

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
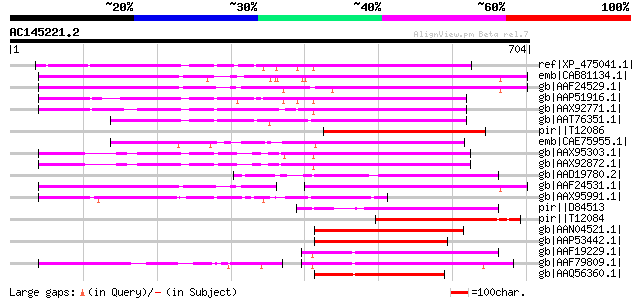
Query= AC145221.2 - phase: 0
(704 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_475041.1| OSJNBa0042F21.11 [Oryza sativa (japonica cultiv... 249 2e-64
emb|CAB81134.1| putative athila transposon protein [Arabidopsis ... 241 5e-62
gb|AAF24529.1| F7F22.15 [Arabidopsis thaliana] 234 9e-60
gb|AAP51916.1| putative polyprotein [Oryza sativa (japonica cult... 233 1e-59
gb|AAX92771.1| Transposable element protein, putative [Oryza sat... 230 1e-58
gb|AAT76351.1| hypothetical protein [Oryza sativa (japonica cult... 205 4e-51
pir||T12086 hypothetical protein - fava bean (fragment) gi|25222... 202 3e-50
emb|CAE75955.1| B1159F04.18 [Oryza sativa (japonica cultivar-gro... 193 2e-47
gb|AAX95303.1| hypothetical protein [Oryza sativa (japonica cult... 188 6e-46
gb|AAX92872.1| Transposable element protein, putative [Oryza sat... 187 8e-46
gb|AAD19780.2| hypothetical protein [Arabidopsis thaliana] 160 2e-37
gb|AAF24531.1| F7F22.17 [Arabidopsis thaliana] 156 2e-36
gb|AAX95991.1| Transposable element protein, putative [Oryza sat... 155 4e-36
pir||D84513 probable retroelement pol polyprotein [imported] - A... 149 4e-34
pir||T12084 hypothetical protein - fava bean (fragment) gi|25222... 144 1e-32
gb|AAN04521.1| Hypothetical protein with similarity to putative ... 143 2e-32
gb|AAP53442.1| hypothetical protein similar to putative retroele... 141 6e-32
gb|AAF19229.1| Similar to Athila ORF 1 [Arabidopsis thaliana] gi... 141 8e-32
gb|AAF79809.1| T32E20.9 [Arabidopsis thaliana] 139 4e-31
gb|AAQ56360.1| hypothetical protein OSJNBa0017M13.8 [Oryza sativ... 134 1e-29
>ref|XP_475041.1| OSJNBa0042F21.11 [Oryza sativa (japonica cultivar-group)]
gi|38347309|emb|CAE02304.2| OSJNBa0042F21.11 [Oryza
sativa (japonica cultivar-group)]
Length = 920
Score = 249 bits (637), Expect = 2e-64
Identities = 188/657 (28%), Positives = 304/657 (45%), Gaps = 85/657 (12%)
Query: 35 EKQVELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSL 94
+KQ+++K + ++ PF G +ED HL F + + +A LRLF SL
Sbjct: 251 DKQIKIKKFKPRDKAS-PFCGKPNEDANAHLLQFLEICSMYTIKGVSPDAVRLRLFPFSL 309
Query: 95 IGKAREWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWER 154
+ +A++W+ + W+ +F +FFP K + I+ F Q DES+ +AWER
Sbjct: 310 LRRAKQWFYANCA-AINTWDKCSTSFLSKFFPIGKTNALRGRISRFQQTRDESIPEAWER 368
Query: 155 YKSLLRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLN 214
+ + CP HG DD + F GL + LDAAA G+ + + + AV++I KM N
Sbjct: 369 LQEYVAACPHHGMDDWLILQNFYNGLTLMSRDHLDAAAGGAFFSKTVQGAVELIEKMVSN 428
Query: 215 DRAAKHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQR 274
+K QR ++ +T+LL +++ +MK + + K + ++
Sbjct: 429 TGWSKERLQTRQRGMHTVK---------ETELLAAKLDLLMKCLDDHEKRPQGTVKALDS 479
Query: 275 HQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQR--FPNNNNF 332
H CE+C G +G + EE Y+G + +++ Q NQ R +P NN
Sbjct: 480 HVT---CEVCGGTGHSGNDCLETHEEAMYMG---KNNEYRPQGGEGWNQPRPYYPGGNN- 532
Query: 333 QRGNNFRQPW---------KSNAGPSNQQPFNNNFQQ------------YPPQQDRTQKI 371
GN QP K+ + N+ + + Q + I
Sbjct: 533 -NGNFSNQPSLMDLVFAQAKTTDALRKKLAANDKILENINVKLDGFASAFQNQLSFNKMI 591
Query: 372 EDTLNQFMQLTMANQQN-----------------------------------NRIHKE-- 394
E L Q L AN+ N + KE
Sbjct: 592 ETQLAQLASLVPANESGRIPGQPDSSIENVKAITTRGGKSTRDPPYPNPAGTNGMSKETP 651
Query: 395 SDVEKDKKKEVGKNVP-----AHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEAL 449
S DK+ + K VP LP+P+ K ++K F F+++ +++ IN+P +A+
Sbjct: 652 STDSDDKEIQPDKTVPQEYCDTRLLPFPQQSRKPSVDKQFACFVEVIQKIHINVPLLDAM 711
Query: 450 EQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVN 509
Q+PTYA+++K+IL+ K E ++L +CS +I +LP+K+KDPG T+ +IG
Sbjct: 712 -QVPTYARYLKDILNNKRPLPTTEVVKLTEHCSNLILHKLPEKKKDPGCPTITCSIGAQQ 770
Query: 510 VGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFV 569
+AL DLG+S++++P V ++ + T M LQLAD SV P+GIAEDV K+ F
Sbjct: 771 FDQALCDLGASVSVMPKDVFDKLNFTVLAPTPMRLQLADSSVHYPVGIAEDVPAKIRDFF 830
Query: 570 FPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEEINFNLHSAMK 626
PVDFVV+D++ + PLILGR F+ A ID G ++ + +E F M+
Sbjct: 831 IPVDFVVLDMDTGKETPLILGRPFLSTAGANIDVGTGSIRFHTNGKEEKFEFQPRME 887
>emb|CAB81134.1| putative athila transposon protein [Arabidopsis thaliana]
gi|25354974|pir||B85075 probable athila transposon
protein [imported] - Arabidopsis thaliana
Length = 866
Score = 241 bits (616), Expect = 5e-62
Identities = 198/748 (26%), Positives = 333/748 (44%), Gaps = 119/748 (15%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
E+KSG I ++ + F GL EDP HL F L N + + LRLF SL KA
Sbjct: 40 EIKSGLISMIQGNKFYGLPMEDPLDHLDEFDRLCNLTKINGVSADGFKLRLFPFSLGDKA 99
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
W + D + W+ +KAF +FF + + + I+ F+Q ES C+AWER+K
Sbjct: 100 HIWEKNLPHDSIITWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGY 159
