

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
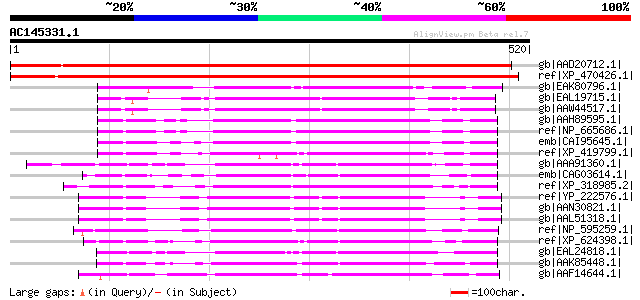
Query= AC145331.1 - phase: 0
(520 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAD20712.1| hypothetical protein [Arabidopsis thaliana] gi|25... 709 0.0
ref|XP_470426.1| putative AFG1-like ATPase [Oryza sativa (japoni... 673 0.0
gb|EAK80796.1| hypothetical protein UM00002.1 [Ustilago maydis 5... 210 8e-53
gb|EAL19715.1| hypothetical protein CNBG3430 [Cryptococcus neofo... 198 4e-49
gb|AAW44517.1| hypothetical protein CNG01350 [Cryptococcus neofo... 197 7e-49
gb|AAH89595.1| Lactation elevated 1 [Mus musculus] 195 3e-48
ref|NP_665686.1| lactation elevated 1 [Mus musculus] gi|21668096... 194 7e-48
emb|CAI95645.1| lactation elevated 1 [Homo sapiens] gi|66347711|... 193 1e-47
ref|XP_419799.1| PREDICTED: similar to lactation elevated 1; CG8... 193 1e-47
gb|AAA91360.1| putative ATPase gi|1171633|sp|P46441|N2B_HAEIR PU... 191 6e-47
emb|CAG03614.1| unnamed protein product [Tetraodon nigroviridis] 186 2e-45
ref|XP_318985.2| ENSANGP00000014807 [Anopheles gambiae str. PEST... 185 3e-45
ref|YP_222576.1| hypothetical protein BruAb1_1905 [Brucella abor... 184 5e-45
gb|AAN30821.1| conserved hypothetical protein [Brucella suis 133... 184 5e-45
gb|AAL51318.1| PUTATIVE ATPASE N2B [Brucella melitensis 16M] gi|... 184 5e-45
ref|NP_595259.1| ATPase [Schizosaccharomyces pombe 972h-] gi|295... 184 6e-45
ref|XP_624398.1| PREDICTED: similar to ENSANGP00000014807 [Apis ... 184 6e-45
gb|EAL24818.1| GA21133-PA [Drosophila pseudoobscura] 174 8e-42
gb|AAK85448.1| Hypothetical protein C30F12.2 [Caenorhabditis ele... 169 1e-40
gb|AAF14644.1| 7138.4 [Leishmania major] 148 5e-34
>gb|AAD20712.1| hypothetical protein [Arabidopsis thaliana] gi|25412329|pir||E84649
hypothetical protein At2g25530 [imported] - Arabidopsis
thaliana gi|15224744|ref|NP_180123.1| AFG1-like ATPase
family protein [Arabidopsis thaliana]
Length = 655
Score = 709 bits (1829), Expect = 0.0
Identities = 335/504 (66%), Positives = 408/504 (80%), Gaps = 3/504 (0%)
Query: 1 MEEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWK--RLNNKITERWTNSRKRPENV 58
ME+YHV L+ WEKKRE ERR+++++E EK+++D W + K+ RW R R NV
Sbjct: 152 MEDYHVRLAEWEKKREEERRKLMVEEAEKKEDDGMWASVNKQGQKLLGRWVLGR-RQMNV 210
Query: 59 DPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVK 118
+PGVGKWVSYL RE+KLDS+VG RP PPAPKGLYIYGNVG GKTMLMDMFY AT+GI++
Sbjct: 211 EPGVGKWVSYLNRERKLDSIVGSRPAVPPAPKGLYIYGNVGCGKTMLMDMFYGATDGIIR 270
Query: 119 HRRRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK 178
HR+R+HFHEAML+INE MHK WK+ EKP+Q ISSWIMNLP D K KEWLA EE YK+
Sbjct: 271 HRQRFHFHEAMLKINEQMHKYWKENGAEKPMQYSISSWIMNLPVDEKVKEWLAGEEFYKQ 330
Query: 179 EVQMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIV 238
++QMKHILP VADKF +D++ +KGA+ILCFDEIQTVDVFAIVALSGI+SRLL++GT++V
Sbjct: 331 QLQMKHILPAVADKFLVDQQSSKKGASILCFDEIQTVDVFAIVALSGIMSRLLTTGTVLV 390
Query: 239 ATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPI 298
ATSNRAP++LN+ M E F +S LE+HCE + +GSE+DYRR A+ S V+YLWP+
Sbjct: 391 ATSNRAPRELNQDGMQKEIFDKFISKLEKHCEIISIGSEVDYRRVAAKNSAENVHYLWPL 450
Query: 299 ERETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGA 358
+ +FEK W+ T ++GG++ S T+ VMFGRT+EVPESC GVARFTF+YLCGRP+GA
Sbjct: 451 NDAVLEEFEKMWRQVTDQYGGEITSATLPVMFGRTVEVPESCSGVARFTFEYLCGRPVGA 510
Query: 359 ADYIAVAENYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQG 418
ADYIAVA+NYHT+FISDIP MSM IRDKARRFITL+DELYNHH CL A ++IDELFQG
Sbjct: 511 ADYIAVAKNYHTIFISDIPAMSMEIRDKARRFITLVDELYNHHCCLVSSAETAIDELFQG 570
Query: 419 TEEGTLFDLESFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVS 478
T EGTLFDLESFQFETE E S+LRRDVLAEG++ + G+P I S+LSG+EE+F F RA S
Sbjct: 571 TAEGTLFDLESFQFETETEDSRLRRDVLAEGSISAAGSPSSIVSMLSGEEEMFAFARAAS 630
Query: 479 RLIEMQTQLYLDGVSNVHPYFQTQ 502
RLIEMQT LYL+GV +HPYF Q
Sbjct: 631 RLIEMQTPLYLEGVRFLHPYFHQQ 654
>ref|XP_470426.1| putative AFG1-like ATPase [Oryza sativa (japonica cultivar-group)]
gi|27573346|gb|AAO20064.1| putative AFG1-like ATPase
[Oryza sativa (japonica cultivar-group)]
Length = 740
Score = 673 bits (1737), Expect = 0.0
Identities = 323/509 (63%), Positives = 397/509 (77%), Gaps = 2/509 (0%)
Query: 1 MEEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDP 60
ME+YH LS WE RE +RRR+L+ E E +Q D W + + SRKR N++P
Sbjct: 224 MEDYHARLSMWENTREKQRRRLLVQEAEDKQRDGVWIDEKRGFLDK--LVSRKRRGNIEP 281
Query: 61 GVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHR 120
GVGKWVSYL REKKLD+LVG++P AP APKG+Y+YGNVGSGKTMLMDMFY ATEG++KHR
Sbjct: 282 GVGKWVSYLNREKKLDTLVGQKPVAPIAPKGIYLYGNVGSGKTMLMDMFYGATEGLIKHR 341
Query: 121 RRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEV 180
