

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.8 + phase: 0
(402 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
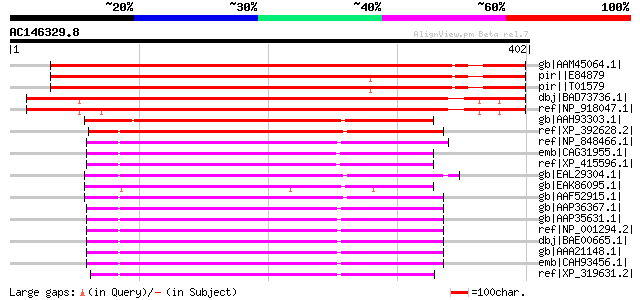
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM45064.1| putative heme A [Arabidopsis thaliana] gi|1502829... 482 e-134
pir||E84879 probable heme A farnesyltransferase [imported] - Ara... 476 e-133
pir||T01579 heme A farnesyltransferase homolog F16B22.1 - Arabid... 476 e-133
dbj|BAD73736.1| putative heme A:farnesyltransferase [Oryza sativ... 464 e-129
ref|NP_918047.1| putative heme A farnesyltransferase homolog [Or... 455 e-126
gb|AAH93303.1| LOC553384 protein [Danio rerio] 205 2e-51
ref|XP_392628.2| PREDICTED: similar to GA18613-PA [Apis mellifera] 205 2e-51
ref|NP_848466.1| COX10 homolog, cytochrome c oxidase assembly pr... 200 7e-50
emb|CAG31955.1| hypothetical protein [Gallus gallus] 200 7e-50
ref|XP_415596.1| PREDICTED: similar to COX10 homolog, cytochrome... 200 7e-50
gb|EAL29304.1| GA18613-PA [Drosophila pseudoobscura] 199 2e-49
gb|EAK86095.1| hypothetical protein UM05692.1 [Ustilago maydis 5... 197 6e-49
gb|AAF52915.1| CG5037-PA [Drosophila melanogaster] gi|19921064|r... 196 1e-48
gb|AAP36367.1| Homo sapiens COX10 homolog, cytochrome c oxidase ... 194 3e-48
gb|AAP35631.1| COX10 homolog, cytochrome c oxidase assembly prot... 194 3e-48
ref|NP_001294.2| heme A:farnesyltransferase [Homo sapiens] 193 9e-48
dbj|BAE00665.1| unnamed protein product [Macaca fascicularis] 192 2e-47
gb|AAA21148.1| heme A:farnesyltransferase 190 6e-47
emb|CAH93456.1| hypothetical protein [Pongo pygmaeus] 190 6e-47
ref|XP_319631.2| ENSANGP00000013941 [Anopheles gambiae str. PEST... 187 4e-46
>gb|AAM45064.1| putative heme A [Arabidopsis thaliana] gi|15028299|gb|AAK76626.1|
putative heme A:farnesyltransferase [Arabidopsis
thaliana] gi|20197183|gb|AAM14960.1| putative heme
A:farnesyltransferase [Arabidopsis thaliana]
gi|20197028|gb|AAC27454.3| putative heme
A:farnesyltransferase [Arabidopsis thaliana]
gi|18406556|ref|NP_566019.1| UbiA prenyltransferase
family protein [Arabidopsis thaliana]
Length = 431
Score = 482 bits (1240), Expect = e-134
Identities = 242/370 (65%), Positives = 291/370 (78%), Gaps = 14/370 (3%)
Query: 32 STFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLLVVATSGTGFVLGSGGA-VDLS 90
+T +++ +S+ L L HY CY +LSKA+LS+LVVATSGTG++LG+G A +
Sbjct: 71 ATATATTGEISSRVAALAGLGHHYARCYWELSKAKLSMLVVATSGTGYILGTGNAAISFP 130
Query: 91 MLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAG 150
L YTC GTMM+AASA++LNQ+FEI ND+ M+RT RPLPSGRI+VPHAV WA+ G +G
Sbjct: 131 GLCYTCAGTMMIAASANSLNQIFEISNDSKMKRTMLRPLPSGRISVPHAVAWATIAGASG 190
Query: 151 TALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDI 210
LLA++TNMLAAGLA++NLVLY+FVYTPLKQ+HP+NTWVGAVVGAIPPLLGWAAASG I
Sbjct: 191 ACLLASKTNMLAAGLASANLVLYAFVYTPLKQLHPINTWVGAVVGAIPPLLGWAAASGQI 250
Query: 211 SLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLI 270
S NS+ILPAALYFWQIPHFMALA++CR DYAAGG+KM SL DPSG++ A VA+RN Y+I
Sbjct: 251 SYNSMILPAALYFWQIPHFMALAHLCRNDYAAGGYKMLSLFDPSGKRIAAVALRNCFYMI 310
Query: 271 PLGFLAYDWGVTSGWFCLESTALTLAISAAAFSFYRDRTKERARRMFHASLLYLPVFMAG 330
PLGF+AYDWG+TS WFCLEST LTLAI+A AFSFYRDRT +AR+MFHASLL+LPVFM+G