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+CP HGF A+ + G+ P +++LD A++G+ E ++++ ++ +D
Sbjct: 160 TNQCPHHGFTKASMLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVDNLAQSDGNY 219
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEE-MMKQMKEMPKLVKE---QIQKEQR 274
+ T R + S+D + + K L +++ ++ Q K + LV + Q+Q +
Sbjct: 220 NEDCDRTVRDTA----DSDDKHRKEIKALNDKLDRILLNQQKHVHFLVDDEQFQVQDGEG 275
Query: 275 HQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNN--F 332
+Q EEV+Y+ N Q SG F NN
Sbjct: 276 NQL---------------------EEVSYINNNQ---------SGYKGYNNFKTNNPNLS 305
Query: 333 QRGNNFRQPWKSNAGPSNQQ-------PFNNNF---QQY---------------PPQQDR 367
R N P P QQ P+N F QQ+ P +++
Sbjct: 306 YRSTNIANPQDQVYPPQQQQVQNKPFVPYNQGFVPKQQFQGNYQPPPPPGGKALPTREEP 365
Query: 368 TQKIEDTLNQFMQ-LTMANQQNNRIHKES-----------------DVEKD--------- 400
ED+ +Q + L++ Q ++ H++ D+ D
Sbjct: 366 KTVTEDSEDQDGEDLSLEKDQADKPHEQPLDQSLEQPLDLSLEQPLDLPLDNVTRPTTRP 425
Query: 401 ---------------KKKEVGKNVPAHK--LPYPKAQSKKDLEKHFKRFLDIFKRLEINI 443
K KE P +K LP+P K +K+ F K +E+ I
Sbjct: 426 IFPAASATAPKPITVKNKEKVFVPPPYKPELPFPGRHKKALADKYRAMFAKNIKEVELRI 485
Query: 444 PFSEALEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRL-PKKEKDPGRVTLP 502
P +AL +P KF+K+++ ++ + + + L CSAIIQ+++ PKK DPG TLP
Sbjct: 486 PLVDALALIPDSHKFLKDLIVERIQEVQGMVV-LSHGCSAIIQKKIIPKKLSDPGSFTLP 544
Query: 503 VAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVL 562
++G + + L DLG+S++L+ SV KR+G K + ++L LAD+SV P G+ ++
Sbjct: 545 CSLGPLAFNRCLCDLGASVSLMALSVAKRLGFTQYKSSNISLILADRSVRIPHGLLKNFG 604
Query: 563 VKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRL-DDEEINFNL 621
+ + P DFVV++++E+ PLILGR F+ A MID G + + L D + F++
Sbjct: 605 ITIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLSLGKDFRMTFDV 664
Query: 622 HSAMKHSKDKGACFKVDAMDEVIMETRKQLHKPTPLERVLTDA-------LSVLSAEEEK 674
AMK +G F + MD++ E ++L + L LT + L ++
Sbjct: 665 KDAMKKPTIEGQLFWIKEMDQLADELLEELAEEDHLNSALTKSGEDGFLHLETFGYQKLL 724
Query: 675 EIEECLKELETLKEIPPKKAKVEELKKE 702
+ + ++E E +E+ +V + +E
Sbjct: 725 DSNKAMEESEPFEELNEPATEVMVMSEE 752
>gb|AAF24529.1| F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 234 bits (596), Expect = 9e-60
Identities = 186/717 (25%), Positives = 325/717 (44%), Gaps = 88/717 (12%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
E++ + + F GL EDP HL F L N + E+ LRLF SL KA
Sbjct: 158 EIRKRRTTVNPGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKA 217
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
W + D + W+ +KAF +FF + + + I+ F+Q ES C+AWER+K
Sbjct: 218 HIWEKNLPHDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGY 277
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+C HGF A+ + G+ P +++LD A++G+ E +++ ++ ++
Sbjct: 278 TNQCSHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSNGNY 337
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEE-MMKQMKEMPKLVKEQIQKEQRHQQ 277
N T R + S+D + + K L +++ ++ Q K + LV ++ + Q +
Sbjct: 338 NENCDRTVRGTA----DSDDKHRKEIKALNDKLDRILLSQHKHVHFLVDDEQYEVQDGEG 393
Query: 278 VHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNN 337
EEV+Y+ N QG Y G N + N N R N
Sbjct: 394 NQL------------------EEVSYINNN------QGGYKG-YNNFKTNNPNLSYRSTN 428
Query: 338 FRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQK-----IEDTLNQFMQLTMANQQNNRIH 392
P P QQ N F Y +T + +++ +T+ + +
Sbjct: 429 VANPQDQVYPPQQQQSQNKPFVPYNQATKQTSQLPGKAVQNPKEYAHAITLHSGKALPTR 488
Query: 393 KE-SDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLEKHFKRFLDIF--------------- 436
+E V +D + + G++ L K Q+ K L+ ++ LD+
Sbjct: 489 EEPKTVTEDSEDQDGED-----LSLKKDQADKPLDLSLEQPLDLSLQQSLDPPLDSFTRP 543
Query: 437 ----------------------KRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDET 474
+++E+ IP +AL +P KF+K+++ ++ + +
Sbjct: 544 TTRPVIPAASPTAPKPVAVKNKEKVELRIPLVDALALIPDSHKFLKDLIVERIQEVQGMV 603
Query: 475 IQLDANCSAIIQRR-LPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVG 533
+ L CSAIIQ++ +PKK DPG TLP ++G + + L DLG+S++L+P SV KR+G
Sbjct: 604 V-LSHECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLG 662
Query: 534 GLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTF 593
K ++L LAD+SV P G+ E++ +++ P DFVV++++E+ PLILGR F
Sbjct: 663 FTQYKSCNISLILADRSVRIPHGLLENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPF 722
Query: 594 MLAARMMIDYDDGLMKVRL-DDEEINFNLHSAMKHSKDKGACFKVDAMDEVIMETRKQLH 652
+ A MID G + + L D + F++ AMK +G F ++ MD++ E ++L
Sbjct: 723 LATAGAMIDVKKGKIDLNLGKDFRMTFDVKDAMKKPTIEGQLFWIEEMDQLADELLEELA 782
Query: 653 KPTPLERVLTDA-------LSVLSAEEEKEIEECLKELETLKEIPPKKAKVEELKKE 702
+ L LT + L L ++ + + ++E E +E+ +V + +E
Sbjct: 783 EEDHLNSALTKSGEDGFLHLETLGYQKLLDSHKAMEESEPFEELNGPATEVMVMSEE 839
>gb|AAP51916.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37530654|ref|NP_919629.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|20042899|gb|AAM08727.1| Putative polyprotein [Oryza
