

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146342.8 - phase: 0
(337 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
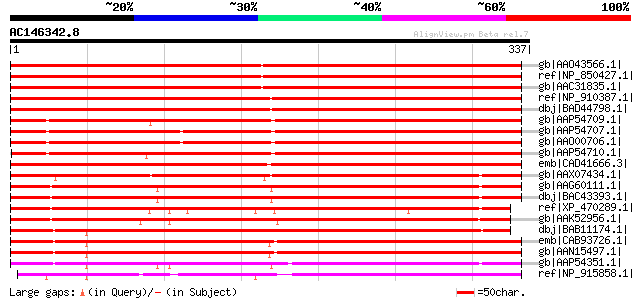
Score E
Sequences producing significant alignments: (bits) Value
gb|AAO43566.1| At2g45510 [Arabidopsis thaliana] gi|2979544|gb|AA... 446 e-124
ref|NP_850427.1| cytochrome P450 family protein [Arabidopsis tha... 446 e-124
gb|AAC31835.1| putative cytochrome P450 [Arabidopsis thaliana] g... 446 e-124
ref|NP_910387.1| ESTs AU056036(S20239),C72753(E2173), AU056035(S... 409 e-113
dbj|BAD44798.1| putative cytochrome P450 [Oryza sativa (japonica... 409 e-113
gb|AAP54709.1| cytochrome P450-like protein [Oryza sativa (japon... 359 6e-98
gb|AAP54707.1| cytochrome P450-like protein [Oryza sativa (japon... 353 5e-96
gb|AAO00706.1| putative cytochrome P450-dependent fatty acid hyd... 353 5e-96
gb|AAP54710.1| cytochrome P450-like protein [Oryza sativa (japon... 352 7e-96
emb|CAD41666.3| OSJNBa0019K04.13 [Oryza sativa (japonica cultiva... 348 2e-94
gb|AAX07434.1| cytochrome P450 CYPD [Pinus taeda] 344 2e-93
gb|AAG60111.1| cytochrome P450, putative [Arabidopsis thaliana] 296 8e-79
dbj|BAC43393.1| unknown protein [Arabidopsis thaliana] gi|306978... 296 8e-79
ref|XP_470289.1| putative plant cytochrome P-450 protein [Oryza ... 271 3e-71
gb|AAK52956.1| cytochrome P450-like protein [Zea mays] 268 2e-70
dbj|BAB11174.1| cytochrome P450-like protein [Arabidopsis thalia... 262 9e-69
emb|CAB93726.1| cytochrome P450-like protein [Arabidopsis thalia... 261 3e-68
gb|AAN15497.1| cytochrome P450-like protein [Arabidopsis thalian... 261 3e-68
gb|AAP54351.1| putative cytochrome P450 protein [Oryza sativa (j... 239 1e-61
ref|NP_915858.1| cytochrome P450-like protein [Oryza sativa (jap... 225 1e-57
>gb|AAO43566.1| At2g45510 [Arabidopsis thaliana] gi|2979544|gb|AAC06153.1| putative
cytochrome P450 [Arabidopsis thaliana]
gi|15225499|ref|NP_182075.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|7430723|pir||T00864 cytochrome
P450 homolog F17K2.4 - Arabidopsis thaliana
Length = 511
Score = 446 bits (1147), Expect = e-124
Identities = 216/332 (65%), Positives = 265/332 (79%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+CTLDSIFKVGFGVEL CL+ SKE FM+AF++ N R++DPLWKLK N GS
Sbjct: 178 MRCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGS 237
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
++ LKK+I ID FV+SLI T+R+ L+ +++ +EDILSRFL+ES+KD NM DKYLRD
Sbjct: 238 QSKLKKSIATIDKFVYSLITTKRKELAKEQNTVVREDILSRFLVESEKDPENMNDKYLRD 297
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILNFMIAGKDT+A LSWF YMLCKNPL+QEK+ QE+ +VT S E +++ FV +I+
Sbjct: 298 IILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTF-SHEKTTDVNGFVESIN 356
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ LD+MHYLHAAL+ETLRLYP VP+ R AE D+LPDG+ V+KG+ +YY++YAMGRM
Sbjct: 357 EEALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRMT 416
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
YIWG DA+EF PERWLKDG+FQPES FKFISFHAGPRICLGKDFAYRQMKIVSMAL+ FF
Sbjct: 417 YIWGQDAEEFKPERWLKDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLHFF 476
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
RFK+ +E + V Y+ M TLH+D GL L A PR
Sbjct: 477 RFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>ref|NP_850427.1| cytochrome P450 family protein [Arabidopsis thaliana]
Length = 505
Score = 446 bits (1146), Expect = e-124
Identities = 216/332 (65%), Positives = 264/332 (79%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
MKCTLDSIFKVGFGVEL CL+ SKE FMKAF++ N R DP WKLK LN GS
Sbjct: 172 MKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGS 231
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
E+ LKK+I IID FV+SLI T+R+ LS +++ S +EDILS+FLLES+KD NM DKYLRD
Sbjct: 232 ESRLKKSIAIIDKFVYSLITTKRKELSKEQNTSVREDILSKFLLESEKDPENMNDKYLRD 291
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILN M+AGKDT+A +LSWF YMLCKNPL+QEK+ QE+ +VTS S E +++ F+ +++