Sbjct: 311 PLGFIAYDWGLTSSWFCLESTLLTLAIAATAFSFYRDRTMHKARKMFHASLLFLPVFMSG 370
Query: 331 LLIHR-RTDNQHFLEVNAEGFMKSPSSVETSEMDDKNGNQKIKGRQGTRARPPVAYASIA 389
LL+HR DNQ L V G S S G K + R+ A+PPVAYAS A
Sbjct: 371 LLLHRVSNDNQQQL-VEEAGLTNSVS-----------GEVKTQRRKKRVAQPPVAYASAA 418
Query: 390 PFPFLPAPSY 399
PFPFLPAPS+
Sbjct: 419 PFPFLPAPSF 428
>pir||E84879 probable heme A farnesyltransferase [imported] - Arabidopsis
thaliana
Length = 434
Score = 476 bits (1226), Expect = e-133
Identities = 242/373 (64%), Positives = 291/373 (77%), Gaps = 17/373 (4%)
Query: 32 STFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLLVVATSGTGFVLGSGGA-VDLS 90
+T +++ +S+ L L HY CY +LSKA+LS+LVVATSGTG++LG+G A +
Sbjct: 71 ATATATTGEISSRVAALAGLGHHYARCYWELSKAKLSMLVVATSGTGYILGTGNAAISFP 130
Query: 91 MLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAG 150
L YTC GTMM+AASA++LNQ+FEI ND+ M+RT RPLPSGRI+VPHAV WA+ G +G
Sbjct: 131 GLCYTCAGTMMIAASANSLNQIFEISNDSKMKRTMLRPLPSGRISVPHAVAWATIAGASG 190
Query: 151 TALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDI 210
LLA++TNMLAAGLA++NLVLY+FVYTPLKQ+HP+NTWVGAVVGAIPPLLGWAAASG I
Sbjct: 191 ACLLASKTNMLAAGLASANLVLYAFVYTPLKQLHPINTWVGAVVGAIPPLLGWAAASGQI 250
Query: 211 SLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLI 270
S NS+ILPAALYFWQIPHFMALA++CR DYAAGG+KM SL DPSG++ A VA+RN Y+I
Sbjct: 251 SYNSMILPAALYFWQIPHFMALAHLCRNDYAAGGYKMLSLFDPSGKRIAAVALRNCFYMI 310
Query: 271 PLGFLAYD---WGVTSGWFCLESTALTLAISAAAFSFYRDRTKERARRMFHASLLYLPVF 327
PLGF+AYD WG+TS WFCLEST LTLAI+A AFSFYRDRT +AR+MFHASLL+LPVF
Sbjct: 311 PLGFIAYDCESWGLTSSWFCLESTLLTLAIAATAFSFYRDRTMHKARKMFHASLLFLPVF 370
Query: 328 MAGLLIHR-RTDNQHFLEVNAEGFMKSPSSVETSEMDDKNGNQKIKGRQGTRARPPVAYA 386
M+GLL+HR DNQ L V G S S G K + R+ A+PPVAYA
Sbjct: 371 MSGLLLHRVSNDNQQQL-VEEAGLTNSVS-----------GEVKTQRRKKRVAQPPVAYA 418
Query: 387 SIAPFPFLPAPSY 399
S APFPFLPAPS+
Sbjct: 419 SAAPFPFLPAPSF 431
>pir||T01579 heme A farnesyltransferase homolog F16B22.1 - Arabidopsis thaliana
(fragment)
Length = 469
Score = 476 bits (1226), Expect = e-133
Identities = 242/373 (64%), Positives = 291/373 (77%), Gaps = 17/373 (4%)
Query: 32 STFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLLVVATSGTGFVLGSGGA-VDLS 90
+T +++ +S+ L L HY CY +LSKA+LS+LVVATSGTG++LG+G A +
Sbjct: 106 ATATATTGEISSRVAALAGLGHHYARCYWELSKAKLSMLVVATSGTGYILGTGNAAISFP 165
Query: 91 MLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAG 150
L YTC GTMM+AASA++LNQ+FEI ND+ M+RT RPLPSGRI+VPHAV WA+ G +G
Sbjct: 166 GLCYTCAGTMMIAASANSLNQIFEISNDSKMKRTMLRPLPSGRISVPHAVAWATIAGASG 225
Query: 151 TALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDI 210
LLA++TNMLAAGLA++NLVLY+FVYTPLKQ+HP+NTWVGAVVGAIPPLLGWAAASG I
Sbjct: 226 ACLLASKTNMLAAGLASANLVLYAFVYTPLKQLHPINTWVGAVVGAIPPLLGWAAASGQI 285
Query: 211 SLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLI 270
S NS+ILPAALYFWQIPHFMALA++CR DYAAGG+KM SL DPSG++ A VA+RN Y+I
Sbjct: 286 SYNSMILPAALYFWQIPHFMALAHLCRNDYAAGGYKMLSLFDPSGKRIAAVALRNCFYMI 345
Query: 271 PLGFLAYD---WGVTSGWFCLESTALTLAISAAAFSFYRDRTKERARRMFHASLLYLPVF 327
PLGF+AYD WG+TS WFCLEST LTLAI+A AFSFYRDRT +AR+MFHASLL+LPVF
Sbjct: 346 PLGFIAYDCESWGLTSSWFCLESTLLTLAIAATAFSFYRDRTMHKARKMFHASLLFLPVF 405
Query: 328 MAGLLIHR-RTDNQHFLEVNAEGFMKSPSSVETSEMDDKNGNQKIKGRQGTRARPPVAYA 386
