

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145512.11 - phase: 0
(127 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
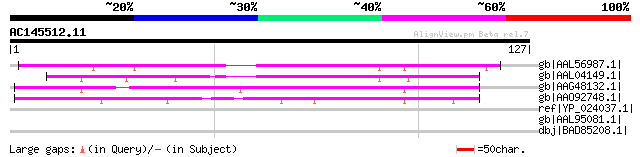
gb|AAL56987.1| functional candidate resistance protein KR1 [Glyc... 79 2e-14
gb|AAL04149.1| NR1 [Glycine max] 65 3e-10
gb|AAG48132.1| putative resistance protein [Glycine max] 61 7e-09
gb|AAO92748.1| candidate disease-resistance protein SR1 [Glycine... 44 7e-04
ref|YP_024037.1| beta-galactosidase [Picrophilus torridus DSM 97... 33 1.2
gb|AAL95081.1| Hemin-binding periplasmic protein hmuT precursor ... 32 4.5
dbj|BAD85208.1| hypothetical protein, conserved [Thermococcus ko... 31 7.7
>gb|AAL56987.1| functional candidate resistance protein KR1 [Glycine max]
Length = 1124
Score = 79.0 bits (193), Expect = 2e-14
Identities = 54/140 (38%), Positives = 77/140 (54%), Gaps = 29/140 (20%)
Query: 3 VSFWFCNKFPSIALGVV--------SAYTWGYLEH-PARVIINDNTFFYTHGRKIDRCSR 53
+SFWF NKFP+IA+ + S+ W + + +VIIN N + C+
Sbjct: 936 ISFWFRNKFPAIAICHIIKRVAEFSSSRGWTFRPNIRTKVIINGNANLFNSVVLGSDCTC 995
Query: 54 PDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF-GFPFMF------SGIHVLKEKSN 106
LF ++ E N+D+ALLEN+WNHAEV GF F F +G+HVLK++SN
Sbjct: 996 -------LFDLRGERVTDNLDEALLENEWNHAEVTCPGFTFTFAPTFIKTGLHVLKQESN 1048
Query: 107 MKDIRFTNP------ENDAN 120
M+DIRF++P +ND N
Sbjct: 1049 MEDIRFSDPCRKTKLDNDFN 1068
>gb|AAL04149.1| NR1 [Glycine max]
Length = 175
Score = 65.5 bits (158), Expect = 3e-10
Identities = 50/123 (40%), Positives = 68/123 (54%), Gaps = 25/123 (20%)
Query: 10 KFPSIAL--------GVVSAYTWGYL-EHPARVIINDNT-FFYTHGRKIDRCSRPDTYHL 59
KFP+IA+ S+ W Y + +VIIN N FY+ D CS
Sbjct: 1 KFPAIAICHSIKRVPEFSSSRGWTYRPDIRTKVIINGNANLFYSMDLGSD-CSV------ 53
Query: 60 HLFHMQVEYFNGNMDKALLENKWNHAEVDF-GFPFMFS------GIHVLKEKSNMKDIRF 112
LF + + N+D+ALLEN+WNHAEV GF F F+ G+HVLK++SNM+DIRF
Sbjct: 54 -LFDPRDDKVTDNLDEALLENEWNHAEVTCPGFTFTFAPTFIKTGLHVLKQESNMEDIRF 112
Query: 113 TNP 115
++P
Sbjct: 113 SDP 115
>gb|AAG48132.1| putative resistance protein [Glycine max]
Length = 1093
Score = 60.8 bits (146), Expect = 7e-09
Identities = 43/127 (33%), Positives = 64/127 (49%), Gaps = 16/127 (12%)
Query: 2 SVSFWFCNKFPSIAL---GVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPD-TY 57
S+SFWF NKFP I+L G++ + +G V IN N R+ P T
Sbjct: 964 SISFWFRNKFPVISLCLAGLMHKHPFGL---KPIVSINGNKMKTEFQRRWFYFEFPVLTD 1020
Query: 58 HLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF---------SGIHVLKEKSNMK 108
H+ +F + F N+D+ + EN WNH V F + +G+HV+K KS+++
Sbjct: 1021 HILIFGERQIKFEDNVDEVVSENDWNHVVVSVDVDFKWNPTEPLVVRTGLHVIKPKSSVE 1080
Query: 109 DIRFTNP 115
DIRF +P
Sbjct: 1081 DIRFIDP 1087
>gb|AAO92748.1| candidate disease-resistance protein SR1 [Glycine max]
Length = 1137
Score = 44.3 bits (103), Expect = 7e-04
Identities = 41/139 (29%), Positives = 67/139 (47%), Gaps = 29/139 (20%)