RR+HFHEAML I++HMH WK++ E+K ++S SWI +LPFD K KEWL EE+YK+
Sbjct: 342 RRFHFHEAMLEIHDHMHDVWKRRDEDKSIESSAFSWISSLPFDGKIKEWLIGEEKYKQNT 401
Query: 181 QMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVAT 240
Q HIL VADKF +DR+ + GA+ILCFDEIQT+DVFA+VALSGILSRLLS+GT++V+T
Sbjct: 402 QQNHILLAVADKFLVDRQANKSGASILCFDEIQTIDVFAVVALSGILSRLLSTGTVLVST 461
Query: 241 SNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIER 300
SN+AP+DLN+ M E F +LLS L+E+C K+LVG+E DYRR I +++Y WP+
Sbjct: 462 SNKAPEDLNQDGMQREIFLDLLSKLDENCNKILVGTETDYRRLIPTDGLTQIHYFWPLTS 521
Query: 301 ETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAAD 360
+ + +E W D T + GG +IS TI VMFGR LE+P+SC GVARF F+YLCGRP+GAAD
Sbjct: 522 DIRSMYEAMWHDITRQTGGNIISVTIPVMFGRYLEIPKSCNGVARFDFEYLCGRPVGAAD 581
Query: 361 YIAVAENYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTE 420
YIA+A NYHT+FISDIP MSM+IRDKARRFITLIDELYNHH L CLA+SSID+LFQGT+
Sbjct: 582 YIAIARNYHTIFISDIPAMSMKIRDKARRFITLIDELYNHHCRLVCLAASSIDDLFQGTD 641
Query: 421 EGTLFDLESFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRL 480
EG LFDLESFQFE EAEG+KLRRDVLAEGNVG+ +P G+ +ILSGQEE+F F+RA+SRL
Sbjct: 642 EGPLFDLESFQFEGEAEGAKLRRDVLAEGNVGAAPSPTGLVAILSGQEEMFAFRRAISRL 701
Query: 481 IEMQTQLYLDGVSNVHPYFQTQHKKFQKN 509
IEMQT LYL+ V VH Q Q K+
Sbjct: 702 IEMQTSLYLERVERVHSSLQQQSSVLTKS 730
>gb|EAK80796.1| hypothetical protein UM00002.1 [Ustilago maydis 521]
gi|49066654|ref|XP_397617.1| hypothetical protein
UM00002.1 [Ustilago maydis 521]
Length = 550
Score = 210 bits (535), Expect = 8e-53
Identities = 142/409 (34%), Positives = 205/409 (49%), Gaps = 41/409 (10%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMH----KTWKKQM 144
PKGLY+YG+VG+GK+MLMD+FY + +RR HFH+ M+ ++ H KT K
Sbjct: 169 PKGLYLYGDVGTGKSMLMDLFYDTLPSNITSKRRIHFHQFMIEAHKRAHFYKSKTHKPSG 228
Query: 145 EEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGA 204
+ SG SS + A E A E +E+ H
Sbjct: 229 IVMMMSSGSSSSASSAGGAASAGEESDAIEAVAREMARNH-------------------- 268
Query: 205 NILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSN 264
++LCFDE Q D+ + L G+L R+L+ G ++V TSNR P +L + + + F +
Sbjct: 269 SVLCFDEFQVTDIADAMILRGLLERMLAYGVVMVMTSNRHPDELYKNGIQRQSFLPCIDL 328
Query: 265 LEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISN 324
L+ + S DYR+ R+ ++V Y P++ +F+K + T V+
Sbjct: 329 LKSRLGVTDLNSGTDYRK--VPRALSKV-YFSPLDDANTREFDKLFDAMTSDPHDPVVEK 385
Query: 325 TISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIR 384
++GRTL+VP S + VARFTFD LCGRP AADYI + N+ T+FI DIP M + R
Sbjct: 386 RPLKIWGRTLQVPRSTQRVARFTFDELCGRPRSAADYIEICNNFSTIFIDDIPKMGLNQR 445
Query: 385 DKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEGSKLRRD 444
D ARRFIT ID Y + L LASS + L +F ++ + A+ + D
Sbjct: 446 DLARRFITFIDAAYESKTKL--LASSEVPIL-------QIFSGDAGDAKPTADQMRALMD 496
Query: 445 VLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLYLDGVS 493
L GG+P I +G EELF F R +SRL EM ++ Y + S
Sbjct: 497 DLGLTMDDLGGSP-----IFTGDEELFAFARVISRLTEMGSRQYAETAS 540
>gb|EAL19715.1| hypothetical protein CNBG3430 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 521
Score = 198 bits (503), Expect = 4e-49
Identities = 143/408 (35%), Positives = 201/408 (49%), Gaps = 68/408 (16%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRR----------RYHFHEAMLRINEHMHK 138
PKGLY+YG+VG+GKTMLMD+F+S I K R R HFH ML + + HK
Sbjct: 160 PKGLYLYGSVGTGKTMLMDLFHST---IPKQFRPTSQGGYGSIRIHFHAFMLDVLQRQHK 216
Query: 139 TWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDRE 198
E K + K +LP+VA L E
Sbjct: 217 L--------------------------------VVEYEKAGLGKKDVLPEVARS--LANE 242
Query: 199 GEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFF 258
G +LCFDE Q D+ + L G+L RL+S G + + TSNR P +L + + F
Sbjct: 243 GR-----VLCFDEFQVTDIVTAMILRGLLERLMSFGVVCIMTSNRHPDELYINGIQRQSF 297
Query: 259 QNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFG 318
+ ++E E V + S DYR+ S+ N L P + INK + +T
Sbjct: 298 IPAIELIKERFEVVDLDSGTDYRKIPRALSKVYYNPLSPTVKSEINKLFDSFA-STDPVS 356
Query: 319 GKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPM 378
+V+ N ++GR L VPES VA+FTF LC +PL AADY+ V + TVF+ DIP
Sbjct: 357 SEVVHNRKVHLWGRELNVPESSGSVAKFTFADLCNKPLSAADYLEVTSKFGTVFVEDIPR 416
Query: 379 MSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEG 438
M + RD+ARRFIT ID Y + + L C + E +F + S + + AE
Sbjct: 417 MGLSERDQARRFITFIDACYENKTKLFC------------SSEVPIFQVFSDKHGSAAED 464
Query: 439 SKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQ 486
+ + ++V+ E +G + VG +S+ SG EELF F R VSRL +M T+
Sbjct: 465 AHM-QEVMDE--LGLDPSAVGSSSLFSGDEELFAFARCVSRLSQMGTK 509
>gb|AAW44517.1| hypothetical protein CNG01350 [Cryptococcus neoformans var.