M+GLL+HR DNQ L V G S S G K + R+ A+PPVAYA
Sbjct: 406 MSGLLLHRVSNDNQQQL-VEEAGLTNSVS-----------GEVKTQRRKKRVAQPPVAYA 453
Query: 387 SIAPFPFLPAPSY 399
S APFPFLPAPS+
Sbjct: 454 SAAPFPFLPAPSF 466
>dbj|BAD73736.1| putative heme A:farnesyltransferase [Oryza sativa (japonica
cultivar-group)] gi|56201999|dbj|BAD73506.1| putative
heme A:farnesyltransferase [Oryza sativa (japonica
cultivar-group)]
Length = 435
Score = 464 bits (1195), Expect = e-129
Identities = 240/401 (59%), Positives = 299/401 (73%), Gaps = 26/401 (6%)
Query: 14 RLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLA---RHYGNCYSQLSKARLSLL 70
R +SSSSSS + + + +A+ S S++ RHYG CY +LSKARLS L
Sbjct: 38 RSSSSSSSSSAPTIAAAVPAVAEATAAAAAVSRQAGSVSDALRHYGRCYFELSKARLSAL 97
Query: 71 VVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMRRTSQRPLP 130
VVATSG G+VLGSG VD++ L TC GTMMVAASA+TLNQVFEIKNDA M+RT +RPLP
Sbjct: 98 VVATSGAGYVLGSGNMVDIAGLCCTCAGTMMVAASANTLNQVFEIKNDAKMKRTMRRPLP 157
Query: 131 SGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQIHPVNTWV 190
SGRI+ HA WA+SVG+AGTALLA + N LAAGLAASNL+LY+FVYTPLKQIHPVNTWV
Sbjct: 158 SGRISPAHAAMWATSVGVAGTALLAWKANGLAAGLAASNLILYAFVYTPLKQIHPVNTWV 217
Query: 191 GAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAGGFKMYSL 250
GAVVGAIPPLLGWAAAS ++SLN++ILPAALY+WQIPHFMALAY+CR DY AGG++M+S
Sbjct: 218 GAVVGAIPPLLGWAAASSELSLNAMILPAALYYWQIPHFMALAYLCRNDYLAGGYRMFSF 277
Query: 251 ADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAAFSFYRDRTK 310
ADP+G++TA V++RN +Y++PLGF AY+WG+TS WF LE++ LTL ++ A SF + T
Sbjct: 278 ADPTGKRTAWVSLRNCLYMLPLGFFAYNWGLTSEWFSLEASLLTLGLTIGALSFVLEPTP 337
Query: 311 ERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNAEGFMKSPSSVETSEM-------- 362
+ ARRMF+ SLLYLP FMAGLL+HR + Q K + +TSE+
Sbjct: 338 KTARRMFYGSLLYLPAFMAGLLLHRLPNEQ-----------KEHNVTQTSEITGILYGAE 386
Query: 363 --DDKNGNQKIKGRQGTR--ARPPVAYASIAPFPFLPAPSY 399
D++ QK + R+ +R +RPPVAYAS+APFPFLP P Y
Sbjct: 387 QQDEERARQKREDRKPSRIHSRPPVAYASVAPFPFLPVPIY 427
>ref|NP_918047.1| putative heme A farnesyltransferase homolog [Oryza sativa (japonica
cultivar-group)]
Length = 449
Score = 455 bits (1171), Expect = e-126
Identities = 240/415 (57%), Positives = 299/415 (71%), Gaps = 40/415 (9%)
Query: 14 RLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLA---RHYGNCYSQLSKARLSL- 69
R +SSSSSS + + + +A+ S S++ RHYG CY +LSKARLS
Sbjct: 38 RSSSSSSSSSAPTIAAAVPAVAEATAAAAAVSRQAGSVSDALRHYGRCYFELSKARLSFQ 97
Query: 70 -------------LVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIK 116
LVVATSG G+VLGSG VD++ L TC GTMMVAASA+TLNQVFEIK
Sbjct: 98 GCISNNSVNCRRALVVATSGAGYVLGSGNMVDIAGLCCTCAGTMMVAASANTLNQVFEIK 157
Query: 117 NDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFV 176
NDA M+RT +RPLPSGRI+ HA WA+SVG+AGTALLA + N LAAGLAASNL+LY+FV
Sbjct: 158 NDAKMKRTMRRPLPSGRISPAHAAMWATSVGVAGTALLAWKANGLAAGLAASNLILYAFV 217
Query: 177 YTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMC 236
YTPLKQIHPVNTWVGAVVGAIPPLLGWAAAS ++SLN++ILPAALY+WQIPHFMALAY+C
Sbjct: 218 YTPLKQIHPVNTWVGAVVGAIPPLLGWAAASSELSLNAMILPAALYYWQIPHFMALAYLC 277
Query: 237 REDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLA 296
R DY AGG++M+S ADP+G++TA V++RN +Y++PLGF AY+WG+TS WF LE++ LTL
Sbjct: 278 RNDYLAGGYRMFSFADPTGKRTAWVSLRNCLYMLPLGFFAYNWGLTSEWFSLEASLLTLG 337