Query: 2 SVSFWFCNKFPSIALGVVSA-------------YTWGYLEHPARVIIND--NTFFYTHGR 46
S SFWF NKFP+ L ++ A ++G+ +V IN F+ H +
Sbjct: 916 SSSFWFRNKFPAKLLCLLIAPVSVPLYSLFPPKVSFGHHVPYPKVFINGKCQAFWGCHWK 975
Query: 47 KIDRCSRPDTYHLHLFHMQ-VEYFNGNM-DKALLENKWNHAEVDFGFPFMF-------SG 97
+ R D H ++F +Q + + N N+ ++ E +WNH EV + SG
Sbjct: 976 Q--RMMELD--HTYIFDLQKLPFENDNLFEEGAWEEEWNHVEVRYESVLELESSLIKGSG 1031
Query: 98 IHVLKEKSNM-KDIRFTNP 115
IH+ +E+ +M +DIRF +P
Sbjct: 1032 IHIFREEGSMEEDIRFDDP 1050
>ref|YP_024037.1| beta-galactosidase [Picrophilus torridus DSM 9790]
gi|48430979|gb|AAT43844.1| beta-galactosidase
[Picrophilus torridus DSM 9790]
Length = 453
Score = 33.5 bits (75), Expect = 1.2
Identities = 20/80 (25%), Positives = 41/80 (51%), Gaps = 4/80 (5%)
Query: 17 GVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSR-PDTYHLHLFHMQVEYFNGNMDK 75
G +S + GYL++ + N + + +TH + + + D Y ++EY N N+ +
Sbjct: 8 GFMSGFHLGYLQNNKNLPENSDWYQWTHDKNVRMMNYIRDGYPED---NKIEYLNDNVIR 64
Query: 76 ALLENKWNHAEVDFGFPFMF 95
L +N +N ++D +P +F
Sbjct: 65 DLHDNNFNLIKIDMDWPSLF 84
>gb|AAL95081.1| Hemin-binding periplasmic protein hmuT precursor [Fusobacterium
nucleatum subsp. nucleatum ATCC 25586]
gi|19704220|ref|NP_603782.1| Hemin-binding periplasmic
protein hmuT precursor [Fusobacterium nucleatum subsp.
nucleatum ATCC 25586]
Length = 286
Score = 31.6 bits (70), Expect = 4.5
Identities = 17/42 (40%), Positives = 25/42 (59%)
Query: 70 NGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIR 111
+GN +K L+EN +D+G S I LKEKS +KD++
Sbjct: 207 DGNWEKVLVENPDIIVIIDYGDQSAESKIKFLKEKSPIKDLK 248
>dbj|BAD85208.1| hypothetical protein, conserved [Thermococcus kodakarensis KOD1]
gi|57640954|ref|YP_183432.1| hypothetical protein TK1019
[Thermococcus kodakaraensis KOD1]
Length = 140
Score = 30.8 bits (68), Expect = 7.7
Identities = 20/85 (23%), Positives = 34/85 (39%), Gaps = 11/85 (12%)
Query: 24 WGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLH-----------LFHMQVEYFNGN 72
W +L+ R+ +N+ +T GRK C+R D L + + YF+G
Sbjct: 29 WVFLDRSKRIDVNEKITIFTDGRKALVCTRLDCERLQKSEILERVRGIMDSYECVYFDGR 88
Query: 73 MDKALLENKWNHAEVDFGFPFMFSG 97
+ K K+ D + F +G
Sbjct: 89 VAKVTRPEKFIETVPDGEYCFYLNG 113
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.326 0.140 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 239,533,221
Number of Sequences: 2540612
Number of extensions: 9850642
Number of successful extensions: 21415
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 21403
Number of HSP's gapped (non-prelim): 7
length of query: 127
length of database: 863,360,394
effective HSP length: 103
effective length of query: 24
effective length of database: 601,677,358
effective search space: 14440256592
effective search space used: 14440256592
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC145512.11