Sbjct: 292 IILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTS-SHEKTTDVNGFIESVT 350
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ L +M YLHAAL+ET+RLYP VP R AE D+LPDG+ V+KG+ +YY+SYAMGRM
Sbjct: 351 EEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRMT 410
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
YIWG DA+EF PERWLKDG+FQPES FKFISFHAGPRIC+GKDFAYRQMKIVSMAL+ FF
Sbjct: 411 YIWGQDAEEFKPERWLKDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMALLHFF 470
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
RFK+ +E + V+Y+ M TLH+D GL L A PR
Sbjct: 471 RFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 502
>gb|AAC31835.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|7430726|pir||T00404 probable cytochrome P450
At2g44890 [imported] - Arabidopsis thaliana
Length = 490
Score = 446 bits (1146), Expect = e-124
Identities = 216/332 (65%), Positives = 264/332 (79%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
MKCTLDSIFKVGFGVEL CL+ SKE FMKAF++ N R DP WKLK LN GS
Sbjct: 157 MKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGS 216
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
E+ LKK+I IID FV+SLI T+R+ LS +++ S +EDILS+FLLES+KD NM DKYLRD
Sbjct: 217 ESRLKKSIAIIDKFVYSLITTKRKELSKEQNTSVREDILSKFLLESEKDPENMNDKYLRD 276
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILN M+AGKDT+A +LSWF YMLCKNPL+QEK+ QE+ +VTS S E +++ F+ +++
Sbjct: 277 IILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTS-SHEKTTDVNGFIESVT 335
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ L +M YLHAAL+ET+RLYP VP R AE D+LPDG+ V+KG+ +YY+SYAMGRM
Sbjct: 336 EEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRMT 395
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
YIWG DA+EF PERWLKDG+FQPES FKFISFHAGPRIC+GKDFAYRQMKIVSMAL+ FF
Sbjct: 396 YIWGQDAEEFKPERWLKDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMALLHFF 455
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
RFK+ +E + V+Y+ M TLH+D GL L A PR
Sbjct: 456 RFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>ref|NP_910387.1| ESTs AU056036(S20239),C72753(E2173), AU056035(S20239) correspond to
a region of the predicted gene.~Similar to putative
cytochrome P-450 (AC003680) [Oryza sativa (japonica
cultivar-group)]
Length = 403
Score = 409 bits (1052), Expect = e-113
Identities = 190/332 (57%), Positives = 255/332 (76%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ T+DSIFKVGFG ELN L S + F KAF+++N+ V+ R++D +WKLKR LN GS
Sbjct: 69 MRATMDSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKLKRYLNIGS 128
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
EA LK+NI+IID FV LI +RE + + D+ KEDILSRF+L S++D M D+YLRD
Sbjct: 129 EAKLKRNIQIIDSFVMKLIHQKREQMKIAADYKTKEDILSRFVLASEQDPGTMDDRYLRD 188
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
I+LNF+IAGKDT+ NTL+WFFY+LCKNP++Q+KVA E+ S+E N ++ F +
Sbjct: 189 IVLNFLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDN-TIESFTKRLD 247
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ + KMHYL A ++ETLRLYPAVP+ + A+E D+LP+GY V KG+ + Y+ YAMGRM
Sbjct: 248 EGAISKMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYMIYAMGRMT 307
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
Y+WG+DAQEF PERWL +G++Q ES FKF+SF+AGPRICLGK+FA+RQMKI++ L+ FF
Sbjct: 308 YLWGEDAQEFRPERWLVNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKIMAATLIHFF 367
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
+F+LE+E+ + Y+TMFTLHID+GL L A PR
Sbjct: 368 KFRLEDESKEPIYKTMFTLHIDNGLHLLANPR 399
>dbj|BAD44798.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 525
Score = 409 bits (1052), Expect = e-113
Identities = 190/332 (57%), Positives = 255/332 (76%), Gaps = 1/332 (0%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ T+DSIFKVGFG ELN L S + F KAF+++N+ V+ R++D +WKLKR LN GS
Sbjct: 191 MRATMDSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKLKRYLNIGS 250
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
EA LK+NI+IID FV LI +RE + + D+ KEDILSRF+L S++D M D+YLRD