sativa] gi|18997221|gb|AAL83338.1| Putative polyprotein
[Oryza sativa (japonica cultivar-group)]
Length = 689
Score = 233 bits (595), Expect = 1e-59
Identities = 171/611 (27%), Positives = 284/611 (45%), Gaps = 68/611 (11%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
+LKS I + PF G +ED HL F + ++ + LRLF SL+G+A
Sbjct: 31 DLKSSHITMAQASPFCGKPNEDANAHLQQFLEICSTYTIKGVSPDVVKLRLFPFSLLGRA 90
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
++W+ V W+ AF +FFP +AWER +
Sbjct: 91 KQWFYANRTAV-NTWDKCSTAFLSKFFP----------------------MEAWERLQEY 127
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+ CP G DD + F GL P + LDAAA G+ + + + AV++I KM N +
Sbjct: 128 VAACPHLGMDDWLILQNFYNGLTPMSRDHLDAAARGAFFSKTVQGAVELIEKMVSNMGWS 187
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQV 278
+ QR ++ +T+LL +++ +MK++ + K + ++ H
Sbjct: 188 EERLQTRQRGMHTVK---------ETELLAAKLDLLMKRLDDHEKGPQRTVKALDSHVM- 237
Query: 279 HFCEMCSGDHPTGYCPPQLEEEVNYVGNQ--------QRQGQFQGQYSGNANQQRFPNNN 330
CE+C +G + EE Y+GN Q Q + Y G N FPN
Sbjct: 238 --CEVCGNTGHSGNDCLETCEEAMYMGNNNGYCPQGGQGWNQPRPYYQGGNNNSNFPNQP 295
Query: 331 NFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQ---KIEDTLNQFMQLTMANQQ 387
+ + F Q K+ S + N+ P + + + + ++ +TM +
Sbjct: 296 SL-KDLVFAQA-KTTDALSKKLAANDKLASLVPANETGRIPGQPDPSIENVKAITMRRGK 353
Query: 388 N------------NRIHK--ESDVEKDKKKEVGKNVP-----AHKLPYPKAQSKKDLEKH 428
+ N I K S+ DK+ + VP L +P+ K +++
Sbjct: 354 STRDPPYPNPVGTNEISKGAPSNDSADKEVQPENTVPQEYCDTRLLSFPQRMRKPSVDEQ 413
Query: 429 FKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRR 488
F RF+++ +++ IN+P +A+ Q+PTYA+++K+IL+ K E ++L CS +I +
Sbjct: 414 FARFVEVIQKIHINVPLLDAM-QVPTYARYLKDILNNKRPLPTTEVVKLTEQCSNLILHK 472
Query: 489 LPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLAD 548
LP+K+KDPG T+ +IG + +LG+S++++ V ++ + T M LQLAD
Sbjct: 473 LPEKKKDPGCPTITCSIGAQQFDQVSCNLGASVSVVLKDVFDKLNFTVLAPTPMRLQLAD 532
Query: 549 KSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLM 608
SV P GIAEDV VK+ F PVDFVV+D++ + PLILGR F+ A ID G +
Sbjct: 533 SSVRYPAGIAEDVPVKIRDFFIPVDFVVLDMDTGKETPLILGRPFLSTAGANIDVGTGSI 592
Query: 609 KVRLDDEEINF 619
+ ++ ++ F
Sbjct: 593 RFHINGKKEKF 603
>gb|AAX92771.1| Transposable element protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 689
Score = 230 bits (586), Expect = 1e-58
Identities = 169/589 (28%), Positives = 276/589 (46%), Gaps = 45/589 (7%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
+LKS I + F G +ED HL F + ++ +A LRLF SL+G+A
Sbjct: 31 DLKSSLIMMAQASLFCGKPNEDANAHLQQFLEICSTYTIKGVSPDAVRLRLFPFSLLGRA 90
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
++W+ V W+ AF +FFP K + I+ F Q DES+ +AWER
Sbjct: 91 KQWFYANHAAV-NTWDKCSTAFLSKFFPMGKTNALRGRISSFQQTRDESIPEAWERL--- 146
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
Q + F GL P + LD G+ + + + AV++I KM N +
Sbjct: 147 -------------QDYNFHNGLTPMSRNHLDTTTGGAFFSKTVQGAVELIEKMVSNMGWS 193
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQV 278
+ QR ++ +T+L+ ++E +MK + + K + ++ H
Sbjct: 194 EERLQTRQRGMHTVK---------ETELVAAKLELLMKHLDDHEKRPQGTVKALDSHVT- 243
Query: 279 HFCEMCSGDHPTGYCPPQLEEEVNYVGNQQ-RQGQFQGQYSGNANQQRFPNNNNFQRGNN 337
CE+C G G P+ EE Y+GN ++ + N N + + FQ +
Sbjct: 244 --CEVCGGTGHLGNDCPETREEAMYMGNNNGKKLAANDKILENINVKLDGFASAFQNQLS 301
Query: 338 FRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDV 397
+ + ++ N + P Q D ++ +TM + KE+
Sbjct: 302 YNKMIETQLARLQSLVPTNETGRIPGQPD------SSIENVKAITMTGGTIG-MSKEAPS 354
Query: 398 EKDKKKEVG--KNVP-----AHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALE 450
KE+ K VP LP+P+ K +++ F RF+++ +++ IN+P +A+
Sbjct: 355 NDSAGKEIQPEKTVPQEYCNTRLLPFPQRMRKPSVDEQFARFVEVIQKIHINVPLLDAM- 413
Query: 451 QMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNV 510
Q+PTYA ++K+IL+ K E ++L CS +I +LP+K+K PG T+ +I
Sbjct: 414 QVPTYAHYLKDILNNKRPLPTMEVVKLTEQCSNVILHKLPEKKKYPGCPTITCSIRAQQF 473
Query: 511 GKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVF 570
+AL DLG+S++++P V ++ + T M LQLAD SV P GIAEDV VK+ F
Sbjct: 474 DQALCDLGASVSVMPKDVFDKLKFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFT 533
Query: 571 PVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEEINF 619
PVDFVV+D++ + PLILGR F+ ID G + ++ +E F
Sbjct: 534 PVDFVVLDMDTGKETPLILGRPFLSTGGANIDMGTGSIHFHINRKEERF 582
>gb|AAT76351.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 585
Score = 205 bits (522), Expect = 4e-51
Identities = 144/493 (29%), Positives = 244/493 (49%), Gaps = 30/493 (6%)
Query: 137 IAMFAQGADESLCDAWERYKSLLRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSL 196
I+ F Q DES+ +AWE + + CP HG DD + F L P + LDAAA G+
Sbjct: 11 ISSFQQTRDESIPEAWELLQEYVAACPHHGMDDWLILQNFYNRLTPMSRDHLDAAAGGAF 70
Query: 197 MALSPREAVDIINKMSLNDRAAKHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMK 256
