

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146552.5 - phase: 0 /pseudo
(43 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
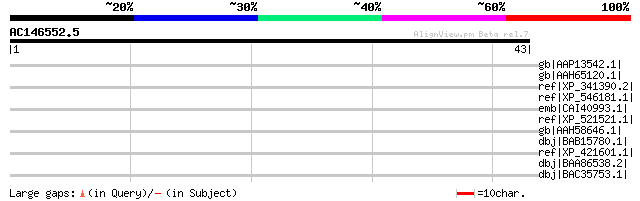
gb|AAP13542.1| PIAS-like protein hZimp10 [Homo sapiens] gi|56404... 32 3.6
gb|AAH65120.1| Retinoic acid induced 17 [Mus musculus] gi|410538... 32 3.6
ref|XP_341390.2| PREDICTED: similar to Rai17 protein [Rattus nor... 32 3.6
ref|XP_546181.1| PREDICTED: similar to Rai17 protein [Canis fami... 32 3.6
emb|CAI40993.1| retinoic acid induced 17 [Homo sapiens] 32 3.6
ref|XP_521521.1| PREDICTED: similar to retinoic acid induced 17;... 32 3.6
gb|AAH58646.1| Rai17 protein [Mus musculus] 32 3.6
dbj|BAB15780.1| FLJ00092 protein [Homo sapiens] 32 3.6
ref|XP_421601.1| PREDICTED: similar to Hypothetical protein E330... 32 3.6
dbj|BAA86538.2| KIAA1224 protein [Homo sapiens] 32 3.6
dbj|BAC35753.1| unnamed protein product [Mus musculus] 32 3.6
>gb|AAP13542.1| PIAS-like protein hZimp10 [Homo sapiens]
gi|56404979|sp|Q9ULJ6|ZIM10_HUMAN Retinoic acid-induced
protein 17 (PIAS-like protein Zimp10)
gi|31543543|ref|NP_065071.1| retinoic acid induced 17
[Homo sapiens]
Length = 1067
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 264 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 301
>gb|AAH65120.1| Retinoic acid induced 17 [Mus musculus]
gi|41053864|ref|NP_899031.2| retinoic acid induced 17
[Mus musculus] gi|56404800|sp|Q6P1E1|ZIM10_MOUSE
Retinoic acid-induced protein 17 (PIAS-like protein
Zimp10)
Length = 1072
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 264 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 301
>ref|XP_341390.2| PREDICTED: similar to Rai17 protein [Rattus norvegicus]
Length = 1066
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 264 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 301
>ref|XP_546181.1| PREDICTED: similar to Rai17 protein [Canis familiaris]
Length = 1842
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 679 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 716
>emb|CAI40993.1| retinoic acid induced 17 [Homo sapiens]
Length = 1064
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 261 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 298
>ref|XP_521521.1| PREDICTED: similar to retinoic acid induced 17; zinc
finger-containing, Miz1, PIAS-like protein on chromosome
10; PIAS-like protein hZimp10 [Pan troglodytes]
Length = 2372
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 1600 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 1637
>gb|AAH58646.1| Rai17 protein [Mus musculus]
Length = 1066
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 264 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 301
>dbj|BAB15780.1| FLJ00092 protein [Homo sapiens]
Length = 369
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 174 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 211
>ref|XP_421601.1| PREDICTED: similar to Hypothetical protein E330020C23 [Gallus gallus]
Length = 2188
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 1408 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 1445
>dbj|BAA86538.2| KIAA1224 protein [Homo sapiens]
Length = 997
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 194 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 231
>dbj|BAC35753.1| unnamed protein product [Mus musculus]
Length = 351
Score = 32.3 bits (72), Expect = 3.6
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 1/39 (2%)
Query: 5 MALPSHRRDHKGDATMPAAASAVASWAALEAVADACSTA 43
M +P H R D T PAAA+A A+ AA A A A +TA
Sbjct: 264 MGIPPHTRP-PADFTQPAAAAAAAAVAAAAATATATATA 301
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.116 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,397,039
Number of Sequences: 2540612
Number of extensions: 1219925
Number of successful extensions: 4376
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 4376
Number of HSP's gapped (non-prelim): 11
length of query: 43
length of database: 863,360,394
effective HSP length: 19
effective length of query: 24
effective length of database: 815,088,766
effective search space: 19562130384
effective search space used: 19562130384
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146552.5