Sbjct: 251 EAKLKRNIQIIDSFVMKLIHQKREQMKIAADYKTKEDILSRFVLASEQDPGTMDDRYLRD 310
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
I+LNF+IAGKDT+ NTL+WFFY+LCKNP++Q+KVA E+ S+E N ++ F +
Sbjct: 311 IVLNFLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDN-TIESFTKRLD 369
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
+ + KMHYL A ++ETLRLYPAVP+ + A+E D+LP+GY V KG+ + Y+ YAMGRM
Sbjct: 370 EGAISKMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYMIYAMGRMT 429
Query: 241 YIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRFF 300
Y+WG+DAQEF PERWL +G++Q ES FKF+SF+AGPRICLGK+FA+RQMKI++ L+ FF
Sbjct: 430 YLWGEDAQEFRPERWLVNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKIMAATLIHFF 489
Query: 301 RFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
+F+LE+E+ + Y+TMFTLHID+GL L A PR
Sbjct: 490 KFRLEDESKEPIYKTMFTLHIDNGLHLLANPR 521
>gb|AAP54709.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536240|ref|NP_922422.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146747|gb|AAM12483.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 515
Score = 359 bits (922), Expect = 6e-98
Identities = 177/335 (52%), Positives = 239/335 (70%), Gaps = 6/335 (1%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ T+DSIF + FG +LN L+ S E F AF+D++ F RYL+P WKL R+LN G+
Sbjct: 184 MRATMDSIFTIAFGQDLNTLDGSG-EGRRFAAAFDDASEFTMLRYLNPFWKLSRLLNVGA 242
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQK--DFSDKEDILSRFLLESKKDSSNMTDKYL 118
EA LK+ IK++D FV+ LI+ R + LS K D ++DIL+RF+ + DS + KYL
Sbjct: 243 EAMLKERIKVVDGFVYKLIRDRSDELSNTKAHDTDSRQDILTRFIQATTSDSGTVDYKYL 302
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSN 178
RDIILN +IAGKDT+A +L+WF YM+CK+P +QEK+ E + T+ + +++ DEF +
Sbjct: 303 RDIILNIVIAGKDTTAGSLAWFLYMMCKHPEVQEKICHEAMEATNAGEAASI--DEFSQS 360
Query: 179 ISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGR 238
++D L+KMHYLHAALTETLRLYPAVP+ + D+LP+G+ V+KG+ V+Y+ YAMGR
Sbjct: 361 LTDEALNKMHYLHAALTETLRLYPAVPLDNKQCFSDDVLPNGFNVSKGDIVFYIPYAMGR 420
Query: 239 MPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
M +WG DA+ F PERWL ++G+FQ ES FKF +F AGPRICLGKDFAYRQMKI + L+
Sbjct: 421 MESLWGKDAESFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKDFAYRQMKIFAAVLL 480
Query: 298 RFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
RFF KL +E ++YRTM TL +D GL L A R
Sbjct: 481 RFFVLKLRDEKEIISYRTMITLSVDQGLHLTAMAR 515
>gb|AAP54707.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536236|ref|NP_922420.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146758|gb|AAM12494.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 495
Score = 353 bits (905), Expect = 5e-96
Identities = 177/333 (53%), Positives = 237/333 (71%), Gaps = 5/333 (1%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
++ T+DSIF + FG +LN L+ S E + F AF+D++ F RY+ PLWKL R+LN G
Sbjct: 167 LRATMDSIFTIAFGTDLNTLDGSG-EGSRFAAAFDDASEFTMLRYISPLWKLARLLNVGV 225
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
EA LK+ IK++D+FV+ LI+ R + LS D ++DILSRFL + DS + KYLRD
Sbjct: 226 EAMLKERIKVVDEFVYRLIRARSDELSNSHDSGSRQDILSRFLQATTSDSV-VDYKYLRD 284
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILN +IAGKDT+A L+WF YM+CK+P +QEK+ E + TS +++ DEF+ +++
Sbjct: 285 IILNIVIAGKDTTAGALAWFLYMVCKHPEVQEKICHEAMVATSAGDTASV--DEFLQSLT 342
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
D L+ MHYLHAALTETLRLYP+VPM + D+LP+G+ V+KG+ V+++ YAMGRM
Sbjct: 343 DQALNNMHYLHAALTETLRLYPSVPMENKQCFSDDVLPNGFNVSKGDIVFFIPYAMGRME 402
Query: 241 YIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRF 299
+WG DA+ F PERWL ++G+FQ ES FKF +F AGPRICLGK+FAYRQMKI + L+RF
Sbjct: 403 SLWGKDAEYFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKEFAYRQMKIFAAVLLRF 462
Query: 300 FRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
F KL +E V+YRT TL ID GL L AT R
Sbjct: 463 FVLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 495
>gb|AAO00706.1| putative cytochrome P450-dependent fatty acid hydroxylase,