+ + + V++I KM N +K QR ++ +T+LL +++ +MK
Sbjct: 71 FSKTVQGTVELIEKMVSNMGWSKERLQTRQRGMHTVK---------ETELLAAKLDLLMK 121
Query: 257 QMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQ 316
++ K + ++ H CE+C G +G P+ EE Y+GN + QG
Sbjct: 122 RLDNHEKRPQGTVKALDSHVT---CEVCGGTDHSGNDCPETREEAMYMGNNNNGYRPQGV 178
Query: 317 YSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQ------QPFNN--NFQQYPPQQDRT 368
N + + NN N + + P+N+ QP+++ N + + ++
Sbjct: 179 QGWNQPRSYYQGGNN----NETQLAQLAYLVPANETGRIPGQPYSSIENVKAITTRGGKS 234
Query: 369 QKIEDTLNQFMQLTMANQ--QNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLE 426
+ N M+ + N+ KE EK +E LP+P+ K ++
Sbjct: 235 TRDPPYPNPAGTNGMSKETPSNDSADKEIQPEKTVPQEY---CDTRLLPFPQWSRKTSVD 291
Query: 427 KHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQ 486
+ F RF+++ +R+ IN+ +A+ Q+P YA+++K+IL+ K E ++L +CS +I
Sbjct: 292 ELFARFVEVIQRIHINVLLLDAM-QVPIYARYLKDILNNKRPLPTTEVVKLTEHCSNVIL 350
Query: 487 RRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQL 546
+LP+K+K T+ +IG +AL DLG+S++++P V R+ + T M LQ+
Sbjct: 351 HKLPEKKKYSECPTITCSIGAQQFDQALCDLGASVSVMPKDVFDRLNFTVLAPTPMRLQM 410
Query: 547 ADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDG 606
AD SV GIAEDV +K+ F PVDFVV+D++ + PLILGR F+ A ID +
Sbjct: 411 ADSSVRYQAGIAEDVPIKIWDFFIPVDFVVLDMDTRKETPLILGRPFLSTAGANIDVETS 470
Query: 607 LMKVRLDDEEINF 619
+ ++++E F
Sbjct: 471 SICFHINEKEEKF 483
>pir||T12086 hypothetical protein - fava bean (fragment)
gi|2522230|dbj|BAA22788.1| retrotransposon-like gene~the
first amino acid was determined to be leucine [Vicia
faba]
Length = 238
Score = 202 bits (514), Expect = 3e-50
Identities = 102/221 (46%), Positives = 157/221 (70%), Gaps = 2/221 (0%)
Query: 426 EKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAII 485
EK+F+ FL++FK+LE+NI EALE+MPTYAKFMK+I+SKK D I + CS+I+
Sbjct: 11 EKNFEIFLEMFKKLELNILVLEALEKMPTYAKFMKDIISKKRTIDTDPIIHTET-CSSIL 69
Query: 486 Q-RRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTL 544
Q ++P K+KD G VT P IG + K I+LG+S++L+P+ + K++G +++ TRMTL
Sbjct: 70 QGMKIPVKKKDRGFVTTPCTIGDRSFKKYFINLGASVSLLPFYIYKKLGISNVQNTRMTL 129
Query: 545 QLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYD 604
+ AD SV P GI EDVLVK+DKFVFPVDFVV+++ ED+++ LILGR F+ R +ID +
Sbjct: 130 RFADHSVKIPYGIVEDVLVKIDKFVFPVDFVVLEMPEDEEITLILGRPFLETRRCLIDIE 189
Query: 605 DGLMKVRLDDEEINFNLHSAMKHSKDKGACFKVDAMDEVIM 645
+G + +++ DEE+ ++ + MK+ D ++ +D+V++
Sbjct: 190 EGTVTLKVYDEELKIDVQNTMKYKDDIATSQHIEVIDQVVV 230
>emb|CAE75955.1| B1159F04.18 [Oryza sativa (japonica cultivar-group)]
gi|38605826|emb|CAE05235.3| OSJNBa0011K22.17 [Oryza
sativa (japonica cultivar-group)]
gi|50923135|ref|XP_471928.1| OSJNBa0011K22.17 [Oryza
sativa (japonica cultivar-group)]
gi|50923097|ref|XP_471909.1| B1159F04.18 [Oryza sativa
(japonica cultivar-group)]
Length = 551
Score = 193 bits (490), Expect = 2e-47
Identities = 153/499 (30%), Positives = 241/499 (47%), Gaps = 53/499 (10%)
Query: 137 IAMFAQGADESLCDAWERYKSLLRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSL 196
I+ F Q DES+ +AWER + + CP HG DD + F GL P LDAAA G+
Sbjct: 11 ISSFQQTRDESIPEAWERLQEYMAACPHHGMDDWLILQNFYNGLTPMSHDHLDAAARGAF 70
Query: 197 MALSPREAVDIINKMS--LNDRAAKHNRS--GTQR---KSGILELGSNDANLAQTKLLTQ 249
+ + + AVD+I KM L R H++ GT + E+ N + L T+
Sbjct: 71 FSKTVQGAVDLIEKMLDLLMKRLDDHDKKPQGTVKVLDSHVTCEVCGNTGHSGNDCLDTR 130
Query: 250 QMEEMMKQMKEMPKLVKEQIQK----EQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVG 305
+ M P+ + Q +++H+ C P L++ V
Sbjct: 131 EAMYMGNNNGYCPQRGQGWNQPCPYVDRKHEAWEIC--------LTPVQPSLKDLV---- 178
Query: 306 NQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQ 365
F + +A ++ N+ N + ++A NQ FN
Sbjct: 179 -------FAQAKTIDALSKKLATNDKILENINVKLDGFASAF-QNQLRFN---------- 220
Query: 366 DRTQKIEDTLNQFMQLTMANQQN-NRIHKE--SDVEKDKKKEVGKNVPAHK-----LPYP 417
+ +E L QF L AN+ N I KE S DK+ ++ K VP L +
Sbjct: 221 ---KMLETQLAQFASLVPANETGTNGIAKEVPSSDSADKEVQLEKIVPQEYCDTWLLSFH 277
Query: 418 KAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDETIQL 477
+ K +++ F RF ++ +++ IN+P +A+ Q+PTYA+++K++L+ K E ++L
Sbjct: 278 QRSRKPSVDEQFARFAEVIQKIHINVPLLDAM-QVPTYARYLKDMLNNKRLLPTMEVVKL 336
Query: 478 DANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDM 537
CS++I +LP+K+KDPG T+ +IG +A DLG+SI+++P V ++ +
Sbjct: 337 TEQCSSVILHKLPEKKKDPGCPTITCSIGAQQFDQAFCDLGASISVMPKDVPDKLNFTVL 396
Query: 538 KLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAA 597
T M LQLAD SV P GIAEDV VK+ F PVDFVV+D++ + PLILGR F+ A
Sbjct: 397 TPTLMCLQLADLSVHYPTGIAEDVPVKIRDFFIPVDFVVLDMDMGKETPLILGRPFLSTA 456
Query: 598 RMMIDYDDGLMKVRLDDEE 616
ID G ++ ++ +E
Sbjct: 457 GANIDVGMGSIRFDINGKE 475
>gb|AAX95303.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 553
Score = 188 bits (477), Expect = 6e-46
Identities = 151/596 (25%), Positives = 260/596 (43%), Gaps = 104/596 (17%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
