

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145219.5 + phase: 0
(130 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
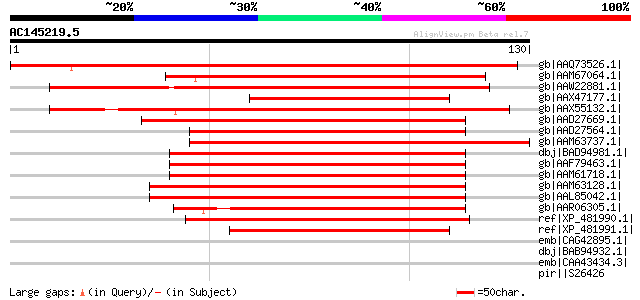
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ73526.1| early flowering 4 [Mesembryanthemum crystallinum] 122 1e-27
gb|AAM67064.1| unknown [Arabidopsis thaliana] gi|17065630|gb|AAL... 113 1e-24
gb|AAW22881.1| putative EARLY flowering 4 protein [Lycopersicon ... 111 4e-24
gb|AAX47177.1| EARLY FLOWERING 4 [Pisum sativum] 97 1e-19
gb|AAX55132.1| hypothetical protein At2g29950 [Arabidopsis thali... 94 7e-19
gb|AAD27669.1| hypothetical protein [Oryza sativa] 86 2e-16
gb|AAD27564.1| hypothetical protein [Sorghum bicolor] 84 6e-16
gb|AAM63737.1| unknown [Arabidopsis thaliana] gi|20197919|gb|AAM... 84 8e-16
dbj|BAD94981.1| hypothetical protein [Arabidopsis thaliana] gi|1... 81 5e-15
gb|AAF79463.1| F1L3.15 [Arabidopsis thaliana] gi|9665116|gb|AAF9... 81 5e-15
gb|AAM61718.1| unknown [Arabidopsis thaliana] 79 3e-14
gb|AAM63128.1| unknown [Arabidopsis thaliana] gi|12323759|gb|AAG... 77 1e-13
gb|AAL85042.1| unknown protein [Arabidopsis thaliana] gi|1453258... 77 1e-13
gb|AAR06305.1| expressed protein [Oryza sativa (japonica cultiva... 75 5e-13
ref|XP_481990.1| unknown protein [Oryza sativa (japonica cultiva... 67 1e-10
ref|XP_481991.1| hypothetical protein [Oryza sativa (japonica cu... 53 1e-06
emb|CAG42895.1| putative cell division protein [Staphylococcus a... 37 0.081
dbj|BAB94932.1| div1b [Staphylococcus aureus subsp. aureus MW2] ... 37 0.081
emb|CAA43434.3| endonuclease [Escherichia coli] gi|52001476|sp|P... 37 0.11
pir||S26426 type IV site-specific deoxyribonuclease (EC 3.1.21.-... 37 0.11
>gb|AAQ73526.1| early flowering 4 [Mesembryanthemum crystallinum]
Length = 139
Score = 122 bits (307), Expect = 1e-27
Identities = 65/128 (50%), Positives = 82/128 (63%), Gaps = 1/128 (0%)
Query: 1 MSTLYSPSSAMEDT-NHRSKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQ 59
M + S+MED NH + RR + NN+ + E GDP W +SF +
Sbjct: 1 MMSYSGAESSMEDGGNHHRQTLAGNRRRRSNNNNNNNNGEEYFSGGDPAEWNAFAESFSK 60
Query: 60 VQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICH 119
VQSVLDRNR +IQQ NEN QSR+PDNMVKNV +IQELNGNISKVAS+YSDL+ +FT +
Sbjct: 61 VQSVLDRNRMLIQQANENHQSRVPDNMVKNVAIIQELNGNISKVASIYSDLSVNFTGVVG 120
Query: 120 QQQRSSHN 127
Q ++ N
Sbjct: 121 QHRQQGGN 128
>gb|AAM67064.1| unknown [Arabidopsis thaliana] gi|17065630|gb|AAL33809.1| unknown
protein [Arabidopsis thaliana]
gi|14334558|gb|AAK59687.1| unknown protein [Arabidopsis
thaliana] gi|2088659|gb|AAB95293.1| expressed protein
[Arabidopsis thaliana] gi|25408665|pir||A84825
hypothetical protein At2g40080 [imported] - Arabidopsis
thaliana gi|18405258|ref|NP_565922.1| expressed protein
[Arabidopsis thaliana]
Length = 111
Score = 113 bits (282), Expect = 1e-24
Identities = 55/81 (67%), Positives = 68/81 (83%), Gaps = 1/81 (1%)
Query: 40 EDAEDG-DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNG 98