5'-partial [Oryza sativa (japonica cultivar-group)]
Length = 404
Score = 353 bits (905), Expect = 5e-96
Identities = 177/333 (53%), Positives = 237/333 (71%), Gaps = 5/333 (1%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
++ T+DSIF + FG +LN L+ S E + F AF+D++ F RY+ PLWKL R+LN G
Sbjct: 76 LRATMDSIFTIAFGTDLNTLDGSG-EGSRFAAAFDDASEFTMLRYISPLWKLARLLNVGV 134
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLRD 120
EA LK+ IK++D+FV+ LI+ R + LS D ++DILSRFL + DS + KYLRD
Sbjct: 135 EAMLKERIKVVDEFVYRLIRARSDELSNSHDSGSRQDILSRFLQATTSDSV-VDYKYLRD 193
Query: 121 IILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNIS 180
IILN +IAGKDT+A L+WF YM+CK+P +QEK+ E + TS +++ DEF+ +++
Sbjct: 194 IILNIVIAGKDTTAGALAWFLYMVCKHPEVQEKICHEAMVATSAGDTASV--DEFLQSLT 251
Query: 181 DATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRMP 240
D L+ MHYLHAALTETLRLYP+VPM + D+LP+G+ V+KG+ V+++ YAMGRM
Sbjct: 252 DQALNNMHYLHAALTETLRLYPSVPMENKQCFSDDVLPNGFNVSKGDIVFFIPYAMGRME 311
Query: 241 YIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVRF 299
+WG DA+ F PERWL ++G+FQ ES FKF +F AGPRICLGK+FAYRQMKI + L+RF
Sbjct: 312 SLWGKDAEYFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKEFAYRQMKIFAAVLLRF 371
Query: 300 FRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
F KL +E V+YRT TL ID GL L AT R
Sbjct: 372 FVLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 404
>gb|AAP54710.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536242|ref|NP_922423.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146744|gb|AAM12480.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 516
Score = 352 bits (904), Expect = 7e-96
Identities = 171/334 (51%), Positives = 241/334 (71%), Gaps = 6/334 (1%)
Query: 2 KCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGSE 61
+ T+DSIF + FG +LN L+ S E F KAF+D+ ++ RYL+P WKL R+LN G+E
Sbjct: 186 RATMDSIFTIAFGQDLNTLDGSG-EGRHFAKAFDDAGEYLLLRYLNPFWKLARLLNVGAE 244
Query: 62 AALKKNIKIIDDFVHSLIKTRRELLS--MQKDFSDKEDILSRFLLESKKDSSNMTDKYLR 119
A LK+ IK++D+FV+ LI+ R + LS M +D ++D+LSRF+ + DS + KYLR
Sbjct: 245 ATLKERIKVVDEFVYKLIRARSDELSNTMAQDHRSRDDLLSRFIQATTSDSGTVDYKYLR 304
Query: 120 DIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNI 179
DI+LN +IA KD+++ +L+WF YM CK P +QEK+ EV+ T+ +++ DEF++++
Sbjct: 305 DIVLNIVIAAKDSTSGSLAWFLYMACKRPEVQEKIFDEVMETTNAGDCASI--DEFLTSL 362
Query: 180 SDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRM 239
+D L+KMHYLHAALTETLRLYP+VP+ + D+LP+G+ V+KG+ V+Y+ YAMGRM
Sbjct: 363 TDQALNKMHYLHAALTETLRLYPSVPLENKQCFSDDVLPNGFSVSKGDGVFYMPYAMGRM 422
Query: 240 PYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVR 298
++WG DA+ F PERWL + G+FQ ES FKF +F AGPRIC+GKDFAYRQMKI + L+R
Sbjct: 423 EFLWGKDAEAFRPERWLDEHGVFQQESPFKFTAFQAGPRICIGKDFAYRQMKIFAAVLIR 482
Query: 299 FFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
F FKL ++ ++V+YRT TL ID L L AT R
Sbjct: 483 SFVFKLRDKKDNVSYRTAITLAIDQDLHLTATAR 516
>emb|CAD41666.3| OSJNBa0019K04.13 [Oryza sativa (japonica cultivar-group)]
gi|50928103|ref|XP_473579.1| OSJNBa0019K04.13 [Oryza
sativa (japonica cultivar-group)]
Length = 506
Score = 348 bits (892), Expect = 2e-94
Identities = 178/334 (53%), Positives = 238/334 (70%), Gaps = 4/334 (1%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ TLDS F+VGFGV L L SSKE +F +AF+D++ V R+ D LWK+KR LN S
Sbjct: 175 MRATLDSFFRVGFGVNLGVLSGSSKEGLVFARAFDDASEQVLFRFFDLLWKVKRFLNISS 234
Query: 61 EAALKKNIKIIDDFVHSLIKTRRELLSM-QKDFSDKEDILSRFLLESKKDSSNMTDKYLR 119
EA +K++I+ I+DFV+S+I + E +S Q +F+ KEDILSRFLLE +KD +KY+R
Sbjct: 235 EATMKQSIRTINDFVYSIIDRKIEQMSREQHEFAKKEDILSRFLLEREKDPGCFDNKYIR 294
Query: 120 DIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQESNLNLDEFVSNI 179
DIILNF+IAG+DT+A TLSWF Y +CKN +Q+K+A+EV + T+ ++ + + +F S +
Sbjct: 295 DIILNFVIAGRDTTAGTLSWFLYAVCKNQRVQDKIAREVRDATTGDRD--VGVQDFSSFL 352
Query: 180 SDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMGRM 239
++ ++KM YLHAALTETLRLYP VP+ + D LPDG+ V KG+ V Y Y MGRM
Sbjct: 353 TEDAINKMQYLHAALTETLRLYPGVPLDVKYCFSDDTLPDGHAVKKGDMVNYQPYPMGRM 412
