

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146342.7 - phase: 0
(512 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
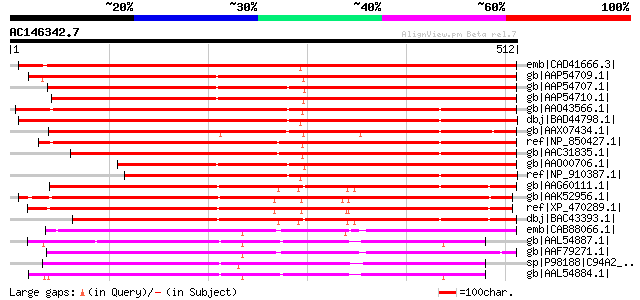
Score E
Sequences producing significant alignments: (bits) Value
emb|CAD41666.3| OSJNBa0019K04.13 [Oryza sativa (japonica cultiva... 568 e-160
gb|AAP54709.1| cytochrome P450-like protein [Oryza sativa (japon... 551 e-155
gb|AAP54707.1| cytochrome P450-like protein [Oryza sativa (japon... 547 e-154
gb|AAP54710.1| cytochrome P450-like protein [Oryza sativa (japon... 533 e-150
gb|AAO43566.1| At2g45510 [Arabidopsis thaliana] gi|2979544|gb|AA... 517 e-145
dbj|BAD44798.1| putative cytochrome P450 [Oryza sativa (japonica... 506 e-142
gb|AAX07434.1| cytochrome P450 CYPD [Pinus taeda] 496 e-139
ref|NP_850427.1| cytochrome P450 family protein [Arabidopsis tha... 495 e-138
gb|AAC31835.1| putative cytochrome P450 [Arabidopsis thaliana] g... 479 e-134
gb|AAO00706.1| putative cytochrome P450-dependent fatty acid hyd... 474 e-132
ref|NP_910387.1| ESTs AU056036(S20239),C72753(E2173), AU056035(S... 451 e-125
gb|AAG60111.1| cytochrome P450, putative [Arabidopsis thaliana] 396 e-108
gb|AAK52956.1| cytochrome P450-like protein [Zea mays] 396 e-108
ref|XP_470289.1| putative plant cytochrome P-450 protein [Oryza ... 394 e-108
dbj|BAC43393.1| unknown protein [Arabidopsis thaliana] gi|306978... 380 e-104
emb|CAB88066.1| cytochrome P450-like protein [Arabidopsis thalia... 333 8e-90
gb|AAL54887.1| cytochrome P450-dependent fatty acid hydroxylase ... 325 3e-87
gb|AAF79271.1| F12K21.15 [Arabidopsis thaliana] gi|15218671|ref|... 320 9e-86
sp|P98188|C94A2_VICSA Cytochrome P450 94A2 (P450-dependent fatty... 316 1e-84
gb|AAL54884.1| cytochrome P450-dependent fatty acid hydroxylase ... 314 5e-84
>emb|CAD41666.3| OSJNBa0019K04.13 [Oryza sativa (japonica cultivar-group)]
gi|50928103|ref|XP_473579.1| OSJNBa0019K04.13 [Oryza
sativa (japonica cultivar-group)]
Length = 506
Score = 568 bits (1464), Expect = e-160
Identities = 285/509 (55%), Positives = 376/509 (72%), Gaps = 10/509 (1%)
Query: 10 LSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMT 69
+ +P A+ A LLLV L L + ++ KK++Y PVAGT+F+Q++NF RL Y T
Sbjct: 1 MESPLSHPAMVALSLLLLVALYLAR---RAVLGKKRRYPPVAGTMFHQLLNFGRLLEYHT 57
Query: 70 DLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVD 129
+L+RKYRT+R+L P + VYT EP+NVE+ILKTNF NYGKG + L+DLLGDGIF VD
Sbjct: 58 ELSRKYRTFRMLTPTCNYVYTVEPANVEHILKTNFANYGKGPMTHDVLEDLLGDGIFNVD 117
Query: 130 GEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSE--AATSNFKLEIQDLLMKS 187
G WR+QRK++S EFST++LRD+S+++FR AA++A I+ AA ++++QDLLM++
Sbjct: 118 GGMWRQQRKVASLEFSTRVLRDYSSAVFRDTAAELAGILERGPAAKGRERVDMQDLLMRA 177
Query: 188 TLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAA 247
TLDS F+V FG L + GSS+EG FA AFD AS L+R+ D+ WK+K+FLNI SEA
Sbjct: 178 TLDSFFRVGFGVNLGVLSGSSKEGLVFARAFDDASEQVLFRFFDLLWKVKRFLNISSEAT 237
Query: 248 LRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKE-----YDTTYLRDII 302
++++ +N+FV +I+ +I+QM+ + K DILSRFL +E +D Y+RDII
Sbjct: 238 MKQSIRTINDFVYSIIDRKIEQMSREQHEFAKKEDILSRFLLEREKDPGCFDNKYIRDII 297
Query: 303 LNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTDEAL 362
LNFVIAG+DTTA TLSWF+Y +CK VQ+K A EVR+AT +F S +T++A+
Sbjct: 298 LNFVIAGRDTTAGTLSWFLYAVCKNQRVQDKIAREVRDATTGDRDVGVQDFSSFLTEDAI 357
Query: 363 EKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWG 422
KM YLHA LTETLRLYP VP+D K CF+DDTLPDG++VKKGDMV+YQPY MGRMKF+WG
Sbjct: 358 NKMQYLHAALTETLRLYPGVPLDVKYCFSDDTLPDGHAVKKGDMVNYQPYPMGRMKFLWG 417
Query: 423 DDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFK 482
D+AEEF+P RWLD++G F AE+PFKFTAFQAGPRICLGKEFAYRQMKI SAVLL FRF+
Sbjct: 418 DNAEEFKPERWLDDSGMFVAESPFKFTAFQAGPRICLGKEFAYRQMKIVSAVLLYFFRFE 477
Query: 483 LNDEKRNVTYKTMINLHIDGGLEIKALHR 511
+ D+ V Y+ M+ L +DG ++AL R
Sbjct: 478 MWDDDATVGYRPMLTLKMDGPFYLRALAR 506
>gb|AAP54709.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536240|ref|NP_922422.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146747|gb|AAM12483.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 515
Score = 551 bits (1419), Expect = e-155
Identities = 274/502 (54%), Positives = 368/502 (72%), Gaps = 11/502 (2%)
Query: 20 SASLTLLLVQLL---LRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYR 76
+A++ L+LV + L + + ++++ PV GT F+Q+ + R+H Y T L+R++
Sbjct: 15 AAAVGLVLVVAICTYLAVVATRKQKRRRRRRPPVVGTAFHQLYHVRRVHDYHTALSREHM 74
Query: 77 TYRLLNPF-RSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWRE 135
T+RLL P R ++YT +P+ VE+IL+TNF NYGKG +N+ N+ DL GDGIFAVDG+KW++
Sbjct: 75 TFRLLVPAGREQIYTCDPAVVEHILRTNFANYGKGSFNHGNMSDLFGDGIFAVDGDKWKQ 134
Query: 136 QRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQV 195
QRKI+S++F+T+ LRDFS +F++NAAK+A +VS A SN ++ Q LM++T+DSIF +
Sbjct: 135 QRKIASYDFTTRALRDFSGDVFKRNAAKLAGVVSSHAASNQSMDFQGFLMRATMDSIFTI 194
Query: 196 AFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAALRKNTEVL 255
AFG +LN++ GS E G+ FA AFD AS T+ RY++ FWK+ + LN+G+EA L++ +V+
Sbjct: 195 AFGQDLNTLDGSGE-GRRFAAAFDDASEFTMLRYLNPFWKLSRLLNVGAEAMLKERIKVV 253
Query: 256 NEFVIKLINTRIQQM-NSKGDSIRKSGDILSRFLQVKEYDT-----TYLRDIILNFVIAG 309
+ FV KLI R ++ N+K DIL+RF+Q D+ YLRDIILN VIAG
Sbjct: 254 DGFVYKLIRDRSDELSNTKAHDTDSRQDILTRFIQATTSDSGTVDYKYLRDIILNIVIAG 313
Query: 310 KDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTDEALEKMNYLH 369