+LKS I + PF G +ED HL F + ++ +A LRLF SL+G+A
Sbjct: 31 DLKSSLITMAQASPFCGKPNEDANAHLQQFLEICSTYTIKGISPDAVRLRLFPFSLLGRA 90
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
++W+ A+ + + W++ +L
Sbjct: 91 KQWFY----------------------------------------ANRAAVNTWDKCSTL 110
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+ + F GL P + LDAA G+ + R AVD+I M N +
Sbjct: 111 I-------------LQNFYNGLMPMSRNHLDAATGGAFFSKMVRGAVDLIENMVSNMGWS 157
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQV 278
+ QR ++ + +LL +++ + K + + K + ++ H
Sbjct: 158 EERLQTHQRGMHTVK---------EIELLATKLDLLRKHLDDDKKRPQVTVKALDSHVTY 208
Query: 279 HFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNF 338
E+CS +G P+ EE Y+GN N ++ Q G +
Sbjct: 209 ---EVCSNMGHSGNDCPETREEAMYMGNNN-------------------NGHHPQGGQGW 246
Query: 339 RQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKI---EDTLNQFMQLTMANQQNNRIHKE- 394
QP + + P + T++I D+ + ++ N I KE
Sbjct: 247 NQP--------RPDYYGDQLASLVPANE-TRRILGQPDSSVENVKAITTRGGTNGIAKEA 297
Query: 395 -SDVEKDKKKEVGKNVP-----AHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEA 448
S+ DK+ + K VP LP+P+ K ++ RF+++ +++ IN+P +A
Sbjct: 298 PSNDSADKEVQPEKTVPQIYCDTRLLPFPQRSRKPLADEQLARFVEVIQKIHINVPLLDA 357
Query: 449 LEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKV 508
+ Q+PTYA+++K+IL+ K E ++L C+ +I +LP+K+KDP T+ +I
Sbjct: 358 M-QVPTYARYLKDILNNKRLLPTTEVVKLTEQCNDVILHKLPEKKKDPRCPTITCSIRAH 416
Query: 509 NVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKF 568
+AL DLG+S++++P V ++ + T M LQLA+ SV P GIAE+V K+ F
Sbjct: 417 QFDRALCDLGASVSVMPKDVFDKLNFTVLASTPMRLQLANSSVRYPAGIAEEVPFKIRDF 476
Query: 569 VFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEEINFNLHSA 624
P DFVV+D++ + PLILGR+F+ ID G + ++ +E F S+
Sbjct: 477 FIPFDFVVLDMDTGKETPLILGRSFLSTTGANIDVGTGSILFHINGKEEKFEFQSS 532
>gb|AAX92872.1| Transposable element protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 548
Score = 187 bits (476), Expect = 8e-46
Identities = 147/591 (24%), Positives = 255/591 (42%), Gaps = 99/591 (16%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
+LKS I + PF G +ED HL F + ++ +A LRLF SL+G+A
Sbjct: 31 DLKSSLITMAQASPFCGKPNEDANAHLQQFLEICSTYTIKGISPDAVRLRLFPFSLLGRA 90
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
++W+ A+ + + W++ +L
Sbjct: 91 KQWFY----------------------------------------ANRAAVNTWDKCSTL 110
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+ + F GL P + LDAA G+ + R AVD+I M N +
Sbjct: 111 I-------------LQNFYNGLMPMSRNHLDAATGGAFFSKMVRGAVDLIENMVSNMGWS 157
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQV 278
+ QR ++ + +LL +++ + K + + K + ++ H
Sbjct: 158 EERLQTHQRGMHTVK---------EIELLATKLDLLRKHLDDDKKRPQVTVKALDSHVTY 208
Query: 279 HFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNF 338
E+CS +G P+ EE Y+GN N ++ Q G +
Sbjct: 209 ---EVCSNMGHSGNDCPETREEAMYMGNNN-------------------NGHHPQGGQGW 246
Query: 339 RQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVE 398
QP + + P + T++I + ++ A S+
Sbjct: 247 NQP--------RPDYYGDQLASLVPANE-TRRILGQPDSSVENVKAITTRGAKEAPSNDS 297
Query: 399 KDKKKEVGKNVP-----AHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMP 453
DK+ + K VP LP+P+ K ++ RF+++ +++ IN+P +A+ Q+P
Sbjct: 298 ADKEVQPEKTVPQIYCDTRLLPFPQRSRKPLADEQLARFVEVIQKIHINVPLLDAM-QVP 356
Query: 454 TYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKA 513
TYA+++K+IL+ K E ++L C+ +I +LP+K+KDP T+ +I +A
Sbjct: 357 TYARYLKDILNNKRLLPTTEVVKLTEQCNDVILHKLPEKKKDPRCPTITCSIRAHQFDRA 416
Query: 514 LIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVD 573
L DLG+S++++P V ++ + T M LQLA+ SV P GIAE+V K+ F P D
Sbjct: 417 LCDLGASVSVMPKDVFDKLNFTVLASTPMRLQLANSSVRYPAGIAEEVPFKIRDFFIPFD 476
Query: 574 FVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEEINFNLHSA 624
FVV+D++ + PLILGR+F+ ID G + ++ +E F S+
Sbjct: 477 FVVLDMDTGKETPLILGRSFLSTTGANIDVGTGSILFHINGKEEKFEFQSS 527
>gb|AAD19780.2| hypothetical protein [Arabidopsis thaliana]
Length = 1048
Score = 160 bits (404), Expect = 2e-37
Identities = 106/359 (29%), Positives = 172/359 (47%), Gaps = 42/359 (11%)
Query: 304 VGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPP 363
VGNQ + + G G A + + P + G K + +++ ++
Sbjct: 19 VGNQNSRSENSG---GRAPKHQNPEKGGHKSGRELNPILKKEKAKEKSKAASDDLEE--- 72
Query: 364 QQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKK 423
D T K DTL EK KK + VP KLP+P Q K
Sbjct: 73 --DNTGKESDTLE---------------------EKAKKTVLPPYVP--KLPFPGRQRKI 107
Query: 424 DLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSA 483
EK + F +I ++L++ +PF + ++ + Y K +K+IL+ K +E + + CSA
Sbjct: 108 QREKEYSLFDEIMRQLQVKLPFLDLVQNVSIYRKHLKDILTNKRTLEEGHVL-ISHECSA 166
Query: 484 IIQRRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMT 543
I+Q IG+ + L DLG+ ++L+P+SV KR+G + T+M+