neoformans JEC21] gi|58269336|ref|XP_571824.1|
hypothetical protein CNG01350 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 521
Score = 197 bits (501), Expect = 7e-49
Identities = 143/408 (35%), Positives = 200/408 (48%), Gaps = 68/408 (16%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRR----------RYHFHEAMLRINEHMHK 138
PKGLY+YG+VG+GKTMLMD+F+S I K R R HFH ML + + HK
Sbjct: 160 PKGLYLYGSVGTGKTMLMDLFHST---IPKQFRPTSQGGYGSIRIHFHAFMLDVLQRQHK 216
Query: 139 TWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDRE 198
E K + K +LP+VA L E
Sbjct: 217 L--------------------------------VVEYEKAGLGKKDVLPEVARS--LANE 242
Query: 199 GEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFF 258
G +LCFDE Q D+ + L G+L RL+S G + + TSNR P +L + + F
Sbjct: 243 GR-----VLCFDEFQVTDIVTAMILRGLLERLMSFGVVCIMTSNRHPDELYINGIQRQSF 297
Query: 259 QNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFG 318
+ ++E E V + S DYR S+ N L P + INK + +T
Sbjct: 298 IPAIELIKERFEVVDLDSGTDYREIPRALSKVYYNPLSPTVKSEINKLFDSFA-STDPVS 356
Query: 319 GKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPM 378
+V+ N ++GR L VPES VA+FTF LC +PL AADY+ V + TVF+ DIP
Sbjct: 357 SEVVHNRKVHLWGRELNVPESSGSVAKFTFADLCNKPLSAADYLEVTSKFGTVFVEDIPR 416
Query: 379 MSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEG 438
M + RD+ARRFIT ID Y + + L C + E +F + S + + AE
Sbjct: 417 MGLSERDQARRFITFIDACYENKTKLFC------------SSEVPIFQVFSDKHGSAAED 464
Query: 439 SKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQ 486
+ + ++V+ E +G + VG +S+ SG EELF F R VSRL +M T+
Sbjct: 465 AHM-QEVMDE--LGLDPSAVGSSSLFSGDEELFAFARCVSRLSQMGTK 509
>gb|AAH89595.1| Lactation elevated 1 [Mus musculus]
Length = 480
Score = 195 bits (495), Expect = 3e-48
Identities = 131/401 (32%), Positives = 202/401 (49%), Gaps = 59/401 (14%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKP 148
PKGLY+YG+VG+GKTM+MDMFY+ E K ++R HFH ML ++ +H + + K
Sbjct: 129 PKGLYVYGDVGTGKTMVMDMFYAYVE--TKRKKRVHFHGFMLDVHRRIHHLKQSLPKRK- 185
Query: 149 LQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILC 208
+ M +D A AEE ++ ++LC
Sbjct: 186 ------AGFMAKSYDPIAP---IAEEISQE-------------------------TSLLC 211
Query: 209 FDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEH 268
FDE Q D+ + L + L +G ++VATSNR P+DL + + F ++ L+E+
Sbjct: 212 FDEFQVTDIADAMILKQLFENLFKNGVVVVATSNRPPEDLYKNGLQRANFVPFIAVLKEY 271
Query: 269 CEKVLVGSEIDYRRFIAQRSENRVNYLWPIERET-INKFEKKWQDATGRFGGKVISNTIS 327
C+ + + S +DYR+ R L+ + E + K D + + S I
Sbjct: 272 CDTLQLDSGVDYRK----RELAPAGKLYYLTSEADVEAVVDKLFDELAQKQNDLTSPRIL 327
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKA 387
M GR L + ++C VA TF+ LC RPLGA+DY+ +++N+ TV I +IP S+ R +A
Sbjct: 328 KMQGRELRLNKACGSVADCTFEELCERPLGASDYLELSKNFDTVIIRNIPQFSLAKRTQA 387
Query: 388 RRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEGSKLRRDVLA 447
RRFITLID Y+ + C AS+ I LF + ++E++ S++ D L
Sbjct: 388 RRFITLIDNFYDFKVRIICSASAPISSLFLHQHQ-----------DSESDQSRILMDDLG 436
Query: 448 EGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+G S+ +G+EE+F FQR +SRL EMQT+ Y
Sbjct: 437 LSQDSAG------LSMFTGEEEIFAFQRTISRLTEMQTEQY 471
>ref|NP_665686.1| lactation elevated 1 [Mus musculus] gi|21668096|gb|AAM74227.1|
lactation elevated 1 [Mus musculus]
Length = 480
Score = 194 bits (492), Expect = 7e-48
Identities = 130/401 (32%), Positives = 202/401 (49%), Gaps = 59/401 (14%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKP 148
PKGLY+YG+VG+GKTM+MDMFY+ E K ++R HFH ML ++ +H + + K
Sbjct: 129 PKGLYVYGDVGTGKTMVMDMFYAYVE--TKRKKRVHFHGFMLDVHRRIHHLKQSLPKRK- 185
Query: 149 LQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILC 208
+ M +D A AEE ++ ++LC
Sbjct: 186 ------AGFMAKSYDPIAP---IAEEISQE-------------------------TSLLC 211
Query: 209 FDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEH 268
FDE Q D+ + L + L +G ++VATSNR P+DL + + F ++ L+E+
Sbjct: 212 FDEFQVTDIADAMILKQLFENLFKNGVVVVATSNRPPEDLYKNGLQRANFVPFIAVLKEY 271
Query: 269 CEKVLVGSEIDYRRFIAQRSENRVNYLWPIERET-INKFEKKWQDATGRFGGKVISNTIS 327
C+ + + S +DYR+ R L+ + E + K D + + S I
Sbjct: 272 CDTLQLDSGVDYRK----RELAPAGKLYYLTSEADVEAVVDKLFDELAQKQNDLTSPRIL 327
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKA 387
+ GR L + ++C VA TF+ LC RPLGA+DY+ +++N+ TV I +IP S+ R +A
Sbjct: 328 KVQGRELRLNKACGSVADCTFEELCERPLGASDYLELSKNFDTVIIRNIPQFSLAKRTQA 387
Query: 388 RRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEGSKLRRDVLA 447
RRFITLID Y+ + C AS+ I LF + ++E++ S++ D L