Query: 297 ISAAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNAEGFMKSPSS 356
++ A SF + T + ARRMF+ SLLYLP FMAGLL+HR + Q K +
Sbjct: 338 LTIGALSFVLEPTPKTARRMFYGSLLYLPAFMAGLLLHRLPNEQ-----------KEHNV 386
Query: 357 VETSEM----------DDKNGNQKIKGRQGTR--ARPPVAYASIAPFPFLPAPSY 399
+TSE+ D++ QK + R+ +R +RPPVAYAS+APFPFLP P Y
Sbjct: 387 TQTSEITGILYGAEQQDEERARQKREDRKPSRIHSRPPVAYASVAPFPFLPVPIY 441
>gb|AAH93303.1| LOC553384 protein [Danio rerio]
Length = 452
Score = 205 bits (521), Expect = 2e-51
Identities = 115/270 (42%), Positives = 166/270 (60%), Gaps = 3/270 (1%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y++LSK +L+ LVV T+ GF + + L + LGT + + +A+++NQ FE+ D
Sbjct: 159 YARLSKLKLTALVVTTAAAGFAMAPVPFDPVGFLMAS-LGTGLSSCTANSINQYFEVPFD 217
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
+ M RT RPL G+I+ HAV +A + G+ G LL N L L A N+ LY+ YT
Sbjct: 218 SNMNRTKNRPLVRGQISPLHAVSFALACGIPGVTLLTLGVNPLTGFLGALNIFLYTCCYT 277
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
PLK++ NTWVGAVVGAIPP++GW AA+G + +L+L A L+ WQ PHF AL++ RE
Sbjct: 278 PLKRLSIANTWVGAVVGAIPPVMGWTAATGSLDPGALLLGAFLFSWQFPHFNALSWNLRE 337
Query: 239 DYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAIS 298
DY+ GG++M S+ P K VA+R+S+ LI L L VT+ F L S + L IS
Sbjct: 338 DYSRGGYRMMSVTHPGLCK--RVALRHSVGLIFLSALGPVLDVTTWTFPLVSLPINLYIS 395
Query: 299 AAAFSFYRDRTKERARRMFHASLLYLPVFM 328
AF FYR ++ +R++F SL +LP+ +
Sbjct: 396 YLAFRFYRHGDRQNSRKLFFCSLWHLPMLL 425
>ref|XP_392628.2| PREDICTED: similar to GA18613-PA [Apis mellifera]
Length = 310
Score = 205 bits (521), Expect = 2e-51
Identities = 114/277 (41%), Positives = 173/277 (62%), Gaps = 6/277 (2%)
Query: 62 LSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIM 121
LSK RL+ LVV T+ G+VL G D +GT +++A+A+ +NQ FE+ DA M
Sbjct: 2 LSKIRLTSLVVITTMAGYVLAPG-VFDAHTFIACSMGTGLLSATANAINQFFEVPFDAQM 60
Query: 122 RRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLK 181
RT R L G +T A+ +A+ G +G LL +Q N L A L ASNL+LY+ +YTP+K
Sbjct: 61 SRTKNRVLVRGHLTPGQAIIFATVSGFSGLLLLYSQVNELTAILGASNLILYTLIYTPMK 120
Query: 182 QIHPVNTWVGAVVGAIPPLLGWAAASGDI-SLNSLILPAALYFWQIPHFMALAYMCREDY 240
+I +NTW+G++VGAIPPL+GWA+ DI S + I+ LY WQ PHF AL++ R DY
Sbjct: 121 RISILNTWIGSIVGAIPPLMGWASCVNDIVSPGAWIMSGILYAWQFPHFNALSWNLRPDY 180
Query: 241 AAGGFKMYSLADPS-GRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
+ G++M ++ +P RKT A+R + L L +LA + VT+ WF L ST L +
Sbjct: 181 SRAGYRMMAVTNPKLCRKT---ALRYTAALTGLCYLAPVFNVTNWWFALTSTPLNVYFLY 237
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
A+ F++ + +R++FH SL++LPV + +L++++
Sbjct: 238 LAWKFHKQSDSKSSRKLFHFSLIHLPVLLILILVNKK 274
>ref|NP_848466.1| COX10 homolog, cytochrome c oxidase assembly protein, heme A:
farnesyltransferase [Mus musculus]
gi|56205099|emb|CAI25610.1| novel protein [Mus musculus]
gi|56238020|emb|CAI26029.1| novel protein [Mus musculus]
gi|23959172|gb|AAH38046.1| COX10 homolog, cytochrome c
oxidase assembly protein, heme A: farnesyltransferase
[Mus musculus]
Length = 443
Score = 200 bits (508), Expect = 7e-50
Identities = 113/281 (40%), Positives = 165/281 (58%), Gaps = 3/281 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D S T LGT + + +A+++NQ FE+ D+
Sbjct: 157 ARLSKIKLTALVVSTTSAGFALAPG-PFDWSCFLLTSLGTGLASCAANSINQFFEVPFDS 215