Sbjct: 167 ILQNV----------ALFSRMIGEYTFDRCLCDLGAGVSLMPFSVAKRLGDTNFTPTKMS 216
Query: 544 LQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDY 603
L L D+S++ P+G+AEDV V+V F P DFV+++++E+ LILGR F+ +ID
Sbjct: 217 LVLGDRSISFPVGVAEDVQVRVGNFYIPTDFVIIELDEEPRHRLILGRPFLNIVAALIDV 276
Query: 604 DDGLMKVRLDDEEINFNLHSAMKHSKDKGACFKVDAMDEVIMETRKQLHKPTPLERVLT 662
+ +R+ D FN+ M + F VD MDE++ E +L+ PL+ VLT
Sbjct: 277 RKSKINLRIGDIVQEFNMERIMSKPTTECQTFWVDIMDELVNELLAELNTEDPLQTVLT 335
>gb|AAF24531.1| F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 156 bits (394), Expect = 2e-36
Identities = 100/313 (31%), Positives = 169/313 (53%), Gaps = 12/313 (3%)
Query: 401 KKKEVGKNVPAHK--LPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKF 458
K KE P +K LP+P K +K+ F K +E+ IP +AL +P KF
Sbjct: 520 KNKEKVFVPPPYKSQLPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIPDSHKF 579
Query: 459 MKEILSKKHRYKEDETIQLDANCSAIIQRRL-PKKEKDPGRVTLPVAIGKVNVGKALIDL 517
+K+++ ++ + + + L CSAIIQ+++ PKK DPG TLP ++G + + L DL
Sbjct: 580 LKDLIVERIQEVQGMVV-LSHECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDL 638
Query: 518 GSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVM 577
G+S++L+P SV KR+G K ++L LAD+SV P G+ E++ +++ P DFVV+
Sbjct: 639 GASVSLMPLSVAKRLGFTQYKSCNISLILADRSVRIPHGLLENLPIRIGAVEIPTDFVVL 698
Query: 578 DIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRL-DDEEINFNLHSAMKHSKDKGACFK 636
+++E+ PLILGR F+ A MID G + + L D + F++ AMK +G F
Sbjct: 699 EMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLNLGKDFRMTFDVKDAMKKPTIEGQLFW 758
Query: 637 VDAMDEVIMETRKQLHKPTPLERVLTDA-------LSVLSAEEEKEIEECLKELETLKEI 689
++ MD++ E ++L + L LT + L L ++ + + ++E E +E+
Sbjct: 759 IEEMDQLADELLEELAEEDHLNSALTKSGEDGFLHLETLGYQKLLDSHKAMEESEPFEEL 818
Query: 690 PPKKAKVEELKKE 702
+V + +E
Sbjct: 819 NGPATEVMVMSEE 831
Score = 112 bits (280), Expect = 4e-23
Identities = 85/324 (26%), Positives = 137/324 (42%), Gaps = 30/324 (9%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
E++ + + F GL EDP HL F L N + E+ LRLF SL KA
Sbjct: 23 EIRKRRTTVNPGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKA 82
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
W + D + W+ +K F +FF + + + I+ F+Q ES C+AWER+K
Sbjct: 83 HIWEKNLPHDSITTWDDCKKVFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGY 142
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+CP HGF A+ + G+ P +++LD A++G+ E +++ ++ +D
Sbjct: 143 TNQCPHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSDGNY 202
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEE-MMKQMKEMPKLVKEQIQKEQRHQQ 277
+ T R + S+D + + K L +++ ++ Q K + LV ++ + Q +
Sbjct: 203 NEDCDRTVRGTA----DSDDKHRKEIKALNDKLDRILLSQHKHVHFLVDDEQYEVQDGEG 258
Query: 278 VHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNN 337
EEV+Y+ N QG Y G N + N N R N
Sbjct: 259 NQL------------------EEVSYINNN------QGGYKG-YNYFKTNNPNLSYRSTN 293
Query: 338 FRQPWKSNAGPSNQQPFNNNFQQY 361
P P QQ N F Y
Sbjct: 294 VANPQDQVYPPQQQQSQNKPFVPY 317
>gb|AAX95991.1| Transposable element protein, putative [Oryza sativa (japonica
cultivar-group)] gi|62733832|gb|AAX95941.1| Transposable
element protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 477
Score = 155 bits (392), Expect = 4e-36
Identities = 134/482 (27%), Positives = 215/482 (43%), Gaps = 54/482 (11%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
+LKS I + P+ G +ED +L F + N+ E +A LRLF SL+ A
Sbjct: 38 DLKSSLITMAQASPYCGKPNEDANAYLQQFLEIYNTYTMKEVSPDAIRLRLFPVSLLWNA 97
Query: 99 REWYLDQSPDVLKDWNALEK---AFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERY 155
++W+ + N +K AF FFP K + I+ F Q DES+ AWER
Sbjct: 98 KQWFYANCATI----NTCDKCSTAFLLMFFPRGKSNALRGRISSFQQTTDESIPKAWERL 153
Query: 156 KSLLRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLND 215
+ + CP HG DD + F G P + L+ A G+ + + REA+D+I KM N
Sbjct: 154 QEYVAACPHHGMDDWLILQNFYIGFTPMSRDHLNMVAGGAFFSKTVREAIDLIEKMVSNM 213
Query: 216 RAAKHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRH 275
++ QR G+ + + ++L +++ +MK++ + K + ++ H
Sbjct: 214 GWSEERLQTRQR--GV-------HTIKEMEILAAKLDFLMKRLDDHDKRPQGTVKALDSH 264
Query: 276 QQVHFCEM-CSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQR 334
CE+ S HP C P+ EE Y+GN + QG G Q P +Q
Sbjct: 265 VT---CEVRGSTGHPRNDC-PETHEEAMYMGNNNNGYRPQG---GQGWNQPCP---YYQG 314
Query: 335 GNNFRQPWK---SNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRI 391
FR+ K + G S + P YP L + +T N
Sbjct: 315 ARIFRRNVKVIITRGGKSTRDP------PYP-----------NLTRTSSITREAPSGNTT 357
Query: 392 HKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQ 451
+E EK +E LP+P+ K +++ F F+++ +++ IN+P +A+ Q
Sbjct: 358 DEEVQQEKTMPQEY---YDTRLLPFPERSRKPSVDEQFTHFVEVIQKIHINVPLLDAM-Q 413
Query: 452 MPTYAKFMKEILSKKHRYKEDETIQLDANCS-AIIQRRLPKKEKDPGRVTLPVAIGKVNV 510
+PTYA ++++IL+ K + ++L CS AI+Q+ L K KDPG P G N