KDTTA +L+WF+YM+CK+P VQEK E EATN +S EF +TDEAL KM+YLH
Sbjct: 314 KDTTAGSLAWFLYMMCKHPEVQEKICHEAMEATNAGEAASIDEFSQSLTDEALNKMHYLH 373
Query: 370 ATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFR 429
A LTETLRLYPAVP+D K CF+DD LP+G++V KGD+V Y PYAMGRM+ +WG DAE FR
Sbjct: 374 AALTETLRLYPAVPLDNKQCFSDDVLPNGFNVSKGDIVFYIPYAMGRMESLWGKDAESFR 433
Query: 430 PARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRN 489
P RWLDENG FQ E+PFKFTAFQAGPRICLGK+FAYRQMKIF+AVLL F KL DEK
Sbjct: 434 PERWLDENGVFQQESPFKFTAFQAGPRICLGKDFAYRQMKIFAAVLLRFFVLKLRDEKEI 493
Query: 490 VTYKTMINLHIDGGLEIKALHR 511
++Y+TMI L +D GL + A+ R
Sbjct: 494 ISYRTMITLSVDQGLHLTAMAR 515
>gb|AAP54707.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536236|ref|NP_922420.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146758|gb|AAM12494.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 495
Score = 547 bits (1410), Expect = e-154
Identities = 271/478 (56%), Positives = 356/478 (73%), Gaps = 7/478 (1%)
Query: 39 SNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLLNPFRSE-VYTSEPSNVE 97
SN K+++ PV GTVF+Q+ N R+H Y T L+R++ T+R+L P + +YT +P+ VE
Sbjct: 20 SNKQKRRRRPPVVGTVFHQLYNVRRIHDYHTALSREHTTFRMLVPAGGDQIYTCDPAVVE 79
Query: 98 YILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIF 157
+ILKTNF NYGKG +N+ N KDL GDGIFA+DGEKW++QRKI+S++FST+ LRDFS ++F
Sbjct: 80 HILKTNFANYGKGPFNHGNAKDLFGDGIFAIDGEKWKQQRKIASYDFSTRALRDFSCAVF 139
Query: 158 RKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANA 217
++NAAK+A IVS A SN ++ Q L++++T+DSIF +AFGT+LN++ GS E G FA A
Sbjct: 140 KRNAAKLAGIVSNHAASNQSMDFQGLMLRATMDSIFTIAFGTDLNTLDGSGE-GSRFAAA 198
Query: 218 FDTASALTLYRYVDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSI 277
FD AS T+ RY+ WK+ + LN+G EA L++ +V++EFV +LI R ++++ DS
Sbjct: 199 FDDASEFTMLRYISPLWKLARLLNVGVEAMLKERIKVVDEFVYRLIRARSDELSNSHDSG 258
Query: 278 RKSGDILSRFLQVKEYDTT----YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEK 333
+ DILSRFLQ D+ YLRDIILN VIAGKDTTA L+WF+YM+CK+P VQEK
Sbjct: 259 SRQ-DILSRFLQATTSDSVVDYKYLRDIILNIVIAGKDTTAGALAWFLYMVCKHPEVQEK 317
Query: 334 AAEEVREATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADD 393
E AT+ +S EF+ +TD+AL M+YLHA LTETLRLYP+VP++ K CF+DD
Sbjct: 318 ICHEAMVATSAGDTASVDEFLQSLTDQALNNMHYLHAALTETLRLYPSVPMENKQCFSDD 377
Query: 394 TLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQA 453
LP+G++V KGD+V + PYAMGRM+ +WG DAE FRP RWLDENG FQ E+PFKFTAFQA
Sbjct: 378 VLPNGFNVSKGDIVFFIPYAMGRMESLWGKDAEYFRPERWLDENGVFQQESPFKFTAFQA 437
Query: 454 GPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
GPRICLGKEFAYRQMKIF+AVLL F KL DEK V+Y+T + L ID GL + A R
Sbjct: 438 GPRICLGKEFAYRQMKIFAAVLLRFFVLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 495
>gb|AAP54710.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536242|ref|NP_922423.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146744|gb|AAM12480.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 516
Score = 533 bits (1372), Expect = e-150
Identities = 262/476 (55%), Positives = 353/476 (74%), Gaps = 8/476 (1%)
Query: 43 KKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLL-NPFRSEVYTSEPSNVEYILK 101
+++++ PV GTVF+Q+ + RLH Y T L R++ T+RLL P R +YT +P+ VE+IL+
Sbjct: 42 RRRRHPPVVGTVFHQLYHVRRLHDYYTALCREHTTFRLLATPGRRNIYTCDPAVVEHILR 101
Query: 102 TNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNA 161
TNF +YGKG N + L DL G+GIFAVDGEKW+ QRKI+S++F+T+ LRDFS+ +F++NA
Sbjct: 102 TNFPSYGKGPLNSEILNDLFGEGIFAVDGEKWKTQRKIASYDFTTRALRDFSSDVFKRNA 161
Query: 162 AKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTA 221
AK+A +VS A SN ++ + LL ++T+DSIF +AFG +LN++ GS E G++FA AFD A
Sbjct: 162 AKLAGVVSNHAASNQSMDFKGLLTRATMDSIFTIAFGQDLNTLDGSGE-GRHFAKAFDDA 220
Query: 222 SALTLYRYVDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQM-NSKGDSIRKS 280
L RY++ FWK+ + LN+G+EA L++ +V++EFV KLI R ++ N+ R
Sbjct: 221 GEYLLLRYLNPFWKLARLLNVGAEATLKERIKVVDEFVYKLIRARSDELSNTMAQDHRSR 280
Query: 281 GDILSRFLQVKEYDT-----TYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAA 335
D+LSRF+Q D+ YLRDI+LN VIA KD+T+ +L+WF+YM CK P VQEK
Sbjct: 281 DDLLSRFIQATTSDSGTVDYKYLRDIVLNIVIAAKDSTSGSLAWFLYMACKRPEVQEKIF 340
Query: 336 EEVREATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTL 395
+EV E TN +S EF++ +TD+AL KM+YLHA LTETLRLYP+VP++ K CF+DD L
Sbjct: 341 DEVMETTNAGDCASIDEFLTSLTDQALNKMHYLHAALTETLRLYPSVPLENKQCFSDDVL 400
Query: 396 PDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGP 455
P+G+SV KGD V Y PYAMGRM+F+WG DAE FRP RWLDE+G FQ E+PFKFTAFQAGP
Sbjct: 401 PNGFSVSKGDGVFYMPYAMGRMEFLWGKDAEAFRPERWLDEHGVFQQESPFKFTAFQAGP 460
Query: 456 RICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
RIC+GK+FAYRQMKIF+AVL+ F FKL D+K NV+Y+T I L ID L + A R
Sbjct: 461 RICIGKDFAYRQMKIFAAVLIRSFVFKLRDKKDNVSYRTAITLAIDQDLHLTATAR 516
>gb|AAO43566.1| At2g45510 [Arabidopsis thaliana] gi|2979544|gb|AAC06153.1| putative
cytochrome P450 [Arabidopsis thaliana]
gi|15225499|ref|NP_182075.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|7430723|pir||T00864 cytochrome
P450 homolog F17K2.4 - Arabidopsis thaliana
Length = 511
Score = 517 bits (1331), Expect = e-145
Identities = 267/512 (52%), Positives = 358/512 (69%), Gaps = 11/512 (2%)
Query: 7 MDFLSNPFLFAALSASLTLLL-VQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLH 65
M+ L++ + A + + L + L++R KS N K+Y PV TVF+ + + + L+