Sbjct: 388 RRFITLIDNFYDFKVRIICSASAPISSLFXHQHQ-----------DSESDQSRILMDDLG 436
Query: 448 EGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+G S+ +G+EE+F FQR +SRL EMQT+ Y
Sbjct: 437 LSQDSAG------LSMFTGEEEIFAFQRTISRLTEMQTEQY 471
>emb|CAI95645.1| lactation elevated 1 [Homo sapiens] gi|66347711|emb|CAI95324.1|
lactation elevated 1 [Homo sapiens]
gi|66347679|emb|CAI95698.1| lactation elevated 1 [Homo
sapiens] gi|66347383|emb|CAI95701.1| lactation elevated
1 [Homo sapiens] gi|21918872|ref|NP_660358.2| lactation
elevated 1 [Homo sapiens] gi|21668098|gb|AAM74228.1|
lactation elevated 1 [Homo sapiens]
gi|37589913|gb|AAH18445.2| Lactation elevated 1 [Homo
sapiens]
Length = 481
Score = 193 bits (490), Expect = 1e-47
Identities = 128/401 (31%), Positives = 204/401 (49%), Gaps = 58/401 (14%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKP 148
P+GLY+YG+VG+GKTM+MDMFY+ E +K ++R HFH ML +++ +H+ + + KP
Sbjct: 129 PRGLYVYGDVGTGKTMVMDMFYAYVE--MKRKKRVHFHGFMLDVHKRIHRLKQSLPKRKP 186
Query: 149 LQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILC 208
F K+ + +A I +++++ A +LC
Sbjct: 187 ------------GFMAKSYDPIAP------------IAEEISEE-----------ACLLC 211
Query: 209 FDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEH 268
FDE Q D+ + L + L +G ++VATSNR P+DL + + F ++ L+E+
Sbjct: 212 FDEFQVTDIADAMILKQLFENLFKNGVVVVATSNRPPEDLYKNGLQRANFVPFIAVLKEY 271
Query: 269 CEKVLVGSEIDYRRFIAQRSENRVNYLWPIERET-INKFEKKWQDATGRFGGKVISNTIS 327
C V + S IDYR+ R L+ + E + K D + + I
Sbjct: 272 CNTVQLDSGIDYRK----RELPAAGKLYYLTSEADVEAVMDKLFDELAQKQNDLTRPRIL 327
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKA 387
+ GR L + ++C VA TF+ LC RPLGA+DY+ +++N+ T+F+ +IP ++ R +
Sbjct: 328 KVQGRELRLNKACGTVADCTFEELCERPLGASDYLELSKNFDTIFLRNIPQFTLANRTQG 387
Query: 388 RRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEGSKLRRDVLA 447
RRFITLID Y+ + C AS+ I LF ++E E S++ D L
Sbjct: 388 RRFITLIDNFYDLKVRIICSASTPISSLFLHQHH-----------DSELEQSRILMDDLG 436
Query: 448 EGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+ G S+ +G+EE+F FQR +SRL EMQT+ Y
Sbjct: 437 LSQDSAEG-----LSMFTGEEEIFAFQRTISRLTEMQTEQY 472
>ref|XP_419799.1| PREDICTED: similar to lactation elevated 1; CG8520 gene product;
lactation elevated-1 [Gallus gallus]
Length = 1272
Score = 193 bits (490), Expect = 1e-47
Identities = 136/413 (32%), Positives = 208/413 (49%), Gaps = 69/413 (16%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKP 148
PKGLY+YG+VG+GKTM+MDMFYS + V+ ++R HFH ML +++ +H+ + + KP
Sbjct: 319 PKGLYVYGDVGTGKTMVMDMFYSHLK--VERKKRVHFHGFMLDVHQRIHRLKQSLPKRKP 376
Query: 149 LQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILC 208
F K+ + +A VA++ + A +LC
Sbjct: 377 ------------GFMAKSYDPIAP----------------VAEEI-------SEEACLLC 401
Query: 209 FDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL-----NEANMVPEFFQNLLS 263
FDE Q D+ + L + L +G ++VATSNR P+DL AN VP L
Sbjct: 402 FDEFQVTDIADAMILKQLFENLFQNGVVVVATSNRPPEDLYKNGLQRANFVPFIAVLKLL 461
Query: 264 NLE--------EHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATG 315
L ++C V + S IDYR+ + + ++ YL + K D
Sbjct: 462 LLGGCVTASSCKYCSTVQLDSGIDYRKRVLPAA-GKLYYL--TSEADVEAVMDKLFDELA 518
Query: 316 RFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISD 375
+ + I + GR L + ++C +A FTF+ LC RPLGA+DY+ +++++ TVF+ D
Sbjct: 519 QKQNDLTRPRILKVQGRELRLNKACGTIADFTFEELCDRPLGASDYLEISKHFDTVFVRD 578
Query: 376 IPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETE 435
IP ++M R +ARRFITLID Y H + C A++ + LF E G++ E
Sbjct: 579 IPPLTMAKRTQARRFITLIDTFYEHKVRIICSAATPLQSLFV-VEAGSI----------E 627
Query: 436 AEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
E S++ D L + G S+ +G+EE+F FQR +SRL EMQT+ Y
Sbjct: 628 LEDSRVLMDDLDLSQDSAKG-----LSMFTGEEEIFAFQRTISRLTEMQTEQY 675
>gb|AAA91360.1| putative ATPase gi|1171633|sp|P46441|N2B_HAEIR PUTATIVE ATPASE N2B
(HFN2B)
Length = 464
Score = 191 bits (484), Expect = 6e-47
Identities = 144/476 (30%), Positives = 230/476 (48%), Gaps = 70/476 (14%)
Query: 18 ERRRILMDEVEKQ--QNDKDWWKRLNNKITERWTNSRKRPENVDP--GVGKWVSYLKREK 73
E + +L D+V+K+ Q +D + L N + +P V+ G G + ++K+E+
Sbjct: 46 ESKELLPDKVQKKTTQELEDLYNTLKNY--------QPKPVRVETSSGGGFFGRFMKKEQ 97
Query: 74 KLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRIN 133
+ TAP KG+YIYG+VG GKT LMDMFYS + I K ++R HF+ M +++
Sbjct: 98 SAPKIELLNTTAP---KGMYIYGSVGGGKTTLMDMFYSCCDDIPK-KQRVHFNSFMSKVH 153
Query: 134 EHMHKTWKKQMEEKPLQSGISSWIMNLPFD-TKAKEWLAAEERYKKEVQMKHILPDVADK 192