Sbjct: 414 VPTYACYLEDILNNKRSLPTTKVVKLMEQCSNAILQKLLEK--KDPGVPRSPAQSGHSNS 471
Query: 511 GK 512
K
Sbjct: 472 TK 473
>pir||D84513 probable retroelement pol polyprotein [imported] - Arabidopsis
thaliana
Length = 841
Score = 149 bits (375), Expect = 4e-34
Identities = 91/273 (33%), Positives = 141/273 (51%), Gaps = 37/273 (13%)
Query: 390 RIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEAL 449
++ K++ EK KK + VP KLP+P Q K EK + F +I ++L+
Sbjct: 2 KLRKQTLEEKAKKTVLPPYVP--KLPFPGRQRKIQREKEYSLFDEIMRQLQ--------- 50
Query: 450 EQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVN 509
RY + CSAI+Q +P K +DPG LP IG+
Sbjct: 51 ------------------RYSPE--------CSAILQNVIPVKREDPGSFVLPSRIGEYT 84
Query: 510 VGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFV 569
+ L DLG+ ++L+P+SV KR+G + T+M+L L D+S++ P+G+AEDV V+V F
Sbjct: 85 FDRCLCDLGAGVSLMPFSVAKRLGDTNFTPTKMSLVLGDRSISFPVGVAEDVQVRVGNFY 144
Query: 570 FPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEEINFNLHSAMKHSK 629
P DFV+++++E+ LILGR F+ +ID + +R+ D FN+ M
Sbjct: 145 IPTDFVIIELDEEPRHRLILGRPFLNIVAALIDVRKSKINLRIGDIVQEFNMERIMSKPT 204
Query: 630 DKGACFKVDAMDEVIMETRKQLHKPTPLERVLT 662
+ F VD MDE++ E +L+ PL+ VLT
Sbjct: 205 TECQTFWVDIMDELVNELLAELNTEDPLQTVLT 237
>pir||T12084 hypothetical protein - fava bean (fragment)
gi|2522227|dbj|BAA22786.1| retrotransposon-like gene~the
first amino acid was determined to be glycine [Vicia
faba]
Length = 298
Score = 144 bits (363), Expect = 1e-32
Identities = 81/197 (41%), Positives = 124/197 (62%), Gaps = 3/197 (1%)
Query: 497 GRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMG 556
G VT+P I KALI+LG+S++L+P S+ K++G +++ TRMTLQ AD SV RP G
Sbjct: 1 GSVTIPCTIRDRFFKKALINLGASVSLMPLSIYKKLGIGNVQNTRMTLQFADHSVKRPYG 60
Query: 557 IAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLDDEE 616
I E VLVK+DKFVF VDFVV+++ ED+ + LILGR F+ R +ID ++G M +++ DEE
Sbjct: 61 IVEYVLVKIDKFVFLVDFVVLEMPEDEKITLILGRPFLETERSLIDIEEGTMNLKVYDEE 120
Query: 617 INFNLHSAMKHSKDKGACFKVDAMDEVIMETRKQLHKPTPLERVLTDALSVLSAEEEKEI 676
+ ++ + MK+ D ++ +D+V+ + PLE+VL LS+ EE +
Sbjct: 121 LKIDVRNTMKYKDDIATSQHIEVIDQVVPNENYLKTQQLPLEKVL--RLSIFEESEEVD- 177
Query: 677 EECLKELETLKEIPPKK 693
E+ ++ L + IP K
Sbjct: 178 EKYMEALAMMDAIPSFK 194
>gb|AAN04521.1| Hypothetical protein with similarity to putative retroelements
[Oryza sativa (japonica cultivar-group)]
gi|19881695|gb|AAM01096.1| Hypothetical protein with
similarity to putative retroelement [Oryza sativa]
Length = 237
Score = 143 bits (360), Expect = 2e-32
Identities = 75/202 (37%), Positives = 123/202 (60%), Gaps = 1/202 (0%)
Query: 414 LPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDE 473
LP+P+ K +++ F F+++ +++ I++P +A+ Q+PTYA ++K+IL+ K E
Sbjct: 33 LPFPQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLPTTE 91
Query: 474 TIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVG 533
++L CS +I +LP+K+KDPG T+ +IG +AL DLG+S++++P V ++
Sbjct: 92 VVKLTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVFDKLN 151
Query: 534 GLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTF 593
+ T M LQLAD SV P GIAEDV VK+ F VDFVV+D++ ++ +ILGR F
Sbjct: 152 FTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIILGRPF 211
Query: 594 MLAARMMIDYDDGLMKVRLDDE 615
+ A I G ++ + E
Sbjct: 212 LGTAGANIHVGTGSIRFQYQRE 233
>gb|AAP53442.1| hypothetical protein similar to putative retroelements [Oryza
sativa (japonica cultivar-group)]
gi|37533706|ref|NP_921155.1| hypothetical protein
similar to putative retroelements [Oryza sativa
(japonica cultivar-group)]
Length = 725
Score = 141 bits (356), Expect = 6e-32
Identities = 71/181 (39%), Positives = 116/181 (63%), Gaps = 1/181 (0%)
Query: 414 LPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDE 473
LP+P+ K +++ F F+++ +++ I++P +A+ Q+PTYA ++K+IL+ K E
Sbjct: 64 LPFPQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLPTTE 122
Query: 474 TIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVG 533
++L CS +I +LP+K+KDPG T+ +IG +AL DLG+S++++P V ++
Sbjct: 123 VVKLTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVFDKLN 182
Query: 534 GLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILGRTF 593
+ T M LQLAD SV P GIAEDV VK+ F VDFVV+D++ ++ +ILGR F
Sbjct: 183 FTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIILGRPF 242
Query: 594 M 594
+
Sbjct: 243 L 243
>gb|AAF19229.1| Similar to Athila ORF 1 [Arabidopsis thaliana]
gi|25518186|pir||B86490 F28L22.6 protein - Arabidopsis
thaliana
Length = 719
Score = 141 bits (355), Expect = 8e-32
Identities = 95/274 (34%), Positives = 156/274 (56%), Gaps = 10/274 (3%)
Query: 397 VEKDKKKEVGKNV---PAHKLPYP-KAQSKKDLEKHFKRFLDI-FKRLEINIPFSEALEQ 451
VEK + +NV P +K P P + KK + + +K L+ K LE+ +P + L
Sbjct: 424 VEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLEVTMPLVDCLAL 483
Query: 452 MPTYAKFMKEILSKKHRYKEDE-TIQLDANCSAIIQRRL-PKKEKDPGRVTLPVAIGKVN 509