Sbjct: 1 MEILTSIAITVATTIFIVLCFTIYLMIRIFTGKSRN--DKRYAPVHATVFDLLFHSDELY 58
Query: 66 HYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGI 125
Y T++AR+ TYR L+P +SE+ T++P NVE+ILKT F+NY KG + +N+ DLLG GI
Sbjct: 59 DYETEIAREKPTYRFLSPGQSEILTADPRNVEHILKTRFDNYSKGHSSRENMADLLGHGI 118
Query: 126 FAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLM 185
FAVDGEKWR+QRK+SS EFST++LRDFS S+FR+NA+K+ VSE A S + QDLLM
Sbjct: 119 FAVDGEKWRQQRKLSSFEFSTRVLRDFSCSVFRRNASKLVGFVSEFALSGKAFDAQDLLM 178
Query: 186 KSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSE 245
+ TLDSIF+V FG EL + G S+EG+ F AFD + T R++D WK+K F NIGS+
Sbjct: 179 RCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGSQ 238
Query: 246 AALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYD-----TTYLRD 300
+ L+K+ +++FV LI T+ +++ + +++ + DILSRFL E D YLRD
Sbjct: 239 SKLKKSIATIDKFVYSLITTKRKELAKEQNTVVRE-DILSRFLVESEKDPENMNDKYLRD 297
Query: 301 IILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREAT-NTKTVSSCTEFVSCVTD 359
IILNF+IAGKDTTAA LSWF+YMLCK P VQEK +E+R+ T + + + FV + +
Sbjct: 298 IILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTFSHEKTTDVNGFVESINE 357
Query: 360 EALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKF 419
EAL++M+YLHA L+ETLRLYP VPVD + DD LPDG+ V KGD + Y YAMGRM +
Sbjct: 358 EALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRMTY 417
Query: 420 IWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCF 479
IWG DAEEF+P RWL ++G FQ E+PFKF +F AGPRICLGK+FAYRQMKI S LL F
Sbjct: 418 IWGQDAEEFKPERWL-KDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLHFF 476
Query: 480 RFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
RFK+ DE V YK M+ LH+DGGL + A+ R
Sbjct: 477 RFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>dbj|BAD44798.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 525
Score = 506 bits (1303), Expect = e-142
Identities = 259/509 (50%), Positives = 351/509 (68%), Gaps = 8/509 (1%)
Query: 10 LSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMT 69
L++ A LS +L ++ V L K + P+ GTVF Q+ NF+R+
Sbjct: 16 LASALAIALLSIALYIIGVVASFAVLCAKEFAERAHDRPPLVGTVFRQLKNFDRMFDEHV 75
Query: 70 DLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVD 129
+ A +RT R++ P EV+TS+P+ VE++LK +F Y KG + +KDL GDGIFA D
Sbjct: 76 NYATAHRTSRIVYPGHCEVFTSDPAVVEHVLKNSFSKYSKGDFLTTAMKDLFGDGIFATD 135
Query: 130 GEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTL 189
G+ WR QRK++S+EFSTK+LRDFS+ FR+NAAK+A +S AA + + IQDLLM++T+
Sbjct: 136 GDMWRHQRKLASYEFSTKVLRDFSSDTFRRNAAKLAEKISCAAANRISINIQDLLMRATM 195
Query: 190 DSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAALR 249
DSIF+V FG ELN++ GS E G F+ AFD A++L YR+VD+ WK+K++LNIGSEA L+
Sbjct: 196 DSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKLKRYLNIGSEAKLK 255
Query: 250 KNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKE-----YDTTYLRDIILN 304
+N ++++ FV+KLI+ + +QM D K DILSRF+ E D YLRDI+LN
Sbjct: 256 RNIQIIDSFVMKLIHQKREQMKIAADYKTKE-DILSRFVLASEQDPGTMDDRYLRDIVLN 314
Query: 305 FVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATN-TKTVSSCTEFVSCVTDEALE 363
F+IAGKDTT TL+WF Y+LCK P VQ+K A E+RE +K ++ F + + A+
Sbjct: 315 FLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDNTIESFTKRLDEGAIS 374
Query: 364 KMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGD 423
KM+YL AT++ETLRLYPAVPVDAK+ DD LP+GY V KGD ++Y YAMGRM ++WG+
Sbjct: 375 KMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYMIYAMGRMTYLWGE 434
Query: 424 DAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKL 483
DA+EFRP RWL NG +Q E+PFKF +F AGPRICLGKEFA+RQMKI +A L+ F+F+L
Sbjct: 435 DAQEFRPERWL-VNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKIMAATLIHFFKFRL 493
Query: 484 NDEKRNVTYKTMINLHIDGGLEIKALHRD 512
DE + YKTM LHID GL + A R+
Sbjct: 494 EDESKEPIYKTMFTLHIDNGLHLLANPRE 522
>gb|AAX07434.1| cytochrome P450 CYPD [Pinus taeda]
Length = 518
Score = 496 bits (1278), Expect = e-139
Identities = 253/490 (51%), Positives = 356/490 (72%), Gaps = 22/490 (4%)
Query: 40 NNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYI 99
N KK Y PV GT+ N +NF RLH Y TD A++Y+T+R++ P S V+T++P NVE+I
Sbjct: 29 NWRKKGVYPPVVGTMLNHAINFERLHDYHTDQAQRYKTFRVVYPTCSYVFTTDPVNVEHI 88
Query: 100 LKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRK 159
LKTNF NY KG +NY +KDLLGDGIF VDG+KWR+QRK++S EF++K+L+DFS+ +F
Sbjct: 89 LKTNFANYDKGTFNYDIMKDLLGDGIFNVDGDKWRQQRKLASSEFASKVLKDFSSGVFCN 148
Query: 160 NAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEG---KNFAN 216
NAAK+ANI+++AA N +E+QDL M+S+LDSI +V FG ++NS+ S E +FA
Sbjct: 149 NAAKLANILAQAAKLNLSVEMQDLFMRSSLDSICKVVFGIDINSLSSSKAESGPEASFAK 208
Query: 217 AFDTASALTLYRY-VDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQM-NSKG 274
AFD A+A+ +R+ V FWK+++F N+GSEA LR N +++++F+ K+I+ R Q+M +++
Sbjct: 209 AFDVANAMVFHRHMVGSFWKVQRFFNVGSEAILRDNIKMVDDFLYKVIHFRRQEMFSAEK 268
Query: 275 DSIRKSGDILSRFLQVKEYDT------TYLRDIILNFVIAGKDTTAATLSWFMYMLCKYP 328
+++R DILSR++ + + +T YLRD+ILNF++A +DTTA LSWF+YMLCK+
Sbjct: 269 ENVRP--DILSRYIIISDKETDGKVSDKYLRDVILNFMVAARDTTAIALSWFIYMLCKHQ 326
Query: 329 AVQEKAAEEVREATNTKTVSSCTE------FVSCVTDEALEKMNYLHATLTETLRLYPAV 382
VQEK EE+ +T+ TE F +TDEAL KM+YLHA+L+ETLRLYPA+
Sbjct: 327 HVQEKLLEEIISSTSVHEDQYSTECNDIASFAQSLTDEALGKMHYLHASLSETLRLYPAL 386
Query: 383 PVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQA 442
PVD K +DTLPDG+ VKKGD V++ PYAMGRM ++WGDDA+EF+P RW+ ++G F