E+AE G DP W+ L+++FRQVQSVLDRNR++IQQVN+N QSRM DNM KNV LIQELNG
Sbjct: 15 EEAEQGEDPAMWENLDRNFRQVQSVLDRNRSLIQQVNDNHQSRMADNMSKNVALIQELNG 74
Query: 99 NISKVASLYSDLNSDFTNICH 119
NISKV ++YSDLN+ F++ H
Sbjct: 75 NISKVVNMYSDLNTSFSSGFH 95
>gb|AAW22881.1| putative EARLY flowering 4 protein [Lycopersicon esculentum]
Length = 121
Score = 111 bits (277), Expect = 4e-24
Identities = 57/110 (51%), Positives = 75/110 (67%), Gaps = 1/110 (0%)
Query: 11 MEDTNHRSKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAI 70
MEDT R ++ + + T D+ E+G+ + W + FRQVQSVLDRNR++
Sbjct: 1 MEDTFKRHRQTLAKPQSLTTDDRRRPRDLS-TEEGNSDVWNNFSNRFRQVQSVLDRNRSL 59
Query: 71 IQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICHQ 120
IQQVNEN QSR DNMV+NV LIQELNGNISKV SLYSD++++F+ + Q
Sbjct: 60 IQQVNENHQSRTTDNMVRNVSLIQELNGNISKVVSLYSDISTNFSTMFRQ 109
>gb|AAX47177.1| EARLY FLOWERING 4 [Pisum sativum]
Length = 50
Score = 96.7 bits (239), Expect = 1e-19
Identities = 49/50 (98%), Positives = 49/50 (98%)
Query: 61 QSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDL 110
QSVLDRNRAIIQQVNENQQSRMPDNMVKNV LIQELNGNISKVASLYSDL
Sbjct: 1 QSVLDRNRAIIQQVNENQQSRMPDNMVKNVSLIQELNGNISKVASLYSDL 50
>gb|AAX55132.1| hypothetical protein At2g29950 [Arabidopsis thaliana]
gi|3420056|gb|AAC31857.1| hypothetical protein
[Arabidopsis thaliana] gi|49823506|gb|AAT68736.1|
hypothetical protein At2g29950 [Arabidopsis thaliana]
gi|15227672|ref|NP_180556.1| expressed protein
[Arabidopsis thaliana] gi|7459222|pir||T02490
hypothetical protein At2g29950 [imported] - Arabidopsis
thaliana
Length = 125
Score = 94.0 bits (232), Expect = 7e-19
Identities = 52/116 (44%), Positives = 72/116 (61%), Gaps = 4/116 (3%)
Query: 11 MEDTNHRSKKNQRRRRPSATPTNNDYSDVE-DAEDGDPEAWQTLNKSFRQVQSVLDRNRA 69
ME + +RS R S ND DV A D E W TL+ F++ Q LD+NR
Sbjct: 1 MEASRNRSLVGNNR---SPEMNENDGEDVAASAAVEDVEVWDTLSNGFKRAQLYLDQNRD 57
Query: 70 IIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICHQQQRSS 125
+IQ+VNEN SR+PDN+ +NVGLI E+NGNIS+V +YSDL+ +F Q++R++
Sbjct: 58 LIQRVNENHMSRIPDNVSRNVGLINEINGNISQVMEIYSDLSLNFAKKFDQRRRTT 113
>gb|AAD27669.1| hypothetical protein [Oryza sativa]
Length = 117
Score = 86.3 bits (212), Expect = 2e-16
Identities = 39/81 (48%), Positives = 59/81 (72%)
Query: 34 NDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLI 93
+ +S + + D + QT +KSF QVQS+LD+NR +I ++N+N +SR PDN+ +NVGLI
Sbjct: 4 DSFSGMANGGQVDNKLIQTFHKSFVQVQSILDQNRMLINEINQNHESRAPDNLTRNVGLI 63
Query: 94 QELNGNISKVASLYSDLNSDF 114
+ELN NI +V LY+DL++ F
Sbjct: 64 RELNNNIRRVVGLYADLSASF 84
>gb|AAD27564.1| hypothetical protein [Sorghum bicolor]
Length = 438
Score = 84.3 bits (207), Expect = 6e-16
Identities = 38/69 (55%), Positives = 54/69 (78%)
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
D + QT +KSF QVQS+LD+NR +I ++N+N +SR PDN+ +NVGLI+ELN NI +V
Sbjct: 10 DGKLIQTFHKSFVQVQSLLDQNRMLISEINQNHESRAPDNLTRNVGLIRELNNNIRRVVG 69
Query: 106 LYSDLNSDF 114
LY+DL++ F
Sbjct: 70 LYADLSASF 78
>gb|AAM63737.1| unknown [Arabidopsis thaliana] gi|20197919|gb|AAM15312.1| Expressed