Query: 240 PYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALVR 298
++WGD+A+EF PERWL D G+F ES FKF +F AGPRICLGK+FAYRQMKIVS L+
Sbjct: 413 KFLWGDNAEEFKPERWLDDSGMFVAESPFKFTAFQAGPRICLGKEFAYRQMKIVSAVLLY 472
Query: 299 FFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
FFRF++ ++ V YR M TL +D L A R
Sbjct: 473 FFRFEMWDDDATVGYRPMLTLKMDGPFYLRALAR 506
>gb|AAX07434.1| cytochrome P450 CYPD [Pinus taeda]
Length = 518
Score = 344 bits (882), Expect = 2e-93
Identities = 180/344 (52%), Positives = 248/344 (71%), Gaps = 15/344 (4%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEAN---IFMKAFNDSNAFVFKRYL-DPLWKLKRVL 56
M+ +LDSI KV FG+++N L S E+ F KAF+ +NA VF R++ WK++R
Sbjct: 174 MRSSLDSICKVVFGIDINSLSSSKAESGPEASFAKAFDVANAMVFHRHMVGSFWKVQRFF 233
Query: 57 NFGSEAALKKNIKIIDDFVHSLIKTRR-ELLSMQKDFSDKEDILSRFLLESKKDSSN-MT 114
N GSEA L+ NIK++DDF++ +I RR E+ S +K+ + + DILSR+++ S K++ ++
Sbjct: 234 NVGSEAILRDNIKMVDDFLYKVIHFRRQEMFSAEKE-NVRPDILSRYIIISDKETDGKVS 292
Query: 115 DKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTST-----SQESN 169
DKYLRD+ILNFM+A +DT+A LSWF YMLCK+ +QEK+ +E+I+ TS S E N
Sbjct: 293 DKYLRDVILNFMVAARDTTAIALSWFIYMLCKHQHVQEKLLEEIISSTSVHEDQYSTECN 352
Query: 170 LNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETV 229
++ F +++D L KMHYLHA+L+ETLRLYPA+P+ G+ D LPDG+ V KG++V
Sbjct: 353 -DIASFAQSLTDEALGKMHYLHASLSETLRLYPALPVDGKYVVNEDTLPDGFKVKKGDSV 411
Query: 230 YYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQM 289
+L YAMGRM Y+WGDDA+EF PERW++DGIF P+S FKF +F AGPR CLGKDFAY QM
Sbjct: 412 NFLPYAMGRMSYLWGDDAKEFKPERWIQDGIFHPKSPFKFPAFQAGPRTCLGKDFAYLQM 471
Query: 290 KIVSMALVRFFRFKLENETNDVTYRTMFTLHI-DHGLPLYATPR 332
KIV+ LVRFF+F+ +T +V YRTM TLH+ + GL + TPR
Sbjct: 472 KIVAAVLVRFFKFEAV-KTKEVRYRTMLTLHMNEDGLNVQVTPR 514
>gb|AAG60111.1| cytochrome P450, putative [Arabidopsis thaliana]
Length = 524
Score = 296 bits (757), Expect = 8e-79
Identities = 168/358 (46%), Positives = 219/358 (60%), Gaps = 28/358 (7%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ TLDSI KVGFGVE+ L E N F KAF+ +N V R++DPLWK+K+ LN GS
Sbjct: 168 MRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIVTLRFIDPLWKMKKFLNIGS 226
Query: 61 EAALKKNIKIIDDFVHSLIKTRR-ELLSMQKDFSD---------KEDILSRFLLESKKDS 110
EA L K+IK+++DF +S+I+ R+ ELL Q ++ K DILSRF+ S
Sbjct: 227 EALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDPD 286
Query: 111 SNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQES-- 168
S T+K LRDI+LNF+IAG+DT+A TL+W YM+ N + EK+ E+ + S E+
Sbjct: 287 SKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEATN 346
Query: 169 --------------NLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEH 214
N + EF ++ +L K+HYLHA +TETLRLYPAVP + E
Sbjct: 347 TSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLED 406
Query: 215 DILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHA 274
D+LP+G V G V Y+ Y+MGRM Y WG DA F PERWLKDG+FQ S FKF +F A
Sbjct: 407 DMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWLKDGVFQNASPFKFTAFQA 466
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
GPRICLGKD AY QMK+ L RF++F L + V YR M L + HGL + + R
Sbjct: 467 GPRICLGKDSAYLQMKMAMAILCRFYKFHLV-PNHPVKYRMMTILSMAHGLKVTVSRR 523
>dbj|BAC43393.1| unknown protein [Arabidopsis thaliana] gi|30697865|ref|NP_177109.2|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 478
Score = 296 bits (757), Expect = 8e-79
Identities = 168/358 (46%), Positives = 219/358 (60%), Gaps = 28/358 (7%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ TLDSI KVGFGVE+ L E N F KAF+ +N V R++DPLWK+K+ LN GS
Sbjct: 122 MRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIVTLRFIDPLWKMKKFLNIGS 180
Query: 61 EAALKKNIKIIDDFVHSLIKTRR-ELLSMQKDFSD---------KEDILSRFLLESKKDS 110
EA L K+IK+++DF +S+I+ R+ ELL Q ++ K DILSRF+ S
Sbjct: 181 EALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDPD 240
Query: 111 SNMTDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQES-- 168
S T+K LRDI+LNF+IAG+DT+A TL+W YM+ N + EK+ E+ + S E+