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G ALL N L L N+ LY+ YTP
Sbjct: 216 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVALLTWGVNPLTGALGVFNIFLYTCCYTP 275
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK++ NTWVGAVVGAIPP++GW AA+G + +L+L LY WQ PHF AL++ RED
Sbjct: 276 LKRVSITNTWVGAVVGAIPPVMGWTAATGSLDAGALLLGGILYSWQFPHFNALSWGLRED 335
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P+ VA+R+ + LI L A +T+ F + S + L IS
Sbjct: 336 YSRGGYCMMSVTHPA--LCRRVALRHCLALIALSTAAPVLDITTWVFPVISLPINLYISY 393
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQ 340
F FY D + +R++F SL +LP+ + +L ++ Q
Sbjct: 394 LGFRFYVDADRRSSRKLFFCSLWHLPLLLLLMLTCKQRPGQ 434
>emb|CAG31955.1| hypothetical protein [Gallus gallus]
Length = 448
Score = 200 bits (508), Expect = 7e-50
Identities = 112/269 (41%), Positives = 162/269 (59%), Gaps = 3/269 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF + DL LGT + + +A+++NQ FE+ D+
Sbjct: 150 ARLSKIKLTALVVSTASAGFAMAPV-PFDLPCFLLASLGTGLASCAANSINQFFEVPFDS 208
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+S G+ G ALL N L L A N+ LY+ YTP
Sbjct: 209 NMNRTKNRPLVRGQISPLLAVCFAASCGVPGIALLTMGVNPLTGALGAFNIFLYTCCYTP 268
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
+K++ NTWVGAVVGAIPP++GW AA+G + +L+L LY WQ PHF AL++ RED
Sbjct: 269 MKRMSIANTWVGAVVGAIPPVMGWTAATGSLDAGALLLGGILYSWQFPHFNALSWGLRED 328
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + LI L +A +T+ F + S L L IS
Sbjct: 329 YSRGGYCMMSVTHP--ELCRRVALRHCVALIGLCTVAPILDITTWTFPIISLPLNLYISY 386
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFM 328
F FYRD + +R++F SL +LP+ +
Sbjct: 387 LGFRFYRDADRSSSRKLFFCSLWHLPLLL 415
>ref|XP_415596.1| PREDICTED: similar to COX10 homolog, cytochrome c oxidase assembly
protein, heme A: farnesyltransferase [Gallus gallus]
Length = 1122
Score = 200 bits (508), Expect = 7e-50
Identities = 112/269 (41%), Positives = 162/269 (59%), Gaps = 3/269 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF + DL LGT + + +A+++NQ FE+ D+
Sbjct: 192 ARLSKIKLTALVVSTASAGFAMAPV-PFDLPCFLLASLGTGLASCAANSINQFFEVPFDS 250
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+S G+ G ALL N L L A N+ LY+ YTP
Sbjct: 251 NMNRTKNRPLVRGQISPLLAVCFAASCGVPGIALLTMGVNPLTGALGAFNIFLYTCCYTP 310
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
+K++ NTWVGAVVGAIPP++GW AA+G + +L+L LY WQ PHF AL++ RED
Sbjct: 311 MKRMSIANTWVGAVVGAIPPVMGWTAATGSLDAGALLLGGILYSWQFPHFNALSWGLRED 370
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + LI L +A +T+ F + S L L IS
Sbjct: 371 YSRGGYCMMSVTHP--ELCRRVALRHCVALIGLCTVAPILDITTWTFPIISLPLNLYISY 428
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFM 328
F FYRD + +R++F SL +LP+ +
Sbjct: 429 LGFRFYRDADRSSSRKLFFCSLWHLPLLL 457
>gb|EAL29304.1| GA18613-PA [Drosophila pseudoobscura]
Length = 341
Score = 199 bits (505), Expect = 2e-49
Identities = 114/291 (39%), Positives = 172/291 (58%), Gaps = 8/291 (2%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y +LSK RL+ LVV T+ G+ + A DLS + LGT M++A+A+ +NQ E+ D
Sbjct: 23 YKKLSKFRLTSLVVVTTMGGYAMAPA-AFDLSSFAMCTLGTGMISAAANAINQYHEVPFD 81
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
+ M RT R L +G+IT +V +A+ G +L N L A L A NL LY+ +YT
Sbjct: 82 SQMSRTKNRVLVTGQITPLKSVLFAAVSASTGLCMLYFGVNGLTAALGAGNLFLYTSIYT 141