+HK + E P +S LPFD T + A E +
Sbjct: 154 GLIHKV---KQERGPQDRAFNSE-KPLPFDPTLPVAEMIANESW---------------- 193
Query: 193 FFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEAN 252
++CFDE Q D+ + L + + L + G + +ATSNR P DL +
Sbjct: 194 -------------LICFDEFQVTDIADAMILKSLFTHLFNEGIVCIATSNRHPNDLYKNG 240
Query: 253 MVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQD 312
+ F + L C + S +DYRR IAQ + NY + + ++ E+ ++
Sbjct: 241 LQRSNFIPFIGVLLNRCNVAAMDSGVDYRR-IAQSGDT--NYFVTTQTDAKSQMERMFKI 297
Query: 313 ATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVF 372
+ + TI+ FGR L +C V F+ LC RPLG +DYI + + +HTV
Sbjct: 298 LCSQENDIIRPRTIT-HFGRDLTFQRTCGQVLDSNFEELCNRPLGGSDYIQIGQFFHTVL 356
Query: 373 ISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQF 432
I D+P +++ ++ + RRFITLID LY++ + A +D+LF T++ DL
Sbjct: 357 IHDVPQLTLLLKSQMRRFITLIDTLYDNRVRVVISAEVPLDQLFSFTDKPK--DL----- 409
Query: 433 ETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
A+ ++ D L G+ + S+ +G+EE+F F R +SRL EMQ + Y
Sbjct: 410 ---ADEQRMLMDDLKLGDTDTS------ASVFTGEEEMFAFDRTISRLYEMQKKEY 456
>emb|CAG03614.1| unnamed protein product [Tetraodon nigroviridis]
Length = 353
Score = 186 bits (471), Expect = 2e-45
Identities = 134/417 (32%), Positives = 212/417 (50%), Gaps = 71/417 (17%)
Query: 74 KLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRIN 133
KL+SL PP PKGLYIYG+VG+GKTMLMD+F+S + ++R HF+ ML I+
Sbjct: 1 KLESL-------PPPPKGLYIYGDVGTGKTMLMDLFHSCV--VTPRKKRVHFNTFMLDIH 51
Query: 134 EHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKF 193
+ +H+ +KQ +LP T K + + + +++++
Sbjct: 52 KRIHR--RKQ---------------SLPKRTLGKLFTYDP--------ISPVAVEISNET 86
Query: 194 FLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANM 253
L LCFDE Q DV + L + L SG ++VATSNR P DL + +
Sbjct: 87 CL-----------LCFDEFQVSDVADALVLKQLFQALFRSGVVLVATSNRPPDDLYKNGL 135
Query: 254 VPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDA 313
+ F + L+E C + S DYRR + + R YL + +++
Sbjct: 136 QRDTFLPFIDMLKERCHIFRLDSGTDYRR-LGKAGAARAFYLTR-NAGAEAALDALFEEL 193
Query: 314 TGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFI 373
+ R T+SV+ GR + + ++C +A TFD LCG+PLGA+DY+ +A ++ TVFI
Sbjct: 194 SFRQKSDTGPQTLSVL-GRPVTLQKTCGSIADCTFDELCGKPLGASDYLEMARHFDTVFI 252
Query: 374 SDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELF--QGTEEGTLFDLESFQ 431
++P ++ ++D+ARRF TLID Y+ + LA++ +D+LF G E+
Sbjct: 253 RNVPRLTRSLKDQARRFTTLIDNFYDKKVRVVLLAAAPVDQLFVLAGGED---------- 302
Query: 432 FETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+L R +L + + + + S+ + +EE+F FQR VSRL EMQT+ Y
Sbjct: 303 --------ELDRQLLDDLGLSAAAEQL---SLFTAEEEIFAFQRTVSRLEEMQTESY 348
>ref|XP_318985.2| ENSANGP00000014807 [Anopheles gambiae str. PEST]
gi|55236088|gb|EAA14419.2| ENSANGP00000014807 [Anopheles
gambiae str. PEST]
Length = 436
Score = 185 bits (469), Expect = 3e-45
Identities = 135/434 (31%), Positives = 198/434 (45%), Gaps = 62/434 (14%)
Query: 55 PENVDPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATE 114
P G+GKW S+ GR A APKGLYIYG+VG GKTMLMDMFY
Sbjct: 57 PPKPSTGIGKWFSF-----------GRAEKAVAAPKGLYIYGSVGGGKTMLMDMFYDCCA 105
Query: 115 GIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEE 174
+ +RR HF+ M ++ +H K + + +S PFD
Sbjct: 106 --IDRKRRVHFNSFMTDVHAKIHDIKSKHVRD-------ASNSKPQPFDP---------- 146
Query: 175 RYKKEVQMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSG 234
+ VA+ D + ++CFDE Q D+ + L + + L ++G
Sbjct: 147 -----------IKPVAELITQD-------SWMICFDEFQVTDIADAMILKRLFTYLFNNG 188
Query: 235 TIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNY 294
I+VATSNRAP DL + + F + L+ HC V + S +DYR A + E++ +
Sbjct: 189 VIVVATSNRAPDDLYKNGLQRSNFVPFIGVLKNHCNIVTLDSGVDYRT-AALKGESKHYF 247
Query: 295 LWPIERETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGR 354
N K +I FGR + ++C V TFD LC +
Sbjct: 248 DKSQGASDANASMDKLFKVLCSQENDMIRPKTFTHFGRNITFAKTCGQVLDSTFDELCDK 307
Query: 355 PLGAADYIAVAENYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDE 414
PLGA+D++ +A+ +HTV I DIP ++++++ + RRFITLID LY+ L A
Sbjct: 308 PLGASDFLQIAQFFHTVLIRDIPQLNLKLKSQTRRFITLIDTLYDSRVRLVVSADVPYKY 367
Query: 415 LFQGTEEGTLFDLESFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQ 474
LF + T E L D+ + T ++I +G+EE+F F+
Sbjct: 368 LFSNDAPDDM--------HTSDEHRMLMDDL----KITKDSTDAS-SNIFTGEEEVFAFE 414
Query: 475 RAVSRLIEMQTQLY 488
R VSRL EMQ+ Y
Sbjct: 415 RTVSRLAEMQSAEY 428
>ref|YP_222576.1| hypothetical protein BruAb1_1905 [Brucella abortus biovar 1 str.