Sbjct: 387 PVDGKYVVNEDTLPDGFKVKKGDSVNFLPYAMGRMSYLWGDDAKEFKPERWI-QDGIFHP 445
Query: 443 ENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHI-D 501
++PFKF AFQAGPR CLGK+FAY QMKI +AVL+ F+F+ K V Y+TM+ LH+ +
Sbjct: 446 KSPFKFPAFQAGPRTCLGKDFAYLQMKIVAAVLVRFFKFEAVKTK-EVRYRTMLTLHMNE 504
Query: 502 GGLEIKALHR 511
GL ++ R
Sbjct: 505 DGLNVQVTPR 514
>ref|NP_850427.1| cytochrome P450 family protein [Arabidopsis thaliana]
Length = 505
Score = 495 bits (1275), Expect = e-138
Identities = 254/488 (52%), Positives = 338/488 (69%), Gaps = 10/488 (2%)
Query: 30 LLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLLNPFRSEVY 89
L +R KS N K+Y PV T+F+ + ++L+ Y T++AR T+R L+P +SE++
Sbjct: 19 LTIRIFTGKSRN--DKRYTPVHATIFDLFFHSHKLYDYETEIARTKPTFRFLSPGQSEIF 76
Query: 90 TSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKML 149
T++P NVE+ILKT F NY KG NL DLLG GIFAVDGEKW++QRK+ S EFST++L
Sbjct: 77 TADPRNVEHILKTRFHNYSKGPVGTVNLADLLGHGIFAVDGEKWKQQRKLVSFEFSTRVL 136
Query: 150 RDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSE 209
R+FS S+FR +A+K+ ++E A S + QD+LMK TLDSIF+V FG EL + G S+
Sbjct: 137 RNFSYSVFRTSASKLVGFIAEFALSGKSFDFQDMLMKCTLDSIFKVGFGVELGCLDGFSK 196
Query: 210 EGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQ 269
EG+ F AFD + T R D FWK+K FLNIGSE+ L+K+ ++++FV LI T+ ++
Sbjct: 197 EGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGSESRLKKSIAIIDKFVYSLITTKRKE 256
Query: 270 MNSKGDSIRKSGDILSRFLQVKEYD-----TTYLRDIILNFVIAGKDTTAATLSWFMYML 324
+ SK + DILS+FL E D YLRDIILN ++AGKDTTAA+LSWF+YML
Sbjct: 257 L-SKEQNTSVREDILSKFLLESEKDPENMNDKYLRDIILNVMVAGKDTTAASLSWFLYML 315
Query: 325 CKYPAVQEKAAEEVREATNT-KTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVP 383
CK P VQEK +E+R+ T++ + + F+ VT+EAL +M YLHA L+ET+RLYP VP
Sbjct: 316 CKNPLVQEKIVQEIRDVTSSHEKTTDVNGFIESVTEEALAQMQYLHAALSETMRLYPPVP 375
Query: 384 VDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAE 443
+ DD LPDG+ V KGD + Y YAMGRM +IWG DAEEF+P RWL ++G FQ E
Sbjct: 376 EHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRMTYIWGQDAEEFKPERWL-KDGVFQPE 434
Query: 444 NPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGG 503
+ FKF +F AGPRIC+GK+FAYRQMKI S LL FRFK+ DE V+YK M+ LH+DGG
Sbjct: 435 SQFKFISFHAGPRICIGKDFAYRQMKIVSMALLHFFRFKMADENSKVSYKKMLTLHVDGG 494
Query: 504 LEIKALHR 511
L + A+ R
Sbjct: 495 LHLCAIPR 502
>gb|AAC31835.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|7430726|pir||T00404 probable cytochrome P450
At2g44890 [imported] - Arabidopsis thaliana
Length = 490
Score = 479 bits (1234), Expect = e-134
Identities = 243/456 (53%), Positives = 322/456 (70%), Gaps = 8/456 (1%)
Query: 62 NRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLL 121
++L+ Y T++AR T+R L+P +SE++T++P NVE+ILKT F NY KG NL DLL
Sbjct: 34 HKLYDYETEIARTKPTFRFLSPGQSEIFTADPRNVEHILKTRFHNYSKGPVGTVNLADLL 93
Query: 122 GDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQ 181
G GIFAVDGEKW++QRK+ S EFST++LR+FS S+FR +A+K+ ++E A S + Q
Sbjct: 94 GHGIFAVDGEKWKQQRKLVSFEFSTRVLRNFSYSVFRTSASKLVGFIAEFALSGKSFDFQ 153
Query: 182 DLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLN 241
D+LMK TLDSIF+V FG EL + G S+EG+ F AFD + T R D FWK+K FLN
Sbjct: 154 DMLMKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLN 213
Query: 242 IGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYD-----TT 296
IGSE+ L+K+ ++++FV LI T+ +++ SK + DILS+FL E D
Sbjct: 214 IGSESRLKKSIAIIDKFVYSLITTKRKEL-SKEQNTSVREDILSKFLLESEKDPENMNDK 272
Query: 297 YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNT-KTVSSCTEFVS 355
YLRDIILN ++AGKDTTAA+LSWF+YMLCK P VQEK +E+R+ T++ + + F+
Sbjct: 273 YLRDIILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFIE 332
Query: 356 CVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMG 415
VT+EAL +M YLHA L+ET+RLYP VP + DD LPDG+ V KGD + Y YAMG
Sbjct: 333 SVTEEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMG 392
Query: 416 RMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVL 475
RM +IWG DAEEF+P RWL ++G FQ E+ FKF +F AGPRIC+GK+FAYRQMKI S L
Sbjct: 393 RMTYIWGQDAEEFKPERWL-KDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMAL 451
Query: 476 LGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
L FRFK+ DE V+YK M+ LH+DGGL + A+ R
Sbjct: 452 LHFFRFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>gb|AAO00706.1| putative cytochrome P450-dependent fatty acid hydroxylase,
5'-partial [Oryza sativa (japonica cultivar-group)]
Length = 404
Score = 474 bits (1221), Expect = e-132
Identities = 235/406 (57%), Positives = 304/406 (73%), Gaps = 6/406 (1%)
Query: 110 GLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVS 169
G +N+ N KDL GDGIFA+DGEKW++QRKI+S++FST+ LRDFS ++F++NAAK+A IVS
Sbjct: 1 GPFNHGNAKDLFGDGIFAIDGEKWKQQRKIASYDFSTRALRDFSCAVFKRNAAKLAGIVS 60
Query: 170 EAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRY 229
A SN ++ Q L++++T+DSIF +AFGT+LN++ GS E G FA AFD AS T+ RY
Sbjct: 61 NHAASNQSMDFQGLMLRATMDSIFTIAFGTDLNTLDGSGE-GSRFAAAFDDASEFTMLRY 119
Query: 230 VDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQ 289
+ WK+ + LN+G EA L++ +V++EFV +LI R ++++ DS + DILSRFLQ
Sbjct: 120 ISPLWKLARLLNVGVEAMLKERIKVVDEFVYRLIRARSDELSNSHDSGSRQ-DILSRFLQ 178
Query: 290 VKEYDTT----YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTK 345
D+ YLRDIILN VIAGKDTTA L+WF+YM+CK+P VQEK E AT+
Sbjct: 179 ATTSDSVVDYKYLRDIILNIVIAGKDTTAGALAWFLYMVCKHPEVQEKICHEAMVATSAG 238