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
P+K+I VNTW+G+VVGAIPPL+GWA +G + ++IL LY WQ PHF AL++ R
Sbjct: 142 PMKRISIVNTWIGSVVGAIPPLMGWAGCAGSLDSGAMILAGILYAWQFPHFNALSWNLRP 201
Query: 239 DYAAGGFKMYSLADPS-GRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAI 297
DY+ G++M ++ +PS R+T A R S+ ++ L +A VT+ WF LE+ L
Sbjct: 202 DYSRAGYRMMAVTNPSLCRRT---AFRYSLAIVGLSLMAPVLDVTNYWFALETIPLNAYF 258
Query: 298 SAAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNAE 348
+ A+ F + + +R +F SLL+LP M L +++ + + + NAE
Sbjct: 259 AYLAYKFNKKSDSKSSRSLFRFSLLHLPALMLLFLANKK---EWYFKKNAE 306
>gb|EAK86095.1| hypothetical protein UM05692.1 [Ustilago maydis 521]
gi|49079292|ref|XP_403307.1| hypothetical protein
UM05692.1 [Ustilago maydis 521]
Length = 1527
Score = 197 bits (500), Expect = 6e-49
Identities = 117/281 (41%), Positives = 168/281 (59%), Gaps = 13/281 (4%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGG-----AVDLSMLSYTCLGTMMVAASASTLNQVF 113
Y QLSK+RL+ LVV T G+ L A ++ L G + +A+A+ LNQ+
Sbjct: 1216 YKQLSKSRLTFLVVLTGMAGYALCPASLTVAVASPVTTLLALTAGMTLCSAAANALNQLV 1275
Query: 114 EIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLY 173
E DA M+RT RPLPS +T HA +AS +G LL T N L A L A+N+VLY
Sbjct: 1276 ESPYDAQMQRTRARPLPSRSVTPLHAFTFASVSAASGVGLLLTTVNPLTALLGAANVVLY 1335
Query: 174 SFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLI----LPAALYFWQIPHF 229
SF YTP+K+ NTWVGAVVGA+PPL+GWAA +G + ++ + L A L+ WQ PHF
Sbjct: 1336 SFTYTPMKRSTIANTWVGAVVGALPPLMGWAACTGTLHTSTDVGGWSLAALLFAWQFPHF 1395
Query: 230 MALAYMCREDYAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWG--VTSGWFC 287
+LA+ R +YA GG++M ++ DP+ + V++R ++ L+P+ L V+ +
Sbjct: 1396 NSLAHTLRAEYARGGYRMMAVTDPALNR--RVSLRYAVALLPICTLMIPLSGIVSPVAYA 1453
Query: 288 LESTALTLAISAAAFSFYRDRTKERARRMFHASLLYLPVFM 328
+ ST L L ++ AA+ FYR+RT + AR F SL++LP M
Sbjct: 1454 VLSTPLNLLMAHAAWKFYRERTVKSARWCFWVSLIHLPAVM 1494
>gb|AAF52915.1| CG5037-PA [Drosophila melanogaster] gi|19921064|ref|NP_609382.1|
CG5037-PA [Drosophila melanogaster]
gi|16198131|gb|AAL13868.1| LD33876p [Drosophila
melanogaster]
Length = 391
Score = 196 bits (498), Expect = 1e-48
Identities = 108/279 (38%), Positives = 167/279 (59%), Gaps = 5/279 (1%)
Query: 59 YSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKND 118
Y +LSK RL+ LVV T+ G+ + A D + + LGT +V+A+A+ +NQ E+ D
Sbjct: 76 YKKLSKFRLTSLVVITTMGGYAMAPA-AFDPTTFAMCTLGTGLVSAAANAINQYHEVPFD 134
Query: 119 AIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYT 178
+ M RT R L +G++T HAV +A+ AG ++L N L A L A NL LY+ +YT
Sbjct: 135 SQMSRTKNRVLVTGQMTPLHAVTFAAVSATAGLSMLYFGVNGLTAALGAGNLFLYTTIYT 194
Query: 179 PLKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRE 238
P+K+I VNTWVG++VGAIPPL+GWA + + ++IL LY WQ PHF AL++ R
Sbjct: 195 PMKRISIVNTWVGSIVGAIPPLMGWAGCAATLDAGAMILAGVLYAWQFPHFNALSWNLRP 254
Query: 239 DYAAGGFKMYSLADPS-GRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAI 297
DY+ G++M ++ +P R+T A+R S+ ++ L +A VT+ WF LE+ L
Sbjct: 255 DYSRAGYRMMAVTNPGLCRRT---ALRYSVAIVGLSAMAPVLDVTNYWFALETLPLNAYF 311
Query: 298 SAAAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
+ A+ F+ +R++F SL++LP M L +++
Sbjct: 312 AYLAYKFHEKSDSGSSRKLFRFSLIHLPALMLLFLTNKK 350
