_BRUSU Elongation factor G
(EF-G)
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>gb|AAL51935.1| Protein Translation Elongation Factor G (EF-G) [Brucella melitensis
16M] gi|17987037|ref|NP_539671.1| Protein Translation
Elongation Factor G (EF-G) [Brucella melitensis 16M]
gi|25299504|pir||AD3346 protein translation elongation
factor G (EF-G) [imported] - Brucella melitensis (strain
16M) gi|21263518|sp|Q8YHP3|EFG_BRUME Elongation factor G
(EF-G)
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>gb|AAL79907.1| elongation factor EfG [Bartonella bacilliformis]
Length = 694
Score = 158 bits (399), Expect = 2e-37
Identities = 80/126 (63%), Positives = 96/126 (75%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPEPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR++REFKVEANVG PQ YRESI++V E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDIIVDRMRREFKVEANVGQPQVAYRESITKVAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>ref|ZP_00376139.1| translation elongation factor [Erythrobacter litoralis HTCC2594]
gi|60737360|gb|EAL75617.1| translation elongation factor
[Erythrobacter litoralis HTCC2594]
Length = 711
Score = 158 bits (399), Expect = 2e-37
Identities = 77/126 (61%), Positives = 96/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV + G+KDT G+TLCDP P +L++M+FP VI++++EPKTKAD EKM L +LA
Sbjct: 393 DIVAIAGMKDTTTGDTLCDPANPIILERMEFPDPVIELSVEPKTKADQEKMGVALNRLAA 452
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D E +TII+G GELHL+ +VDR+KREFKVEANVGAPQ YRES++ EV
Sbjct: 453 EDPSFRVSTDHESGQTIIKGMGELHLDILVDRMKREFKVEANVGAPQVAYRESLAREVEV 512
Query: 121 RYVFNK 126
Y K
Sbjct: 513 TYTHKK 518
>ref|YP_033838.1| Elongation factor g (EF-g) [Bartonella henselae str. Houston-1]
gi|49238604|emb|CAF27845.1| Elongation factor g (EF-g)
[Bartonella henselae str. Houston-1]
gi|22203346|gb|AAM92279.1| elongation factor G
[Bartonella henselae] gi|62286487|sp|Q8KQB3|EFG_BARHE
Elongation factor G (EF-G)
Length = 694
Score = 157 bits (398), Expect = 2e-37
Identities = 79/126 (62%), Positives = 96/126 (75%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPEPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR++REFKVEANVG PQ YRESI+++ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDIIVDRMRREFKVEANVGQPQVAYRESITKIAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>emb|CAF26309.1| Elongation factor g (EF-g) [Bartonella quintana str. Toulouse]
gi|49474407|ref|YP_032449.1| Elongation factor g (EF-g)
[Bartonella quintana str. Toulouse]
gi|62286478|sp|Q6FZB9|EFG_BARQU Elongation factor G
(EF-G)
Length = 694
Score = 156 bits (395), Expect = 5e-37
Identities = 79/126 (62%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P VL++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLRPVVLERMEFPEPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR++REFKVEAN+G PQ YRESI++ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDIIVDRMRREFKVEANIGQPQVAYRESITKAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>ref|ZP_00193057.2| COG0480: Translation elongation factors (GTPases) [Mesorhizobium
sp. BNC1]
Length = 696
Score = 156 bits (394), Expect = 7e-37
Identities = 78/126 (61%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTK D EKM L +LA
Sbjct: 378 DIVALAGLKETTTGDTLCDPLKPVILERMEFPEPVIQIAIEPKTKGDQEKMGLALNRLAA 437
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR+KREFKVEAN+GAPQ YRE+I++ E+
Sbjct: 438 EDPSFRVKTDEESGQTIIAGMGELHLDIIVDRMKREFKVEANIGAPQVAYRETITKTAEI 497
Query: 121 RYVFNK 126
Y K
Sbjct: 498 DYTHKK 503
>sp|Q5NQ66|EFG_ZYMMO Elongation factor G (EF-G) gi|56542985|gb|AAV89139.1| translation