Query: 346 TVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGD 405
+S EF+ +TD+AL M+YLHA LTETLRLYP+VP++ K CF+DD LP+G++V KGD
Sbjct: 239 DTASVDEFLQSLTDQALNNMHYLHAALTETLRLYPSVPMENKQCFSDDVLPNGFNVSKGD 298
Query: 406 MVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAY 465
+V + PYAMGRM+ +WG DAE FRP RWLDENG FQ E+PFKFTAFQAGPRICLGKEFAY
Sbjct: 299 IVFFIPYAMGRMESLWGKDAEYFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKEFAY 358
Query: 466 RQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHR 511
RQMKIF+AVLL F KL DEK V+Y+T + L ID GL + A R
Sbjct: 359 RQMKIFAAVLLRFFVLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 404
>ref|NP_910387.1| ESTs AU056036(S20239),C72753(E2173), AU056035(S20239) correspond to
a region of the predicted gene.~Similar to putative
cytochrome P-450 (AC003680) [Oryza sativa (japonica
cultivar-group)]
Length = 403
Score = 451 bits (1161), Expect = e-125
Identities = 224/402 (55%), Positives = 295/402 (72%), Gaps = 8/402 (1%)
Query: 117 LKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNF 176
+KDL GDGIFA DG+ WR QRK++S+EFSTK+LRDFS+ FR+NAAK+A +S AA +
Sbjct: 1 MKDLFGDGIFATDGDMWRHQRKLASYEFSTKVLRDFSSDTFRRNAAKLAEKISCAAANRI 60
Query: 177 KLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKI 236
+ IQDLLM++T+DSIF+V FG ELN++ GS E G F+ AFD A++L YR+VD+ WK+
Sbjct: 61 SINIQDLLMRATMDSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKL 120
Query: 237 KKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKE---- 292
K++LNIGSEA L++N ++++ FV+KLI+ + +QM D K DILSRF+ E
Sbjct: 121 KRYLNIGSEAKLKRNIQIIDSFVMKLIHQKREQMKIAADYKTKE-DILSRFVLASEQDPG 179
Query: 293 -YDTTYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATN-TKTVSSC 350
D YLRDI+LNF+IAGKDTT TL+WF Y+LCK P VQ+K A E+RE +K ++
Sbjct: 180 TMDDRYLRDIVLNFLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDNTI 239
Query: 351 TEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQ 410
F + + A+ KM+YL AT++ETLRLYPAVPVDAK+ DD LP+GY V KGD ++Y
Sbjct: 240 ESFTKRLDEGAISKMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYM 299
Query: 411 PYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKI 470
YAMGRM ++WG+DA+EFRP RWL NG +Q E+PFKF +F AGPRICLGKEFA+RQMKI
Sbjct: 300 IYAMGRMTYLWGEDAQEFRPERWL-VNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKI 358
Query: 471 FSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALHRD 512
+A L+ F+F+L DE + YKTM LHID GL + A R+
Sbjct: 359 MAATLIHFFKFRLEDESKEPIYKTMFTLHIDNGLHLLANPRE 400
>gb|AAG60111.1| cytochrome P450, putative [Arabidopsis thaliana]
Length = 524
Score = 396 bits (1017), Expect = e-108
Identities = 222/504 (44%), Positives = 312/504 (61%), Gaps = 37/504 (7%)
Query: 41 NMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYIL 100
N K P+ G Q+ NF+R+H ++ + RT + PF + Y ++P NVEY+L
Sbjct: 24 NKSGPKTWPLVGAAIEQLTNFDRMHDWLVEYLYNSRTVVVPMPFTTYTYIADPINVEYVL 83
Query: 101 KTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKN 160
KTNF NY KG + ++ LLGDGIF DGE WR+QRK +S EF++K LRDFST +F++
Sbjct: 84 KTNFSNYPKGETYHSYMEVLLGDGIFNSDGELWRKQRKTASFEFASKNLRDFSTVVFKEY 143
Query: 161 AAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDT 220
+ K+ I+S+A+ ++++Q+LLM+ TLDSI +V FG E+ ++ E +FA AFDT
Sbjct: 144 SLKLFTILSQASFKEQQVDMQELLMRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDT 202
Query: 221 ASALTLYRYVDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQM---------N 271
A+ + R++D WK+KKFLNIGSEA L K+ +V+N+F +I R ++ N
Sbjct: 203 ANIIVTLRFIDPLWKMKKFLNIGSEALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNN 262
Query: 272 SKGDSIRKSGDILSRFLQV------KEYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLC 325
+ ++ + DILSRF+++ KE + + LRDI+LNFVIAG+DTTA TL+W +YM+
Sbjct: 263 NNNNNNKVKHDILSRFIEISDDPDSKETEKS-LRDIVLNFVIAGRDTTATTLTWAIYMIM 321
Query: 326 KYPAVQEKAAEEVR-------EATNTKT-----------VSSCTEFVSCVTDEALEKMNY 367
V EK E++ EATNT TEF + ++L K++Y
Sbjct: 322 MNENVAEKLYSELQELEKESAEATNTSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHY 381
Query: 368 LHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEE 427
LHA +TETLRLYPAVP D K DD LP+G VK G MV+Y PY+MGRM++ WG DA
Sbjct: 382 LHAVITETLRLYPAVPQDPKGVLEDDMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAAL 441
Query: 428 FRPARWLDENGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEK 487
F+P RWL ++G FQ +PFKFTAFQAGPRICLGK+ AY QMK+ A+L ++F L
Sbjct: 442 FKPERWL-KDGVFQNASPFKFTAFQAGPRICLGKDSAYLQMKMAMAILCRFYKFHL-VPN 499
Query: 488 RNVTYKTMINLHIDGGLEIKALHR 511
V Y+ M L + GL++ R
Sbjct: 500 HPVKYRMMTILSMAHGLKVTVSRR 523
>gb|AAK52956.1| cytochrome P450-like protein [Zea mays]
Length = 543
Score = 396 bits (1017), Expect = e-108
Identities = 220/529 (41%), Positives = 324/529 (60%), Gaps = 38/529 (7%)
Query: 10 LSNPFLFAALSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMT 69
L+ P + AL L ++L +L+++ + + K + PV G Q+ N++R+H ++
Sbjct: 17 LAGPHKYIAL---LLVVLSWILVQRWSLRKQ--KGPRSWPVIGATVEQLRNYHRMHDWLV 71
Query: 70 DLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVD 129
++RT + PF S Y ++P NVE++LKTNF NY KG+ + LLGDGIF D
Sbjct: 72 GYLSRHRTVTVDMPFTSYTYIADPVNVEHVLKTNFTNYPKGIVYRSYMDVLLGDGIFNAD 131
Query: 130 GEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTL 189
GE WR+QRK +S EF++K LRDFS +FR+ + K++ I+S+A+ + +++Q+L M+ TL
Sbjct: 132 GELWRKQRKTASFEFASKNLRDFSAIVFREYSLKLSGILSQASKAGKVVDMQELYMRMTL 191
Query: 190 DSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAALR 249