9-941] gi|62196915|gb|AAX75215.1| conserved hypothetical
protein [Brucella abortus biovar 1 str. 9-941]
Length = 387
Score = 184 bits (468), Expect = 5e-45
Identities = 130/424 (30%), Positives = 198/424 (46%), Gaps = 95/424 (22%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
++ L L G+R KGLY++G VG GKTMLMDMF+ V+ +RR HF++ M
Sbjct: 53 RKSSALGWLFGKRRETAATIKGLYVHGEVGRGKTMLMDMFFQLLP--VERKRRAHFNDFM 110
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK-EVQMKHILPD 188
++E ++ A + +K+ E + +P
Sbjct: 111 ADVHERIY---------------------------------AHRQAHKRGETKQDDPIPP 137
Query: 189 VADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL 248
VA E + A +LCFDE D+ + LS + S L S G ++VATSN AP +L
Sbjct: 138 VA-------EALSQQAWVLCFDEFAVTDIADAMILSRLFSALFSRGVVLVATSNVAPDNL 190
Query: 249 NEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEK 308
+ + F + L+ H + + + S DYR ++ + + YL P+ ET + +
Sbjct: 191 YRDGLNRQLFLPFIDILKSHVDVINLDSRTDYR---LEKLDRQPVYLSPLGPETERRMDA 247
Query: 309 KWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENY 368
W A + G + + I + GR +EVP + G ARF+FD LC RPLGA+DYIA+A Y
Sbjct: 248 AW--AAHKNGAEEKPDVIHIK-GRDIEVPRAAAGAARFSFDDLCARPLGASDYIAIANRY 304
Query: 369 HTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLE 428
T+FI ++P++ R++A+RFI LID LY+HH+ L A + ++L+
Sbjct: 305 PTLFIDNVPVLDYSRRNEAKRFILLIDVLYDHHARLFVSAEAQPEKLY------------ 352
Query: 429 SFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+ + GT E F F R SRL EMQ+ Y
Sbjct: 353 ----------------------IANSGT------------EAFEFDRTASRLFEMQSAEY 378
Query: 489 LDGV 492
LDG+
Sbjct: 379 LDGI 382
>gb|AAN30821.1| conserved hypothetical protein [Brucella suis 1330]
gi|23502779|ref|NP_698906.1| hypothetical protein BR1929
[Brucella suis 1330]
Length = 387
Score = 184 bits (468), Expect = 5e-45
Identities = 130/424 (30%), Positives = 198/424 (46%), Gaps = 95/424 (22%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
++ L L G+R KGLY++G VG GKTMLMDMF+ V+ +RR HF++ M
Sbjct: 53 RKSSALGWLFGKRRETAATIKGLYVHGEVGRGKTMLMDMFFQLLP--VERKRRAHFNDFM 110
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK-EVQMKHILPD 188
++E ++ A + +K+ E + +P
Sbjct: 111 ADVHERIY---------------------------------AHRQAHKRGETKQDDPIPP 137
Query: 189 VADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL 248
VA E + A +LCFDE D+ + LS + S L S G ++VATSN AP +L
Sbjct: 138 VA-------EALSQQAWVLCFDEFTVTDIADAMILSRLFSALFSRGVVLVATSNVAPDNL 190
Query: 249 NEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEK 308
+ + F + L+ H + + + S DYR ++ + + YL P+ ET + +
Sbjct: 191 YRDGLNRQLFLPFIDILKSHVDVINLDSRTDYR---LEKLDRQPVYLSPLGPETERRMDA 247
Query: 309 KWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENY 368
W A + G + + I + GR +EVP + G ARF+FD LC RPLGA+DYIA+A Y
Sbjct: 248 AW--AAHKNGAEEKPDVIHIK-GRDIEVPRAAAGAARFSFDDLCARPLGASDYIAIANRY 304
Query: 369 HTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLE 428
T+FI ++P++ R++A+RFI LID LY+HH+ L A + ++L+
Sbjct: 305 PTLFIDNVPVLDYSRRNEAKRFILLIDVLYDHHARLFVSAEAQPEKLY------------ 352
Query: 429 SFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+ + GT E F F R SRL EMQ+ Y
Sbjct: 353 ----------------------IANSGT------------EAFEFDRTASRLFEMQSAEY 378
Query: 489 LDGV 492
LDG+
Sbjct: 379 LDGI 382
>gb|AAL51318.1| PUTATIVE ATPASE N2B [Brucella melitensis 16M]
gi|17986420|ref|NP_539054.1| PUTATIVE ATPASE N2B
[Brucella melitensis 16M] gi|25355785|pir||AC3269
probable ATPase N2B BMEI0136 [imported] - Brucella
melitensis (strain 16M)
Length = 403
Score = 184 bits (468), Expect = 5e-45
Identities = 130/424 (30%), Positives = 198/424 (46%), Gaps = 95/424 (22%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
++ L L G+R KGLY++G VG GKTMLMDMF+ V+ +RR HF++ M
Sbjct: 69 RKSSALGWLFGKRRETAATIKGLYVHGEVGRGKTMLMDMFFQLLP--VERKRRAHFNDFM 126
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK-EVQMKHILPD 188
++E ++ A + +K+ E + +P
Sbjct: 127 ADVHERIY---------------------------------AHRQAHKRGETKQDDPIPP 153
Query: 189 VADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL 248
VA E + A +LCFDE D+ + LS + S L S G ++VATSN AP +L
Sbjct: 154 VA-------EALSQQAWVLCFDEFTVTDIADAMILSRLFSALFSRGVVLVATSNVAPDNL 206
Query: 249 NEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEK 308
+ + F + L+ H + + + S DYR ++ + + YL P+ ET + +
Sbjct: 207 YRDGLNRQLFLPFIDILKSHVDVINLDSRTDYR---LEKLDRQPVYLSPLGPETERRMDA 263
Query: 309 KWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENY 368
W A + G + + I + GR +EVP + G ARF+FD LC RPLGA+DYIA+A Y
Sbjct: 264 AW--AAHKNGAEEKPDVIHIK-GRDIEVPRAAAGAARFSFDDLCARPLGASDYIAIANRY 320
