

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.2 - phase: 0 /pseudo
(875 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
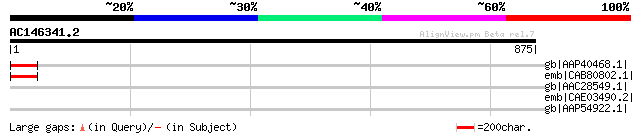
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP40468.1| unknown protein [Arabidopsis thaliana] gi|4256619... 57 3e-06
emb|CAB80802.1| putative protein [Arabidopsis thaliana] gi|60498... 57 3e-06
gb|AAC28549.1| hypothetical protein [Arabidopsis thaliana] gi|15... 48 0.002
emb|CAE03490.2| OSJNBa0065O17.15 [Oryza sativa (japonica cultiva... 39 0.75
gb|AAP54922.1| unknown protein [Oryza sativa (japonica cultivar-... 37 3.7
>gb|AAP40468.1| unknown protein [Arabidopsis thaliana]
gi|42566192|ref|NP_191954.3| expressed protein
[Arabidopsis thaliana]
Length = 831
Score = 56.6 bits (135), Expect = 3e-06
Identities = 25/47 (53%), Positives = 35/47 (74%), Gaps = 1/47 (2%)
Query: 1 MAKKSQRRAV-RYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSS 46
MAKK+QRRA R E+++ GCMWGF++M FRHG KL+ D++ +S
Sbjct: 1 MAKKTQRRAAARQEREQLGCMWGFMNMFSFRHGPLAHKLLLDQKHAS 47
>emb|CAB80802.1| putative protein [Arabidopsis thaliana] gi|6049884|gb|AAF02799.1|
F5I10.23 gene product [Arabidopsis thaliana]
gi|2252842|gb|AAB62841.1| A_IG005I10.23 gene product
[Arabidopsis thaliana] gi|7485266|pir||T01527
hypothetical protein A_IG005I10.23 - Arabidopsis
thaliana
Length = 770
Score = 56.6 bits (135), Expect = 3e-06
Identities = 25/47 (53%), Positives = 35/47 (74%), Gaps = 1/47 (2%)
Query: 1 MAKKSQRRAV-RYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSS 46
MAKK+QRRA R E+++ GCMWGF++M FRHG KL+ D++ +S
Sbjct: 1 MAKKTQRRAAARQEREQLGCMWGFMNMFSFRHGPLAHKLLLDQKHAS 47
>gb|AAC28549.1| hypothetical protein [Arabidopsis thaliana]
gi|15225907|ref|NP_182114.1| expressed protein
[Arabidopsis thaliana] gi|7486456|pir||T02457
hypothetical protein At2g45900 [imported] - Arabidopsis
thaliana
Length = 720
Score = 47.8 bits (112), Expect = 0.002
Identities = 23/47 (48%), Positives = 32/47 (67%), Gaps = 2/47 (4%)
Query: 1 MAKKSQRRAVRYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSSK 47
M K+ RR + ++ GC+W F+SM DFRHG S +KL+ DK+R SK
Sbjct: 1 MEKEICRRHL--SREGLGCVWVFMSMFDFRHGGSNQKLLMDKKRRSK 45
>emb|CAE03490.2| OSJNBa0065O17.15 [Oryza sativa (japonica cultivar-group)]
gi|50927901|ref|XP_473478.1| OSJNBa0065O17.15 [Oryza
sativa (japonica cultivar-group)]
Length = 859
Score = 38.9 bits (89), Expect = 0.75
Identities = 26/54 (48%), Positives = 32/54 (59%), Gaps = 5/54 (9%)
Query: 1 MAKKSQRRAVRYEKDKAGCMWGFISMLDFRHGHSTRKLIADK--RRSSKHSEGK 52
MAKKS ++ + EK GCM G I M DFR + KLI+D RRSS S+ K
Sbjct: 1 MAKKSYKQRCKSEKVHMGCMSGLIHMFDFRR---SPKLISDGTIRRSSVRSDLK 51
>gb|AAP54922.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37536666|ref|NP_922635.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|13876517|gb|AAK43493.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 901
Score = 36.6 bits (83), Expect = 3.7
Identities = 16/49 (32%), Positives = 29/49 (58%), Gaps = 2/49 (4%)
Query: 1 MAKKSQRRAVRYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSSKHS 49
M ++S +R+ E++ GC+WG + ML FR + L+ K+ S +H+
Sbjct: 1 MGRRSHKRSTEQEENNVGCVWGLMRMLYFR--RDAKFLLDTKQVSRRHT 47
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.374 0.166 0.667
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,135,494,641
Number of Sequences: 2540612
Number of extensions: 37465452
Number of successful extensions: 176262
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 176257
Number of HSP's gapped (non-prelim): 5
length of query: 875
length of database: 863,360,394
effective HSP length: 137
effective length of query: 738
effective length of database: 515,296,550
effective search space: 380288853900
effective search space used: 380288853900
T: 11
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 80 (35.4 bits)
Medicago: description of AC146341.2