DSI +V FG E+ ++ E +FA AFD A+ + R++D W+IK+F ++GSEA L
Sbjct: 192 DSICKVGFGVEIGTLSPDLPEN-SFAQAFDAANIIITLRFIDPLWRIKRFFHVGSEALLA 250
Query: 250 KNTEVLNEFVIKLINTR---IQQMNSKGDSIRKSGDILSRFLQVKEY--------DTTYL 298
++ ++++EF +I R I ++ + G + DILSRF+++ E D L
Sbjct: 251 QSIKLVDEFTYSVIRRRKAEIVEVRASGKQEKMKHDILSRFIELGEAGDDGGGFGDDKSL 310
Query: 299 RDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKA--------AEEVRE---------- 340
RD++LNFVIAG+DTTA TLSWF +M +P V EK AE RE
Sbjct: 311 RDVVLNFVIAGRDTTATTLSWFTHMAMSHPDVAEKLRRELCAFEAERAREEGVTLVLCGG 370
Query: 341 --ATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDG 398
A + + +F +T ++L K+ YLHA +TETLRLYPAVP D K DD LPDG
Sbjct: 371 ADADDKAFAARVAQFAGLLTYDSLGKLVYLHACVTETLRLYPAVPQDPKGILEDDVLPDG 430
Query: 399 YSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTAFQAGPRIC 458
V+ G MV+Y PY+MGRM++ WG DA FRP RW++E+G F+ +PFKFTAFQAGPRIC
Sbjct: 431 TKVRAGGMVTYVPYSMGRMEYNWGPDAASFRPERWINEDGAFRNASPFKFTAFQAGPRIC 490
Query: 459 LGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIK 507
LGK+ AY QMK+ A+L + F+L E V Y+ M L + GL+++
Sbjct: 491 LGKDSAYLQMKMALAILFRFYSFRLL-EGHPVQYRMMTILSMAHGLKVR 538
>ref|XP_470289.1| putative plant cytochrome P-450 protein [Oryza sativa (japonica
cultivar-group)] gi|19071651|gb|AAL84318.1| putative
plant cytochrome P-450 protein [Oryza sativa (japonica
cultivar-group)]
Length = 544
Score = 394 bits (1012), Expect = e-108
Identities = 221/519 (42%), Positives = 317/519 (60%), Gaps = 34/519 (6%)
Query: 19 LSASLTLLLVQLLLRKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTY 78
L A ++L +L+ K + + N K + P+ G Q+ N++R+H ++ + K RT
Sbjct: 25 LIAIFLVVLSWILVHKWSLR--NQKGPRSWPIIGATVEQLKNYHRMHDWLVEYLSKDRTV 82
Query: 79 RLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRK 138
+ PF S Y ++P NVE++LKTNF NY KG + LLGDGIF DGE WR+QRK
Sbjct: 83 TVDMPFTSYTYIADPVNVEHVLKTNFTNYPKGEVYRSYMDVLLGDGIFNADGEMWRKQRK 142
Query: 139 ISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFG 198
+S EF++K LRDFST +FR+ + K+++I+S+A + +++Q+L M+ TLDSI +V FG
Sbjct: 143 TASFEFASKNLRDFSTVVFREYSLKLSSILSQACKAGRVVDMQELFMRMTLDSICKVGFG 202
Query: 199 TELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIGSEAALRKNTEVLNEF 258
E+ ++ E +FA AFD A+ + R++D W++KKFL++GSEA L ++ +++++F
Sbjct: 203 VEIGTLSPDLPE-NSFAQAFDAANIIVTLRFIDPLWRLKKFLHVGSEALLEQSMKLVDDF 261
Query: 259 VIKLINTR---IQQMNSKGDSIRKSGDILSRFLQVKEY----------DTTYLRDIILNF 305
+I R I Q + G + DILSRF+++ E D LRD++LNF
Sbjct: 262 TYSVIRRRKAEILQARASGKQEKIKHDILSRFIELGEAGGDEGGGSFGDDKSLRDVVLNF 321
Query: 306 VIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEV--------RE--------ATNTKTVSS 349
VIAG+DTTA TLSWF YM +PAV +K E+ RE A +
Sbjct: 322 VIAGRDTTATTLSWFTYMAMTHPAVADKLRRELAAFEDERAREEGVALADAAGEASFAAR 381
Query: 350 CTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSY 409
+F S ++ +A+ K+ YLHA +TETLRLYPAVP D K DD LPDG V+ G MV+Y
Sbjct: 382 VAQFASLLSYDAVGKLVYLHACVTETLRLYPAVPQDPKGIVEDDVLPDGTKVRAGGMVTY 441
Query: 410 QPYAMGRMKFIWGDDAEEFRPARWLD-ENGNFQAENPFKFTAFQAGPRICLGKEFAYRQM 468
PY+MGRM++ WG DA FRP RWL + G F+ +PFKFTAFQAGPRICLGK+ AY QM
Sbjct: 442 VPYSMGRMEYNWGPDAASFRPERWLSGDGGAFRNASPFKFTAFQAGPRICLGKDSAYLQM 501
Query: 469 KIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIK 507
K+ A+L + F L E V Y+ M L + GL+++
Sbjct: 502 KMALAILFRFYTFDL-VEDHPVKYRMMTILSMAHGLKVR 539
>dbj|BAC43393.1| unknown protein [Arabidopsis thaliana] gi|30697865|ref|NP_177109.2|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 478
Score = 380 bits (976), Expect = e-104
Identities = 214/481 (44%), Positives = 301/481 (62%), Gaps = 37/481 (7%)
Query: 64 LHHYMTDLARKYRTYRLLNPFRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGD 123
+H ++ + RT + PF + Y ++P NVEY+LKTNF NY KG + ++ LLGD
Sbjct: 1 MHDWLVEYLYNSRTVVVPMPFTTYTYIADPINVEYVLKTNFSNYPKGETYHSYMEVLLGD 60
Query: 124 GIFAVDGEKWREQRKISSHEFSTKMLRDFSTSIFRKNAAKVANIVSEAATSNFKLEIQDL 183
GIF DGE WR+QRK +S EF++K LRDFST +F++ + K+ I+S+A+ ++++Q+L
Sbjct: 61 GIFNSDGELWRKQRKTASFEFASKNLRDFSTVVFKEYSLKLFTILSQASFKEQQVDMQEL 120
Query: 184 LMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVFWKIKKFLNIG 243
LM+ TLDSI +V FG E+ ++ E +FA AFDTA+ + R++D WK+KKFLNIG
Sbjct: 121 LMRMTLDSICKVGFGVEIGTLAPELPE-NHFAKAFDTANIIVTLRFIDPLWKMKKFLNIG 179
Query: 244 SEAALRKNTEVLNEFVIKLINTRIQQM---------NSKGDSIRKSGDILSRFLQV---- 290
SEA L K+ +V+N+F +I R ++ N+ ++ + DILSRF+++
Sbjct: 180 SEALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDP 239
Query: 291 --KEYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVR-------EA 341
KE + + LRDI+LNFVIAG+DTTA TL+W +YM+ V EK E++ EA
Sbjct: 240 DSKETEKS-LRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEA 298
Query: 342 TNTKT-----------VSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAKICF 390
TNT TEF + ++L K++YLHA +TETLRLYPAVP D K
Sbjct: 299 TNTSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVL 358
Query: 391 ADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDENGNFQAENPFKFTA 450
DD LP+G VK G MV+Y PY+MGRM++ WG DA F+P RWL ++G FQ +PFKFTA
Sbjct: 359 EDDMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWL-KDGVFQNASPFKFTA 417
Query: 451 FQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEIKALH 510
FQAGPRICLGK+ AY QMK+ A+L ++F L V Y+ M L + GL++
Sbjct: 418 FQAGPRICLGKDSAYLQMKMAMAILCRFYKFHL-VPNHPVKYRMMTILSMAHGLKVTVSR 476
Query: 511 R 511
R
Sbjct: 477 R 477