Query: 369 HTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLE 428
T+FI ++P++ R++A+RFI LID LY+HH+ L A + ++L+
Sbjct: 321 PTLFIDNVPVLDYSRRNEAKRFILLIDVLYDHHARLFVSAEAQPEKLY------------ 368
Query: 429 SFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+ + GT E F F R SRL EMQ+ Y
Sbjct: 369 ----------------------IANSGT------------EAFEFDRTASRLFEMQSAEY 394
Query: 489 LDGV 492
LDG+
Sbjct: 395 LDGI 398
>ref|NP_595259.1| ATPase [Schizosaccharomyces pombe 972h-] gi|2956750|emb|CAA17914.1|
SPBC115.02c [Schizosaccharomyces pombe]
gi|7492423|pir||T39297 probable atpase - fission yeast
(Schizosaccharomyces pombe)
Length = 454
Score = 184 bits (467), Expect = 6e-45
Identities = 131/429 (30%), Positives = 191/429 (43%), Gaps = 73/429 (17%)
Query: 65 WVSYLKR---EKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRR 121
W+S LK+ KK +L P P PKG+Y+YG+VG GKT LMD+FY V +
Sbjct: 92 WISPLKKMFSRKKSPTLTSSLPV-PGMPKGIYLYGDVGCGKTALMDLFYHNLPPNVTRSQ 150
Query: 122 RYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQ 181
R HFH M++++ H ++RY E+
Sbjct: 151 RIHFHAFMMQVHRTSHDL---------------------------------QDRYGFEID 177
Query: 182 -MKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVAT 240
+ HI +A K +LCFDE+Q DV + L + L+ G +I T
Sbjct: 178 FIDHIASGIA-----------KETTVLCFDELQVTDVADALLLRRLFEALMKYGVVIFIT 226
Query: 241 SNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIER 300
SNRAP DL + + E F + LE + + + S DYRR +S+ YL+P
Sbjct: 227 SNRAPSDLYKNGIQRESFIPCIKLLEHRLQVICLDSPNDYRRL---KSKTEDTYLYPANS 283
Query: 301 ETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAAD 360
+ K + W + + V FGR + VP++ VA FTF+ LCG P AAD
Sbjct: 284 PEVKKALENWFLCYADEKDPAHQDEVEV-FGRKIIVPKASGNVAWFTFEQLCGEPKSAAD 342
Query: 361 YIAVAENYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTE 420
Y+++A YH +SDIP +S+ +D RFIT ID LY+ H L + + E++
Sbjct: 343 YLSLASRYHVFIVSDIPKLSIESKDLIHRFITFIDALYDTHGKLILSSEVPVQEIYP--- 399
Query: 421 EGTLFDLESFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRL 480
+E D A+G + S G EE+FTF R +SRL
Sbjct: 400 ------------TAPSEVLSSTADPAAKGKIES-----HYHGAFGGIEEVFTFTRCLSRL 442
Query: 481 IEMQTQLYL 489
EM+ Q ++
Sbjct: 443 SEMKKQSWI 451
>ref|XP_624398.1| PREDICTED: similar to ENSANGP00000014807 [Apis mellifera]
Length = 436
Score = 184 bits (467), Expect = 6e-45
Identities = 133/416 (31%), Positives = 212/416 (49%), Gaps = 62/416 (14%)
Query: 75 LDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINE 134
LD +GR+ PP KGLY+YG VG GKTMLMD+FY + +K+++R HFH ML ++
Sbjct: 73 LDKWLGRKRKQPP--KGLYLYGAVGGGKTMLMDLFYQCCQ--IKNKKRVHFHSFMLDVHN 128
Query: 135 HMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFF 194
+H+ +K + +SS + PFD +P VA
Sbjct: 129 KVHEV------KKNIVRDVSSTKLQ-PFDP---------------------IPPVARSI- 159
Query: 195 LDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMV 254
+ A +LCFDE Q D+ + L + + L ++G I++ATSNRAP DL + +
Sbjct: 160 ------TEEAWLLCFDEFQVTDIADAMILKRLFTELFNNGVIVIATSNRAPDDLYKNGLQ 213
Query: 255 PEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDAT 314
F + L+ +C + S IDYR +E ++ ++ ++ I+ +K ++ +
Sbjct: 214 RGNFIPFIQVLKNYCIINSLDSGIDYRLKTGLGNE-KIYFI--KGKDAISDVDKVFKYLS 270
Query: 315 GRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFIS 374
+ V S TI + GR + ++C + TF+ LC RPLGA+DY+ +++ +HTV I
Sbjct: 271 SKENDVVRSRTICIR-GRNVTFKKTCGQILDSTFEELCDRPLGASDYLELSQAFHTVIIR 329
Query: 375 DIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELF--QGTEEGTLFDLESFQF 432
D+P ++ R++ + RRFITLID LY++ + A+ +LF +G E T
Sbjct: 330 DVPQLNFRLKSQTRRFITLIDTLYDNKVRVVISAAVPHTKLFVPEGNNEYT--------- 380
Query: 433 ETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
E L D+ S G+ +++ +G+EELF F R +SRL EMQT Y
Sbjct: 381 ---DEKRMLMDDLKI-----SHGSDDYKSNLFTGEEELFAFDRTISRLSEMQTTQY 428
>gb|EAL24818.1| GA21133-PA [Drosophila pseudoobscura]
Length = 468
Score = 174 bits (440), Expect = 8e-42
Identities = 123/402 (30%), Positives = 194/402 (47%), Gaps = 55/402 (13%)
Query: 88 APKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEK 147
AP GLY+YG+VG GKT LMD+FY I + ++R HF M ++ +H+ ++Q
Sbjct: 113 APMGLYLYGSVGVGKTTLMDLFYDCCTQIPR-KQRVHFTAFMNSVHGRIHEAKERQ---G 168
Query: 148 PLQSGISSWIMNLPFD-TKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANI 206
P+ +S PFD TK + A E + +
Sbjct: 169 PVDRAFNSE-KPAPFDPTKPVADIIARESW-----------------------------L 198
Query: 207 LCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLE 266
+CFDE Q D+ + L + + L G +I+ATSNR P+DL + + F ++ L+
Sbjct: 199 ICFDEFQVTDIADAMVLKRLFTHLFRKGIVIIATSNRHPEDLYKNGLQRTNFLPFIALLQ 258
Query: 267 EHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTI 326