Sbjct: 241 SKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEATN 300
Query: 169 --------------NLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEH 214
N + EF ++ +L K+HYLHA +TETLRLYPAVP + E
Sbjct: 301 TSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLED 360
Query: 215 DILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHA 274
D+LP+G V G V Y+ Y+MGRM Y WG DA F PERWLKDG+FQ S FKF +F A
Sbjct: 361 DMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWLKDGVFQNASPFKFTAFQA 420
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
GPRICLGKD AY QMK+ L RF++F L + V YR M L + HGL + + R
Sbjct: 421 GPRICLGKDSAYLQMKMAMAILCRFYKFHLV-PNHPVKYRMMTILSMAHGLKVTVSRR 477
>ref|XP_470289.1| putative plant cytochrome P-450 protein [Oryza sativa (japonica
cultivar-group)] gi|19071651|gb|AAL84318.1| putative
plant cytochrome P-450 protein [Oryza sativa (japonica
cultivar-group)]
Length = 544
Score = 271 bits (692), Expect = 3e-71
Identities = 160/350 (45%), Positives = 216/350 (61%), Gaps = 27/350 (7%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ TLDSI KVGFGVE+ L E N F +AF+ +N V R++DPLW+LK+ L+ GS
Sbjct: 189 MRMTLDSICKVGFGVEIGTLSPDLPE-NSFAQAFDAANIIVTLRFIDPLWRLKKFLHVGS 247
Query: 61 EAALKKNIKIIDDFVHSLIKTRR-ELLSMQ---KDFSDKEDILSRF--LLESKKDSSNMT 114
EA L++++K++DDF +S+I+ R+ E+L + K K DILSRF L E+ D +
Sbjct: 248 EALLEQSMKLVDDFTYSVIRRRKAEILQARASGKQEKIKHDILSRFIELGEAGGDEGGGS 307
Query: 115 ---DKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEV------------I 159
DK LRD++LNF+IAG+DT+A TLSWF YM +P + +K+ +E+ +
Sbjct: 308 FGDDKSLRDVVLNFVIAGRDTTATTLSWFTYMAMTHPAVADKLRRELAAFEDERAREEGV 367
Query: 160 NVTSTSQESNL--NLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDIL 217
+ + E++ + +F S +S + K+ YLHA +TETLRLYPAVP + E D+L
Sbjct: 368 ALADAAGEASFAARVAQFASLLSYDAVGKLVYLHACVTETLRLYPAVPQDPKGIVEDDVL 427
Query: 218 PDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWLK--DGIFQPESSFKFISFHAG 275
PDG V G V Y+ Y+MGRM Y WG DA F PERWL G F+ S FKF +F AG
Sbjct: 428 PDGTKVRAGGMVTYVPYSMGRMEYNWGPDAASFRPERWLSGDGGAFRNASPFKFTAFQAG 487
Query: 276 PRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGL 325
PRICLGKD AY QMK+ L RF+ F L E + V YR M L + HGL
Sbjct: 488 PRICLGKDSAYLQMKMALAILFRFYTFDLV-EDHPVKYRMMTILSMAHGL 536
>gb|AAK52956.1| cytochrome P450-like protein [Zea mays]
Length = 543
Score = 268 bits (685), Expect = 2e-70
Identities = 157/351 (44%), Positives = 214/351 (60%), Gaps = 28/351 (7%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDPLWKLKRVLNFGS 60
M+ TLDSI KVGFGVE+ L E N F +AF+ +N + R++DPLW++KR + GS
Sbjct: 187 MRMTLDSICKVGFGVEIGTLSPDLPE-NSFAQAFDAANIIITLRFIDPLWRIKRFFHVGS 245
Query: 61 EAALKKNIKIIDDFVHSLIKTRR----ELLSMQKDFSDKEDILSRF--LLESKKDSSNM- 113
EA L ++IK++D+F +S+I+ R+ E+ + K K DILSRF L E+ D
Sbjct: 246 EALLAQSIKLVDEFTYSVIRRRKAEIVEVRASGKQEKMKHDILSRFIELGEAGDDGGGFG 305
Query: 114 TDKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTST-SQESNLNL 172
DK LRD++LNF+IAG+DT+A TLSWF +M +P + EK+ +E+ + ++E + L
Sbjct: 306 DDKSLRDVVLNFVIAGRDTTATTLSWFTHMAMSHPDVAEKLRRELCAFEAERAREEGVTL 365
Query: 173 -----------------DEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHD 215
+F ++ +L K+ YLHA +TETLRLYPAVP + E D
Sbjct: 366 VLCGGADADDKAFAARVAQFAGLLTYDSLGKLVYLHACVTETLRLYPAVPQDPKGILEDD 425
Query: 216 ILPDGYIVNKGETVYYLSYAMGRMPYIWGDDAQEFLPERWL-KDGIFQPESSFKFISFHA 274
+LPDG V G V Y+ Y+MGRM Y WG DA F PERW+ +DG F+ S FKF +F A
Sbjct: 426 VLPDGTKVRAGGMVTYVPYSMGRMEYNWGPDAASFRPERWINEDGAFRNASPFKFTAFQA 485
Query: 275 GPRICLGKDFAYRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGL 325
GPRICLGKD AY QMK+ L RF+ F+L E + V YR M L + HGL
Sbjct: 486 GPRICLGKDSAYLQMKMALAILFRFYSFRL-LEGHPVQYRMMTILSMAHGL 535
>dbj|BAB11174.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|15237250|ref|NP_197710.1| cytochrome P450 family
protein [Arabidopsis thaliana]
gi|25083267|gb|AAN72056.1| cytochrome P450-like protein
[Arabidopsis thaliana] gi|13549071|gb|AAK29622.1|
CYP86B1 [Arabidopsis thaliana]