>gb|AAP36367.1| Homo sapiens COX10 homolog, cytochrome c oxidase assembly protein,
heme A: farnesyltransferase (yeast) [synthetic
construct] gi|60653197|gb|AAX29293.1| cytochrome c
oxidase assembly protein [synthetic construct]
gi|60653195|gb|AAX29292.1| cytochrome c oxidase assembly
protein [synthetic construct]
Length = 444
Score = 194 bits (494), Expect = 3e-48
Identities = 109/277 (39%), Positives = 160/277 (57%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
+QLSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 AQLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RED
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>gb|AAP35631.1| COX10 homolog, cytochrome c oxidase assembly protein, heme A:
farnesyltransferase (yeast) [Homo sapiens]
gi|60656255|gb|AAX32691.1| COX10-like [synthetic
construct] gi|60656253|gb|AAX32690.1| COX10-like
[synthetic construct] gi|13623563|gb|AAH06394.1| Heme
A:farnesyltransferase [Homo sapiens]
gi|12652629|gb|AAH00060.1| Heme A:farnesyltransferase
[Homo sapiens] gi|20141419|sp|Q12887|COX10_HUMAN
Protoheme IX farnesyltransferase, mitochondrial
precursor (Heme O synthase) gi|2138300|gb|AAC51330.1|
heme A: farnesyltransferase [Homo sapiens]
Length = 443
Score = 194 bits (494), Expect = 3e-48
Identities = 109/277 (39%), Positives = 160/277 (57%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
+QLSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 AQLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RED
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>ref|NP_001294.2| heme A:farnesyltransferase [Homo sapiens]
Length = 443
Score = 193 bits (490), Expect = 9e-48
Identities = 108/277 (38%), Positives = 160/277 (56%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 ARLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RED
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>dbj|BAE00665.1| unnamed protein product [Macaca fascicularis]
Length = 298
Score = 192 bits (487), Expect = 2e-47
Identities = 109/277 (39%), Positives = 160/277 (57%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D +GT +V+ +A+++NQ FE+ D+
Sbjct: 13 ARLSKIKLTGLVVSTTAAGFALAPG-PFDWPCFLLASVGTGLVSCAANSINQFFEVPFDS 71
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 72 NMNRTKNRPLVRGQISPLLAVAFATCCAVPGVAILTWGVNPLTGALGLFNIFLYTCCYTP 131
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I VNTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RED
Sbjct: 132 LKRISIVNTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 191
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + IS
Sbjct: 192 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPFVALPINAYISY 249
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 250 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 286
>gb|AAA21148.1| heme A:farnesyltransferase
Length = 443
Score = 190 bits (483), Expect = 6e-47
Identities = 107/277 (38%), Positives = 158/277 (56%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
+QLSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 AQLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW A +G + + +L LY WQ PHF AL++ RED