C+ + S IDYRR IAQ + NY + + + ++ + TI
Sbjct: 259 RRCQLAKLDS-IDYRR-IAQSGDT--NYFVKGQSDAEGSMNRMFKILCAEENDIIRPRTI 314
Query: 327 SVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDK 386
+ FGR L +C V +F+ LC RPL +DY+ +++ +HTV I D+P +++ I+ +
Sbjct: 315 T-HFGRDLTFVRTCGQVLDSSFEELCNRPLAGSDYLQISQFFHTVLIRDVPSLNLNIKSQ 373
Query: 387 ARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEGSKLRRDVL 446
RRFITLID LY++ + A +D LFQ T+ + D + +L
Sbjct: 374 MRRFITLIDTLYDNRVRVVISADYPLDNLFQVTDPADISDTDR---------------IL 418
Query: 447 AEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+ GT +S+ +G+EELF F+R +SRL EMQ + Y
Sbjct: 419 MDDLKIKHGTHESKSSVFTGEEELFAFERTISRLYEMQKREY 460
>gb|AAK85448.1| Hypothetical protein C30F12.2 [Caenorhabditis elegans]
gi|17505769|ref|NP_491986.1| atpase (51.4 kD) (1H544)
[Caenorhabditis elegans]
Length = 445
Score = 169 bits (429), Expect = 1e-40
Identities = 118/401 (29%), Positives = 194/401 (47%), Gaps = 58/401 (14%)
Query: 88 APKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEK 147
+P+G+Y+YG+VG GKTMLMD+F+ + +RR HF++ M +++ MH E
Sbjct: 89 SPRGIYLYGSVGCGKTMLMDLFFENCP--IDKKRRVHFNDFMQNVHKRMH--------EL 138
Query: 148 PLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANIL 207
+QS + FD +P + D+ + N+L
Sbjct: 139 KMQSNKARG----KFDP---------------------VPVIVDEIM-------ETTNLL 166
Query: 208 CFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEE 267
CFDE Q D+ + L S L G ++VATSNRAP +L + + F ++ LE+
Sbjct: 167 CFDEFQVTDIADAMILKRFFSMLFDRGLVMVATSNRAPSELYKNGLQRHQFMPFITILED 226
Query: 268 HCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTIS 327
C + + S +DYRR A N+++ ++ + + ++ + V S T+
Sbjct: 227 KCASLALDSGMDYRRSAAGDG----NHVYFSSEDSNTQCDIVFKQSAANENDTVRSKTLE 282
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKA 387
++ GR + V + C GVA F LC GA DY+ A +HTV + +IP+M+ + +
Sbjct: 283 IL-GRRVIVEKCCGGVADVDFKELCMTAKGAVDYLVYARVFHTVIVRNIPIMNQDMWNAM 341
Query: 388 RRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQFETEAEGSKLRRDVLA 447
RRFIT+ID Y+ + A++ +DELFQ F+ + + ++ ++ D L
Sbjct: 342 RRFITMIDTFYDQKVRVVIGAAAPLDELFQ-------FEGHNTSHDALSDSKRMLMDDLG 394
Query: 448 EGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
+ G + ++ SG EE F + R VSRL EMQT+ Y
Sbjct: 395 IKSDHEGMS----ANVFSGDEEAFAYSRTVSRLYEMQTEKY 431
>gb|AAF14644.1| 7138.4 [Leishmania major]
Length = 478
Score = 148 bits (373), Expect = 5e-34
Identities = 122/426 (28%), Positives = 187/426 (43%), Gaps = 47/426 (11%)
Query: 70 KREKKLDSLVGRRPTAPPAP----KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHF 125
++EKK++ + A P KGLY++G VG GKTMLMD+ Y ++ +RR HF
Sbjct: 82 EQEKKVEQALSDDNKAVYHPLSQVKGLYVWGGVGCGKTMLMDLLYDNAPPEIR-KRRLHF 140
Query: 126 HEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFD-TKAKEWLAAEERYKKEVQMKH 184
H+ ML + + + K EE M P + T + ++ R + +
Sbjct: 141 HQFMLDMQKTSNSIRYKSKEE-----------MQDPANRTNMVRYNTSDNRRRTPDAEIN 189
Query: 185 ILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRA 244
+ +VA + D E +LCFDE+ DV + L + G +++ TSNR
Sbjct: 190 LFDEVAQRMISDVE-------LLCFDEVAVSDVAHAMILKRLFHSFYKIGLVVIFTSNRP 242
Query: 245 PKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETIN 304
+DL + + F + +++ C + S +D+R Q YL P+ E +
Sbjct: 243 LEDLYKDGLNRGGFIPFIDLVKKQCIIHHMKSNVDHRLLGHQAD----TYLTPMNSENNS 298
Query: 305 KFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAV 364
K EK + + + +FGR + VP +C GV F F LCG AADY +
Sbjct: 299 KLEKLFLEMCKAMPA---TERKLEVFGRDVIVPRACGGVCYFDFYELCGGEKSAADYEVI 355
Query: 365 AENYHTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTL 424
A+ +HT+FI+ +P D RF+ LID LY H + A+ +L EE
Sbjct: 356 AKTFHTIFINGVPQFPYENSDVKSRFLLLIDTLYGHRCKVMIHAAVEPPQLQAPKEEAA- 414
Query: 425 FDLESFQFETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQ 484
EG R D L+E SG V + + F R VSRL EM+
Sbjct: 415 ---------GRIEGDAQRVDQLSEFERESGNRLVDV------DDSAFQMDRCVSRLFEMR 459
Query: 485 TQLYLD 490
T+ YL+
Sbjct: 460 TKEYLE 465
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 896,527,409
Number of Sequences: 2540612
Number of extensions: 39158932
Number of successful extensions: 141737
Number of sequences better than 10.0: 234
Number of HSP's better than 10.0 without gapping: 193
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 140859
Number of HSP's gapped (non-prelim): 571
length of query: 520
length of database: 863,360,394
effective HSP length: 133
effective length of query: 387
effective length of database: 525,458,998
effective search space: 203352632226
effective search space used: 203352632226
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC145331.1