Length = 559
Score = 262 bits (670), Expect = 9e-69
Identities = 133/328 (40%), Positives = 205/328 (61%), Gaps = 5/328 (1%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D++ + FGV+ CL F KAF D+ R++ P +WK R L+
Sbjct: 208 LRLTFDNVCMIAFGVDPGCLGPDQPVIP-FAKAFEDATEAAVVRFVMPTCVWKFMRYLDI 266
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYL 118
G+E LK++IK +DDF +I+TR++ LS++ + + + D+L+ F+ + + +DK+L
Sbjct: 267 GTEKKLKESIKGVDDFADEVIRTRKKELSLEGETTKRSDLLTVFMGLRDEKGESFSDKFL 326
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQE-SNLNLDEFVS 177
RDI +NF++AG+DTS+ LSWFF++L KNP ++EK+ E+ + + N ++
Sbjct: 327 RDICVNFILAGRDTSSVALSWFFWLLEKNPEVEEKIMVEMCKILRQRDDHGNAEKSDYEP 386
Query: 178 NISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
+ KM YL AAL+E LRLYP+VP+ + +E D+ PDG ++ KG+ V Y YAMG
Sbjct: 387 VFGPEEIKKMDYLQAALSEALRLYPSVPVDHKEVQEDDVFPDGTMLKKGDKVIYAIYAMG 446
Query: 238 RMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMALV 297
RM IWG D EF PERWL+DG F ES++KF +F+ GPR+CLGKDFAY QMK + A+V
Sbjct: 447 RMEAIWGKDCLEFRPERWLRDGRFMSESAYKFTAFNGGPRLCLGKDFAYYQMKSTAAAIV 506
Query: 298 RFFRFKLENETNDVTYRTMFTLHIDHGL 325
++ K+ N + V + T+++ HGL
Sbjct: 507 YRYKVKVVN-GHKVEPKLALTMYMKHGL 533
>emb|CAB93726.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|11249681|pir||T50510 cytochrome P450-like protein -
Arabidopsis thaliana
Length = 550
Score = 261 bits (666), Expect = 3e-68
Identities = 132/338 (39%), Positives = 212/338 (62%), Gaps = 8/338 (2%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D++ + FGV+ CL E F KAF D+ R++ P +WKL R LN
Sbjct: 205 LRLTFDNVCMIAFGVDPGCLSPKLPEIP-FAKAFEDATEATVVRFVMPKFVWKLMRSLNL 263
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYL 118
G+E LK++I +DDF +I+TR++ +S++ + + + D+L+ F+ ++ +DK+L
Sbjct: 264 GTEKKLKESINGVDDFAEEVIRTRKKEMSLETEIAKRPDLLTIFMGLRDENGQKFSDKFL 323
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQE---SNLNLDEF 175
RDI +NF++AG+DTS+ LSWFF+++ KNP ++EK+ + + + + N+ E+
Sbjct: 324 RDICVNFILAGRDTSSVALSWFFWLIEKNPEVEEKIMMGICKILEQRVDHGDTKKNM-EY 382
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
+ KM YL AAL+ETLRLYP+VP+ + E D+ PDG + KGE V Y YA
Sbjct: 383 EPVFRPEEIKKMDYLQAALSETLRLYPSVPVDHKEVLEDDVFPDGTKLKKGEKVIYAIYA 442
Query: 236 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
MGRM IWG D +EF PERWL+DG + ES++KF +F+ GPR+CLGKDFAY QM+ V+ A
Sbjct: 443 MGRMETIWGKDCREFKPERWLRDGRYMSESAYKFTAFNGGPRLCLGKDFAYYQMRYVAAA 502
Query: 296 LVRFFRFKLENE-TNDVTYRTMFTLHIDHGLPLYATPR 332
++ ++ +++++ + V + T+++ HGL + R
Sbjct: 503 IIYRYKVRVDDKGGHKVEPKMALTMYMKHGLKVNMVKR 540
>gb|AAN15497.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|22531050|gb|AAM97029.1| cytochrome P450-like protein
[Arabidopsis thaliana] gi|30682301|ref|NP_196442.2|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 488
Score = 261 bits (666), Expect = 3e-68
Identities = 132/338 (39%), Positives = 212/338 (62%), Gaps = 8/338 (2%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D++ + FGV+ CL E F KAF D+ R++ P +WKL R LN
Sbjct: 143 LRLTFDNVCMIAFGVDPGCLSPKLPEIP-FAKAFEDATEATVVRFVMPKFVWKLMRSLNL 201
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYL 118
G+E LK++I +DDF +I+TR++ +S++ + + + D+L+ F+ ++ +DK+L
Sbjct: 202 GTEKKLKESINGVDDFAEEVIRTRKKEMSLETEIAKRPDLLTIFMGLRDENGQKFSDKFL 261
Query: 119 RDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQE---SNLNLDEF 175
RDI +NF++AG+DTS+ LSWFF+++ KNP ++EK+ + + + + N+ E+
Sbjct: 262 RDICVNFILAGRDTSSVALSWFFWLIEKNPEVEEKIMMGICKILEQRVDHGDTKKNM-EY 320
Query: 176 VSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYA 235
+ KM YL AAL+ETLRLYP+VP+ + E D+ PDG + KGE V Y YA
Sbjct: 321 EPVFRPEEIKKMDYLQAALSETLRLYPSVPVDHKEVLEDDVFPDGTKLKKGEKVIYAIYA 380
Query: 236 MGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMA 295
MGRM IWG D +EF PERWL+DG + ES++KF +F+ GPR+CLGKDFAY QM+ V+ A
Sbjct: 381 MGRMETIWGKDCREFKPERWLRDGRYMSESAYKFTAFNGGPRLCLGKDFAYYQMRYVAAA 440
Query: 296 LVRFFRFKLENE-TNDVTYRTMFTLHIDHGLPLYATPR 332
++ ++ +++++ + V + T+++ HGL + R