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTATTGSLDAGAFLLGGILYSWQFPHFNALSWGLRED 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ G+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRDGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISH 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>emb|CAH93456.1| hypothetical protein [Pongo pygmaeus]
Length = 443
Score = 190 bits (483), Expect = 6e-47
Identities = 107/277 (38%), Positives = 159/277 (56%), Gaps = 3/277 (1%)
Query: 60 SQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDA 119
++LSK +L+ LVV+T+ GF L G D T +GT + + +A+++NQ FE+ D+
Sbjct: 158 ARLSKIKLTALVVSTTAAGFALAPG-PFDWPCFLLTSVGTGLASCAANSINQFFEVPFDS 216
Query: 120 IMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTP 179
M RT RPL G+I+ AV +A+ + G A+L N L L N+ LY+ YTP
Sbjct: 217 NMNRTKNRPLVRGQISPLLAVSFATCCAVPGVAILTLGVNPLTGALGLFNIFLYTCCYTP 276
Query: 180 LKQIHPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCRED 239
LK+I NTWVGAVVGAIPP++GW AA+G + + +L LY WQ PHF AL++ RE
Sbjct: 277 LKRISIANTWVGAVVGAIPPVMGWTAATGSLDAGAFLLGGILYSWQFPHFNALSWGLREG 336
Query: 240 YAAGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISA 299
Y+ GG+ M S+ P VA+R+ + L+ L A +T+ F + + + IS
Sbjct: 337 YSRGGYCMMSVTHPG--LCRRVALRHCLALLVLSAAAPVLDITTWTFPIMALPINAYISY 394
Query: 300 AAFSFYRDRTKERARRMFHASLLYLPVFMAGLLIHRR 336
F FY D + +RR+F SL +LP+ + +L +R
Sbjct: 395 LGFRFYVDADRRSSRRLFFCSLWHLPLLLLLMLTCKR 431
>ref|XP_319631.2| ENSANGP00000013941 [Anopheles gambiae str. PEST]
gi|55235180|gb|EAA14890.2| ENSANGP00000013941 [Anopheles
gambiae str. PEST]
Length = 271
Score = 187 bits (476), Expect = 4e-46
Identities = 104/268 (38%), Positives = 158/268 (58%), Gaps = 4/268 (1%)
Query: 63 SKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASASTLNQVFEIKNDAIMR 122
S A + LVV T+ G+ + G + +L GT +V+ +A+++NQV E DA M
Sbjct: 4 SLAPFAALVVMTAMAGYAMAPG-SFELGTFLLCSAGTTLVSGAANSINQVIETSFDAQMP 62
Query: 123 RTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQ 182
RT R L G ++ HAVG+A G ++L N + A L A+NL+LY+ +YTP+K+
Sbjct: 63 RTRNRVLVKGHLSRLHAVGFALGASTIGCSMLYYGVNEMTAALGAANLILYTSIYTPMKR 122
Query: 183 IHPVNTWVGAVVGAIPPLLGWAAAS-GDISLNSLILPAALYFWQIPHFMALAYMCREDYA 241
+NTWVG+VVGAIPPL+GWAA + GD+ + IL LY WQ PHF AL++ R +Y
Sbjct: 123 YSILNTWVGSVVGAIPPLMGWAACTGGDLGAGAWILAGLLYCWQFPHFNALSWNIRPEYL 182
Query: 242 AGGFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAA 301
G+KM + + P+ V++R++ ++ L LA VT+ WF LES L + A
Sbjct: 183 KAGYKMMANSHPA--LCTRVSLRHTGFIAGLSLLAPALDVTNVWFALESLPLNAYFAYLA 240
Query: 302 FSFYRDRTKERARRMFHASLLYLPVFMA 329
+ F++ + +R++F SLL+LP+ MA
Sbjct: 241 YDFHQKADSKSSRKLFRFSLLHLPLLMA 268
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 642,863,064
Number of Sequences: 2540612
Number of extensions: 25393482
Number of successful extensions: 91206
Number of sequences better than 10.0: 493
Number of HSP's better than 10.0 without gapping: 348
Number of HSP's successfully gapped in prelim test: 145
Number of HSP's that attempted gapping in prelim test: 90156
Number of HSP's gapped (non-prelim): 563
length of query: 402
length of database: 863,360,394
effective HSP length: 130
effective length of query: 272
effective length of database: 533,080,834
effective search space: 144997986848
effective search space used: 144997986848
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC146329.8