>emb|CAB88066.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|15228981|ref|NP_191222.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|11249683|pir||T49064
cytochrome P450-like protein - Arabidopsis thaliana
Length = 499
Score = 333 bits (854), Expect = 8e-90
Identities = 195/487 (40%), Positives = 288/487 (59%), Gaps = 24/487 (4%)
Query: 37 NKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLL--NPFRSE-VYTSEP 93
N S+ K Y P+ G++ + N +R + + + T + P + + V T+ P
Sbjct: 24 NSSSEFGFKSY-PIVGSLPGLVNNRHRFLDWTVETLSRCPTQTAIFRRPGKLQFVMTANP 82
Query: 94 SNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFS 153
+NVEY+LKT FE++ KG L+D LG GIF DGE W +QRK +S+EFSTK LRDF
Sbjct: 83 ANVEYMLKTKFESFPKGERFISILEDFLGRGIFNSDGEMWWKQRKTASYEFSTKSLRDFV 142
Query: 154 TS-IFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGK 212
S + + ++ +++EAAT+ +++QD+L + D+I ++AF + + G
Sbjct: 143 MSNVTVEINTRLVPVLAEAATNGKLIDLQDILERFAFDNICKLAFNVDSACLGDDGAAGV 202
Query: 213 NFANAFDTASALTLYRYVDVF---WKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQ 269
NF AF+TA+ + R+ V WKIKK LNIGSE LR++ ++++F +++ RI+Q
Sbjct: 203 NFMQAFETAATIISQRFQSVISYSWKIKKKLNIGSERVLRESIMIVHKFADEIVRNRIEQ 262
Query: 270 MNSKGDSIRKSGDILSRFLQVKEYDTT-YLRDIILNFVIAGKDTTAATLSWFMYMLCKYP 328
G D+LSRF+ +E ++ LRDI+++F++AG+DTT++ LSWF ++L +P
Sbjct: 263 ----GKVSDHKEDLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFFWLLSMHP 318
Query: 329 AVQEKAAEE---VREATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVD 385
V++K +E +RE T K + F E L+ MNYLHA +TE+LRLYP VPVD
Sbjct: 319 EVKDKILQELNSIRERTG-KRIGEVYGF------EDLKLMNYLHAAITESLRLYPPVPVD 371
Query: 386 AKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDE-NGNFQAEN 444
C D+ LPDG + K +SY YAMGRM+ IWG D + F P RW+DE NG F+ EN
Sbjct: 372 TMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMESIWGKDCDRFDPERWIDETNGGFRGEN 431
Query: 445 PFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGL 504
P+KF AF AGPR+CLGKE AY QMK A +L F ++ +K + L I GGL
Sbjct: 432 PYKFPAFHAGPRMCLGKEMAYIQMKSIVAAVLERFVVEVPGKKERPEILMSVTLRIRGGL 491
Query: 505 EIKALHR 511
++ R
Sbjct: 492 NVRVQER 498
>gb|AAL54887.1| cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 511
Score = 325 bits (832), Expect = 3e-87
Identities = 188/486 (38%), Positives = 292/486 (59%), Gaps = 39/486 (8%)
Query: 19 LSASLTLLLVQLLL-----------RKLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHY 67
LS SL LV LL L + +N+ K + +P+ G+ F+ + N +R +
Sbjct: 8 LSTSLLFCLVPLLFLFFVKFKKTITNTLLSSNNSSKIPRSYPLIGSYFSILANHDRRIKW 67
Query: 68 MTD--LARKYRTYRLLNP--FRSEVYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGD 123
++D L T+ L+ P FR+ ++T+ PSNV+++LKTNF+ Y KG +Y LKD L +
Sbjct: 68 ISDIILTTPNLTFTLIRPLNFRT-IFTANPSNVQHVLKTNFQVYQKGHGSYSTLKDFLSN 126
Query: 124 GIFAVDGEKWREQRKISSHEFSTKMLRDF-STSIFRKNAAKVANIVSEAATSNFKLEIQD 182
GIF VDG+ W+ QR+++SHEF+T+ LR F T + + + ++ I++ AA + L+ QD
Sbjct: 127 GIFNVDGDIWKYQRQVASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQD 186
Query: 183 LLMKSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVF---WKIKKF 239
+L + D+I ++AFG + + S E + FA AF+ A L+ R++ F WKIK+
Sbjct: 187 ILQRFAFDNICKIAFGYDPGYLLPSLPEAE-FAVAFEDAVRLSTERFIVPFSLIWKIKRA 245
Query: 240 LNIGSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYDTTYLR 299
LNIGSE LR E + EF +++ + +++N K S S D+LSRFL D ++
Sbjct: 246 LNIGSEKKLRVAVEQVREFAKEIVREKQKELNDK--SSLDSADLLSRFLSTGHSDEDFVT 303
Query: 300 DIILNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTD 359
DI+++F++AG+DTT+A L+WF +++ K+P V+ + +EV E + S +
Sbjct: 304 DIVISFILAGRDTTSAALTWFFWLISKHPEVESQIMKEVGEKSE-----------SLLLY 352
Query: 360 EALEKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKF 419
+ ++ M Y HA+L E++R YP VP+D+K DD LPDG VKKG V+Y PYAMGR++
Sbjct: 353 DEVKNMMYTHASLCESMRFYPPVPMDSKEATKDDILPDGTFVKKGTRVTYHPYAMGRVEK 412
Query: 420 IWGDDAEEFRPARWLDE-----NGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAV 474
+WG+D EF+P RWLD+ N F ++ + + FQAGPRICLGKE A+ QMK A
Sbjct: 413 VWGEDWAEFKPERWLDKDEVTGNWTFVPKDAYTYPVFQAGPRICLGKEMAFLQMKRVVAG 472
Query: 475 LLGCFR 480
+L F+
Sbjct: 473 VLRRFK 478
>gb|AAF79271.1| F12K21.15 [Arabidopsis thaliana] gi|15218671|ref|NP_174713.1|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 498
Score = 320 bits (819), Expect = 9e-86
Identities = 190/485 (39%), Positives = 286/485 (58%), Gaps = 22/485 (4%)
Query: 38 KSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKYRTYRLL--NPFRSE-VYTSEPS 94
KS++ K +P+ G+ + N +R + + + T + P + + + T+ PS
Sbjct: 24 KSSSEFGFKSYPIVGSFPGLVNNRHRFLDWTVETLSRCPTQTAIFRRPGKQQLIMTANPS 83
Query: 95 NVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLRDFST 154
NVEY+LKT FE++ KG L+D LG GIF DG+ W +QRK +S+EFSTK LRDF
Sbjct: 84 NVEYMLKTKFESFPKGQQFTSVLEDFLGHGIFNSDGDMWWKQRKTASYEFSTKSLRDFVM 143
Query: 155 S-IFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSEEGKN 213
S + + ++ ++ EAAT+ +++QD+L + D+I ++AF + + G N
Sbjct: 144 SNVTVEINTRLVPVLVEAATTGKLIDLQDILERFAFDNICKLAFNVDCACLGHDGAVGVN 203
Query: 214 FANAFDTASALTLYRYVDVF---WKIKKFLNIGSEAALRKNTEVLNEFVIKLINTRIQQM 270
F AF+TA+ + R+ V W+IKK LNIGSE LR++ +++F +++ RI Q
Sbjct: 204 FMRAFETAATIISQRFRSVASCAWRIKKKLNIGSERVLRESIATVHKFADEIVRNRIDQ- 262
Query: 271 NSKGDSIRKSGDILSRFLQVKEYDTT-YLRDIILNFVIAGKDTTAATLSWFMYMLCKYPA 329
G S D+LSRF+ +E ++ LRDI+++F++AG+DTT++ LSWF ++L +P
Sbjct: 263 ---GRSSDHKEDLLSRFISKEEMNSPEILRDIVISFILAGRDTTSSALSWFFWLLSMHPE 319