+P K++K+++++ R KE + + L CSAIIQ+++ PKK DPG TLP A+G +
Sbjct: 484 IPDSNKYVKDMITE--RIKEVQGMVVLSHECSAIIQQKIIPKKLGDPGSFTLPCALGPLA 541
Query: 510 VGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFV 569
K L DLG+S++L+P V K++G K ++L LAD+SV G+ ED+ V +
Sbjct: 542 FSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVRISHGLLEDLPVMIGVVE 601
Query: 570 FPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLD-DEEINFNLHSAMKHS 628
P DFVV++++E+ PLILGR F+ AR +ID G + + L D ++ F++ + MK
Sbjct: 602 VPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNLGRDLKMTFDITNTMKKP 661
Query: 629 KDKGACFKVDAMDEVIMETRKQLHKPTPLERVLT 662
+G F ++ MD + + ++L + L+ LT
Sbjct: 662 TIEGNIFWIEEMDMLADKMLEELGETDHLQSALT 695
>gb|AAF79809.1| T32E20.9 [Arabidopsis thaliana]
Length = 1586
Score = 139 bits (349), Expect = 4e-31
Identities = 99/299 (33%), Positives = 163/299 (54%), Gaps = 14/299 (4%)
Query: 397 VEKDKKKEVGKNV---PAHKLPYP-KAQSKKDLEKHFKRFLDI-FKRLEINIPFSEALEQ 451
VEK + +NV P +K P P + KK + + +K L+ K LE+ +P + L
Sbjct: 527 VEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLEVTMPLVDCLAL 586
Query: 452 MPTYAKFMKEILSKKHRYKEDE-TIQLDANCSAIIQRRL-PKKEKDPGRVTLPVAIGKVN 509
+P K++K+++++ R KE + + L CSAIIQ+++ PKK DPG TLP A+G +
Sbjct: 587 IPDSNKYVKDMITE--RIKEVQGMVVLSHECSAIIQQKIIPKKLGDPGSFTLPCALGPLA 644
Query: 510 VGKALIDLGSSINLIPYSVVKRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFV 569
K L DLG+S++L+P V K++G K ++L LAD+SV G+ ED+ V +
Sbjct: 645 FSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVRISHGLLEDLPVMIGVVE 704
Query: 570 FPVDFVVMDIEEDDDVPLILGRTFMLAARMMIDYDDGLMKVRLD-DEEINFNLHSAMKHS 628
P DFVV++++E+ PLILGR F+ AR +ID G + + L D ++ F++ + MK
Sbjct: 705 VPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNLGRDLKMTFDITNTMKKP 764
Query: 629 KDKGACFKVDAMD----EVIMETRKQLHKPTPLERVLTDALSVLSAEEEKEIEECLKEL 683
+G F ++ MD ++ K + R+ + S S E ++ + LKE+
Sbjct: 765 TIEGNIFWIEEMDMKAIGYSLDDIKGISPTLCTHRIHLENESYSSIEPQRRLNPNLKEV 823
Score = 95.9 bits (237), Expect = 4e-18
Identities = 84/347 (24%), Positives = 141/347 (40%), Gaps = 59/347 (17%)
Query: 39 ELKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKA 98
E+KSG I +V ++ F GL EDP HL F L + + E+ LRLF SL KA
Sbjct: 72 EIKSGLIAMVQSNKFHGLPMEDPLDHLDEFDRLCSLTKINRVSEDGFKLRLFPFSLGDKA 131
Query: 99 REWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSL 158
+W + WN +KAF +FF + + + I+ F Q +E+ +AWER+K
Sbjct: 132 HQWEKSLPQGSITSWNDCKKAFLAKFFSNSRTARLRNDISGFTQTNNETFYEAWERFKGY 191
Query: 159 LRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAA 218
+CP H +++LD A++G+ + + +++ ++ +D
Sbjct: 192 QTQCPHH-------------------EMLLDTASNGNFLNKDVEDGWEVVENLAQSDGNY 232
Query: 219 KHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQV 278
+ + R S + E+ ++MK M + + + +Q+H +
Sbjct: 233 NEDYDRSIRTS------------------SDSDEKHRREMKAMNDKLDKLLLMQQKH--I 272
Query: 279 HFCEMCSGDHPTGYCPP----QLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQR 334
HF GD T QL EEV+YV NQ + + N + + N R
Sbjct: 273 HFL----GDDETLQVQDGETLQL-EEVSYVQNQGGYNKGFNNFKQNHPNLSYRSTNVANR 327
Query: 335 GNNFRQ-----------PWKSNAGPSNQQPFNNNFQQYPPQQDRTQK 370
+ P+ G +Q + N+QQ P TQ+
Sbjct: 328 QDQVYPSQQQNQPKPFVPYNQGQGYVPKQQYQGNYQQQLPPPGFTQQ 374
>gb|AAQ56360.1| hypothetical protein OSJNBa0017M13.8 [Oryza sativa (japonica
cultivar-group)]
Length = 731
Score = 134 bits (337), Expect = 1e-29
Identities = 71/177 (40%), Positives = 111/177 (62%), Gaps = 2/177 (1%)
Query: 414 LPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDE 473
LP+ + K +++ F RF+++ +++ IN+P + + QMPTYA+++K+ L+ K R E
Sbjct: 164 LPFSERSRKPSVDEQFARFVEVIQKIHINVPLLDTI-QMPTYARYLKDKLNNK-RPLPTE 221
Query: 474 TIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSVVKRVG 533
++L CS II +LPKK++DPG T+ +IG +AL DLG++++++P V ++
Sbjct: 222 VVKLTEQCSNIILHKLPKKKEDPGCPTITCSIGAQQFDQALCDLGANVSVMPKDVFDKLN 281
Query: 534 GLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLILG 590
T M LQLA+ SV P GIAEDV VK+ F PV FVV+D + + PLILG
Sbjct: 282 FTVWAPTPMHLQLANSSVRYPAGIAEDVPVKIRDFFIPVGFVVLDTDTGKETPLILG 338
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,223,065,861
Number of Sequences: 2540612
Number of extensions: 55192042
Number of successful extensions: 289842
Number of sequences better than 10.0: 1280
Number of HSP's better than 10.0 without gapping: 284
Number of HSP's successfully gapped in prelim test: 1093
Number of HSP's that attempted gapping in prelim test: 262236
Number of HSP's gapped (non-prelim): 9419
length of query: 704
length of database: 863,360,394
effective HSP length: 135
effective length of query: 569
effective length of database: 520,377,774
effective search space: 296094953406
effective search space used: 296094953406
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 79 (35.0 bits)
Medicago: description of AC145221.2