Sbjct: 441 IIYRYKVRVDDKGGHKVEPKMALTMYMKHGLKVNMVKR 478
>gb|AAP54351.1| putative cytochrome P450 protein [Oryza sativa (japonica
cultivar-group)] gi|37535524|ref|NP_922064.1| putative
cytochrome P450 protein [Oryza sativa (japonica
cultivar-group)] gi|18087871|gb|AAL59025.1| putative
cytochrome P450 protein [Oryza sativa]
Length = 560
Score = 239 bits (609), Expect = 1e-61
Identities = 124/347 (35%), Positives = 209/347 (59%), Gaps = 19/347 (5%)
Query: 1 MKCTLDSIFKVGFGVELNCLEKSSKEANIFMKAFNDSNAFVFKRYLDP--LWKLKRVLNF 58
++ T D++ + FGV+ CL E F KAF D+ R++ P +W+ R L
Sbjct: 213 LRLTFDNVCMIAFGVDPGCLRPGLPEIP-FAKAFEDATEATIVRFVTPTAVWRAMRALGV 271
Query: 59 GSEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSD--KEDILSRF--LLESKKDSSNMT 114
G E L++++ +D F + +I+ R+E ++ + D+L+ F + ++ ++ +
Sbjct: 272 GHERVLQRSLAGVDRFAYDVIRQRKEEVAAGGGGGGGGRSDLLTIFTKMRDADTGAAAYS 331
Query: 115 DKYLRDIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEVINVTSTSQES------ 168
DK+LRDI +NF++AG+DTS+ L+WFF++L KNP ++ K+ +E+ ++ + + S
Sbjct: 332 DKFLRDICVNFILAGRDTSSVALAWFFWLLNKNPAVEAKILEEIDDIVAARRSSPPAPAV 391
Query: 169 ---NLNLDEFVSNISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNK 225
+ D+ V + + + KM YLHAAL+E LRLYP+VP+ + E ++ PDG ++ K
Sbjct: 392 AANGADEDDLVFHPEE--VKKMEYLHAALSEALRLYPSVPVDHKEVVEDEVFPDGTVLKK 449
Query: 226 GETVYYLSYAMGRMPYIWGDDAQEFLPERWLKDGIFQPESSFKFISFHAGPRICLGKDFA 285
G V Y Y MGRM IWG+D +E+ PERWL+DG F ES++KF +F+ GPR+CLGKDFA
Sbjct: 450 GTKVIYAMYTMGRMESIWGEDCREYKPERWLRDGRFMGESAYKFTAFNGGPRLCLGKDFA 509
Query: 286 YRQMKIVSMALVRFFRFKLENETNDVTYRTMFTLHIDHGLPLYATPR 332
Y QMK + +++R + ++ + + V + T+++ HGL + T R
Sbjct: 510 YYQMKFAAASILRRYHVRVV-DGHPVAPKMALTMYMKHGLKVKLTKR 555
>ref|NP_915858.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20804566|dbj|BAB92258.1| putative
cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa (japonica cultivar-group)]
Length = 511
Score = 225 bits (574), Expect = 1e-57
Identities = 124/337 (36%), Positives = 200/337 (58%), Gaps = 26/337 (7%)
Query: 6 DSIFKVGFGVELNCLEK---SSKEANIFMKAFNDSNAFVFKRYLDP---LWKLKRVLNFG 59
D+I V FG + CL K ++ ++ FM+AFND+ + R+ P LW++K++ N
Sbjct: 190 DNICLVAFGEDPACLTKERMAAPQSAEFMRAFNDAQNAILARFNSPAKSLWRVKKLFNME 249
Query: 60 SEAALKKNIKIIDDFVHSLIKTRRELLSMQKDFSDKEDILSRFLLESKKDSSNMTDKYLR 119
E +++ + I F +++ RRE + + +D LSRF S +D+ LR
Sbjct: 250 PERRMREALATIHGFAERIVRERRE--RGEAGLARGDDFLSRFAA-----SGEHSDESLR 302
Query: 120 DIILNFMIAGKDTSANTLSWFFYMLCKNPLIQEKVAQEV--INVTSTSQESNLNLDEFVS 177
D++ NF++AG+DT+++ L+WFF+++ P ++++V +E+ + +S S ++ + DE
Sbjct: 303 DVVTNFVLAGRDTTSSALTWFFWIVSGRPDVEDRVVREIRAVRASSGSTDATFSFDE--- 359
Query: 178 NISDATLDKMHYLHAALTETLRLYPAVPMSGRTAEEHDILPDGYIVNKGETVYYLSYAMG 237
L + HYLHAA+TE++RLYP V + + +E D LPDG V KG V Y +YAMG
Sbjct: 360 ------LREKHYLHAAITESMRLYPPVAIDTHSCKEDDFLPDGTFVGKGWLVMYSAYAMG 413
Query: 238 RMPYIWGDDAQEFLPERWLKD-GIFQPESSFKFISFHAGPRICLGKDFAYRQMKIVSMAL 296
RM IWG D +E+ PERWL + G F+PES+FK+ F+AGPRIC+GK+ AY QMK + +
Sbjct: 414 RMEGIWGADCEEYRPERWLDEAGAFRPESTFKYPVFNAGPRICIGKEMAYIQMKSIVACV 473
Query: 297 VRFFRFKLENETNDVTYRTM-FTLHIDHGLPLYATPR 332
+ F + ++ N+ + TL + GLP+ T R
Sbjct: 474 LEKFSLRYASDANERPRSVLSLTLRMKWGLPMKVTIR 510
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.138 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 542,824,924
Number of Sequences: 2540612
Number of extensions: 22076909
Number of successful extensions: 65362
Number of sequences better than 10.0: 4692
Number of HSP's better than 10.0 without gapping: 2633
Number of HSP's successfully gapped in prelim test: 2061
Number of HSP's that attempted gapping in prelim test: 55485
Number of HSP's gapped (non-prelim): 4958
length of query: 337
length of database: 863,360,394
effective HSP length: 128
effective length of query: 209
effective length of database: 538,162,058
effective search space: 112475870122
effective search space used: 112475870122
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC146342.8