Query: 330 VQEKAAEEVRE--ATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVDAK 387
V++K +E+ A K + F E L+ MNYLHA +TE+LRLYP VPVD K
Sbjct: 320 VEDKILQELNSIRARTGKRIGEVYGF------EHLKMMNYLHAAITESLRLYPPVPVDIK 373
Query: 388 ICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWLDE-NGNFQAENPF 446
C D+ LPDG V KG ++Y +AMGRM+ IWG D + F P RW+DE NG F+ E+P
Sbjct: 374 SCAEDNVLPDGTFVGKGWAITYNIFAMGRMESIWGKDCDRFDPERWIDETNGCFRGEDPS 433
Query: 447 KFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFRFKLNDEKRNVTYKTMINLHIDGGLEI 506
KF AF AGPR+C+GK+ AY QMK A +L F ++ ++R +M L I GGL
Sbjct: 434 KFPAFHAGPRMCVGKDMAYIQMKSIVAAVLERFVVEVPGKERPEILLSM-TLRIKGGLFA 492
Query: 507 KALHR 511
+ R
Sbjct: 493 RVQER 497
>sp|P98188|C94A2_VICSA Cytochrome P450 94A2 (P450-dependent fatty acid omega-hydroxylase)
gi|11245504|gb|AAG33645.1| cytochrome P450-dependent
fatty acid hydroxylase [Vicia sativa]
Length = 513
Score = 316 bits (810), Expect = 1e-84
Identities = 178/459 (38%), Positives = 278/459 (59%), Gaps = 25/459 (5%)
Query: 34 KLNNKSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMTDLARKY--RTYRLLNPFRS-EVYT 90
K + + N K +P+ G+ F+ + NF+R + +D+ + T+ L PF + +V+T
Sbjct: 32 KTPSSTTNTPIPKSYPIFGSAFSLLANFHRRIQWTSDILQTIPSSTFVLHRPFGARQVFT 91
Query: 91 SEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIFAVDGEKWREQRKISSHEFSTKMLR 150
++P+ V++IL+TNF YGKGL YQ++ D LGDGIF DGE W+ QR+ISSHEF+T+ LR
Sbjct: 92 AQPAVVQHILRTNFTCYGKGLTFYQSINDFLGDGIFNADGESWKFQRQISSHEFNTRSLR 151
Query: 151 DF-STSIFRKNAAKVANIVSEAATSNFKLEIQDLLMKSTLDSIFQVAFGTELNSMCGSSE 209
F T + + + ++ ++S+A+ S L+ QD+L + T D+I +AFG + + S
Sbjct: 152 KFVETVVDVELSDRLVPVLSQASNSQTTLDFQDILQRLTFDNICMIAFGYDPEYLLPSLP 211
Query: 210 EGKNFANAFDTASALTLYRY---VDVFWKIKKFLNIGSEAALRKNTEVLNEFVIKLINTR 266
E FA AFD +S L++ R + + WK+K+FLNIG E L++ + K++ +
Sbjct: 212 EIP-FAKAFDESSQLSIERLNALIPLLWKVKRFLNIGVERQLKEAVAEVRGLATKIVKNK 270
Query: 267 IQQMNSKG-DSIRKSGDILSRFLQVKEYDTTYLRDIILNFVIAGKDTTAATLSWFMYMLC 325
+++ K S +S D+LSRFL D +++ D++++ ++AG+DTT+A L+WF ++L
Sbjct: 271 KKELKEKALQSESESVDLLSRFLSSGHSDESFVTDMVISIILAGRDTTSAALTWFFWLLS 330
Query: 326 KYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTDEALEKMNYLHATLTETLRLYPAVPVD 385
K+ V+ + +E+ + T V + ++ M Y HA L E++RLYP +PVD
Sbjct: 331 KHSHVENEILKEITGKSET------------VGYDEVKDMVYTHAALCESMRLYPPLPVD 378
Query: 386 AKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWGDDAEEFRPARWL--DENG--NFQ 441
K+ DD LPDG VKKG V+Y YAMGR + IWG D EFRP RWL DE G +F
Sbjct: 379 TKVAVHDDVLPDGTLVKKGWRVTYHIYAMGRSEKIWGPDWAEFRPERWLSRDEVGKWSFV 438
Query: 442 AENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLGCFR 480
+ + + FQAGPR+C+GKE A+ QMK A ++G FR
Sbjct: 439 GIDYYSYPVFQAGPRVCIGKEMAFLQMKRVVAGIMGRFR 477
>gb|AAL54884.1| cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 510
Score = 314 bits (804), Expect = 5e-84
Identities = 182/483 (37%), Positives = 287/483 (58%), Gaps = 36/483 (7%)
Query: 20 SASLTLLLVQLLLR---KLNN-------KSNNMKKKKYHPVAGTVFNQMMNFNRLHHYMT 69
S SL LV LL K N SN+ K + +P+ G+ F+ + N ++ +++
Sbjct: 9 STSLLFCLVPLLFLFFIKFNKTITNTLLSSNSSKIPRSYPLIGSYFSILANHDQRIKWIS 68
Query: 70 D--LARKYRTYRLLNPFRSE-VYTSEPSNVEYILKTNFENYGKGLYNYQNLKDLLGDGIF 126
D L+ T+ L+ P ++T+ PSNV+++LKTNF+ Y KG + LKD L +GIF
Sbjct: 69 DIILSTPNLTFTLIRPLNFHTIFTANPSNVQHMLKTNFQVYQKGHNSNTTLKDFLSNGIF 128
Query: 127 AVDGEKWREQRKISSHEFSTKMLRDF-STSIFRKNAAKVANIVSEAATSNFKLEIQDLLM 185
VDG+ W+ QR+++SHEF+T+ LR F T + + + ++ I++ AA + L+ QD+L
Sbjct: 129 NVDGDIWKYQRQVASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQDILQ 188
Query: 186 KSTLDSIFQVAFGTELNSMCGSSEEGKNFANAFDTASALTLYRYVDVF---WKIKKFLNI 242
+ D+I ++AFG + + S E + FA AF+ A L+ R++ F WK+K+ LNI
Sbjct: 189 RFAFDNICKIAFGYDPGYLLPSLPEAE-FAVAFEDAVRLSTERFILPFPLIWKMKRALNI 247
Query: 243 GSEAALRKNTEVLNEFVIKLINTRIQQMNSKGDSIRKSGDILSRFLQVKEYDTTYLRDII 302
GSE LR E + EF +++ + +++ K S S D+LSRFL D ++ DI+
Sbjct: 248 GSEKKLRFAVEQVREFAKEIVREKQRELKDK--SSLDSADLLSRFLSTGHSDENFVTDIV 305
Query: 303 LNFVIAGKDTTAATLSWFMYMLCKYPAVQEKAAEEVREATNTKTVSSCTEFVSCVTDEAL 362
++F++AG+DTT+A L+WF +++ K+P V+ + +E+ E + S + + +
Sbjct: 306 ISFILAGRDTTSAALTWFFWLISKHPEVESQILKEIGEKSE-----------SLLLYDEV 354
Query: 363 EKMNYLHATLTETLRLYPAVPVDAKICFADDTLPDGYSVKKGDMVSYQPYAMGRMKFIWG 422
+ M Y HA+L E++R YP VP+D K DD LPDG VKKG+ V+Y PYAMGR++ +WG
Sbjct: 355 KNMIYTHASLCESMRFYPPVPMDTKEATKDDILPDGTFVKKGNRVTYHPYAMGRVEKVWG 414
Query: 423 DDAEEFRPARWLDE-----NGNFQAENPFKFTAFQAGPRICLGKEFAYRQMKIFSAVLLG 477
D EFRP RWLD+ N F +++ + + FQAGPR+CLGKE A+ QMK A +L
Sbjct: 415 KDWAEFRPERWLDKDEVTGNWTFVSKDAYTYPVFQAGPRVCLGKEMAFLQMKRVVAGVLR 474
Query: 478 CFR 480
F+
Sbjct: 475 RFK 477
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 810,560,021
Number of Sequences: 2540612
Number of extensions: 33645806
Number of successful extensions: 93254
Number of sequences better than 10.0: 4851
Number of HSP's better than 10.0 without gapping: 2361
Number of HSP's successfully gapped in prelim test: 2490
Number of HSP's that attempted gapping in prelim test: 80854
Number of HSP's gapped (non-prelim): 5581
length of query: 512
length of database: 863,360,394
effective HSP length: 132
effective length of query: 380
effective length of database: 527,999,610
effective search space: 200639851800
effective search space used: 200639851800
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 77 (34.3 bits)
Medicago: description of AC146342.7