elongation factor [Zymomonas mobilis subsp. mobilis ZM4]
gi|56551411|ref|YP_162250.1| translation elongation
factor [Zymomonas mobilis subsp. mobilis ZM4]
Length = 690
Score = 155 bits (393), Expect = 9e-37
Identities = 77/126 (61%), Positives = 96/126 (76%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV LVG+K+T G+TLC P P +L++M+FP VI++A+EPKTKAD EKM L +LA
Sbjct: 372 DIVALVGMKETTTGDTLCAPNAPIILERMEFPEPVIEVAVEPKTKADQEKMGLALNRLAA 431
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV+ D E +TII+G GELHL+ +VDR+KREFKVEANVGAPQ YRES++ EV
Sbjct: 432 EDPSFRVASDFESGQTIIKGMGELHLDILVDRMKREFKVEANVGAPQVAYRESLARPVEV 491
Query: 121 RYVFNK 126
Y K
Sbjct: 492 DYTHKK 497
Score = 34.3 bits (77), Expect = 4.1
Identities = 14/25 (56%), Positives = 18/25 (72%)
Query: 254 TEGRASYSMQFDRFDTVSRNIQNEL 278
T+GRA YSMQF +D V N+ +EL
Sbjct: 661 TQGRAQYSMQFSHYDEVPANVADEL 685
>gb|AAV96724.1| translation elongation factor G [Silicibacter pomeroyi DSS-3]
gi|56698321|ref|YP_168694.1| translation elongation
factor G [Silicibacter pomeroyi DSS-3]
gi|62286649|sp|Q5LMR4|EFG_SILPO Elongation factor G
(EF-G)
Length = 705
Score = 155 bits (392), Expect = 1e-36
Identities = 79/126 (62%), Positives = 93/126 (73%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDT G+TLCD K P VL+ M FP VI+IA+EPKTK D EKM GL +LA
Sbjct: 387 DIIALAGLKDTTTGDTLCDAKDPVVLETMTFPDPVIEIAVEPKTKGDQEKMSQGLARLAA 446
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D E +TI++G GELHL+ +VDRLKREFKVEAN+GAPQ YRE+IS+ E
Sbjct: 447 EDPSFRVETDIESGQTIMKGMGELHLDILVDRLKREFKVEANIGAPQVAYRETISKEVEH 506
Query: 121 RYVFNK 126
Y K
Sbjct: 507 TYTHKK 512
>gb|AAM92275.1| elongation factor G [Rhodobacter capsulatus]
Length = 708
Score = 154 bits (390), Expect = 2e-36
Identities = 77/126 (61%), Positives = 94/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLK+T G+TLCDP P VL+ M FPV VI+IA+EPKTKAD EKM L +LA
Sbjct: 390 DIIALGGLKETTTGDTLCDPSAPVVLETMTFPVPVIEIAVEPKTKADQEKMGIALARLAA 449
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D E +TI++G GELHL+ +VDR++REFKVEAN+GAPQ YRE+IS E+
Sbjct: 450 EDPSFRVETDVESGQTIMKGMGELHLDILVDRMRREFKVEANIGAPQVAYRETISREHEI 509
Query: 121 RYVFNK 126
Y K
Sbjct: 510 DYTHKK 515
>ref|ZP_00338489.1| COG0480: Translation elongation factors (GTPases) [Silicibacter sp.
TM1040]
Length = 706
Score = 153 bits (387), Expect = 5e-36
Identities = 78/126 (61%), Positives = 91/126 (71%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDT G+TLCD K P VL+ M FP VI+IA+EPKTK D EKM GL +LA
Sbjct: 388 DIIALAGLKDTTTGDTLCDAKEPVVLETMTFPDPVIEIAVEPKTKGDQEKMSQGLARLAA 447
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D E +TI++G GELHL+ +VDRLKREFKVEAN+GAPQ YRE+I E
Sbjct: 448 EDPSFRVETDLESGQTIMKGMGELHLDILVDRLKREFKVEANIGAPQVAYRETIGHEVEH 507
Query: 121 RYVFNK 126
Y K
Sbjct: 508 TYTHKK 513
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 433,190,669
Number of Sequences: 2540612
Number of extensions: 16440183
Number of successful extensions: 57673
Number of sequences better than 10.0: 1359
Number of HSP's better than 10.0 without gapping: 921
Number of HSP's successfully gapped in prelim test: 438
Number of HSP's that attempted gapping in prelim test: 55326
Number of HSP's gapped (non-prelim): 1882
length of query: 281
length of database: 863,360,394
effective HSP length: 126
effective length of query: 155
effective length of database: 543,243,282
effective search space: 84202708710
effective search space used: 84202708710
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 74 (33.1 bits)
Medicago: description of AC146564.3