protein [Arabidopsis thaliana]
gi|18396289|ref|NP_565334.1| expressed protein
[Arabidopsis thaliana]
Length = 109
Score = 84.0 bits (206), Expect = 8e-16
Identities = 37/85 (43%), Positives = 58/85 (67%)
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
D + QT KSF QVQ++LD NR +I ++N+N +S++PDN+ +NVGLI+ELN N+ +VA
Sbjct: 17 DGKILQTFEKSFVQVQNILDHNRLLINEINQNHESKIPDNLGRNVGLIRELNNNVRRVAH 76
Query: 106 LYSDLNSDFTNICHQQQRSSHNRGK 130
LY DL+++F+ + G+
Sbjct: 77 LYVDLSNNFSKSMEASSEGDSSEGR 101
>dbj|BAD94981.1| hypothetical protein [Arabidopsis thaliana]
gi|18394486|ref|NP_564024.1| expressed protein
[Arabidopsis thaliana]
Length = 114
Score = 81.3 bits (199), Expect = 5e-15
Identities = 37/74 (50%), Positives = 53/74 (71%)
Query: 41 DAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
D + D + Q+ KSF VQ +LD+NR +I ++N+N +S+ PDN+ +NVGLI+ELN NI
Sbjct: 11 DRHNMDGKLLQSFQKSFVDVQDILDQNRLLINEINQNHESKQPDNLGRNVGLIKELNNNI 70
Query: 101 SKVASLYSDLNSDF 114
+VASLY DL+ F
Sbjct: 71 RRVASLYGDLSHSF 84
>gb|AAF79463.1| F1L3.15 [Arabidopsis thaliana] gi|9665116|gb|AAF97300.1|
Hypothetical protein [Arabidopsis thaliana]
Length = 141
Score = 81.3 bits (199), Expect = 5e-15
Identities = 37/74 (50%), Positives = 53/74 (71%)
Query: 41 DAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
D + D + Q+ KSF VQ +LD+NR +I ++N+N +S+ PDN+ +NVGLI+ELN NI
Sbjct: 11 DRHNMDGKLLQSFQKSFVDVQDILDQNRLLINEINQNHESKQPDNLGRNVGLIKELNNNI 70
Query: 101 SKVASLYSDLNSDF 114
+VASLY DL+ F
Sbjct: 71 RRVASLYGDLSHSF 84
>gb|AAM61718.1| unknown [Arabidopsis thaliana]
Length = 114
Score = 78.6 bits (192), Expect = 3e-14
Identities = 36/74 (48%), Positives = 52/74 (69%)
Query: 41 DAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
D + D + Q+ KSF VQ +LD+NR +I ++N+ +S+ PDN+ +NVGLI+ELN NI
Sbjct: 11 DRHNMDGKLLQSFQKSFVDVQDILDQNRLLINEINQXHESKQPDNLGRNVGLIKELNNNI 70
Query: 101 SKVASLYSDLNSDF 114
+VASLY DL+ F
Sbjct: 71 RRVASLYGDLSHSF 84
>gb|AAM63128.1| unknown [Arabidopsis thaliana] gi|12323759|gb|AAG51839.1|
hypothetical protein; 65517-65170 [Arabidopsis thaliana]
gi|25406197|pir||H96750 hypothetical protein F28P22.18
[imported] - Arabidopsis thaliana
Length = 115
Score = 76.6 bits (187), Expect = 1e-13
Identities = 37/79 (46%), Positives = 52/79 (64%)
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
YS + D + Q KSF QVQ +LD+NR +I ++N+N +S+ D++ +NVGLI+E
Sbjct: 6 YSGFGERYQMDGKLLQNFQKSFVQVQDILDQNRLLINEINQNHESKQADHLGRNVGLIRE 65
Query: 96 LNGNISKVASLYSDLNSDF 114
LN NI VASLY DL+ F
Sbjct: 66 LNNNIRTVASLYGDLSHSF 84
>gb|AAL85042.1| unknown protein [Arabidopsis thaliana] gi|14532584|gb|AAK64020.1|
unknown protein [Arabidopsis thaliana]
gi|18410139|ref|NP_565044.1| expressed protein
[Arabidopsis thaliana]
Length = 119
Score = 76.6 bits (187), Expect = 1e-13
Identities = 37/79 (46%), Positives = 52/79 (64%)
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
YS + D + Q KSF QVQ +LD+NR +I ++N+N +S+ D++ +NVGLI+E
Sbjct: 10 YSGFGERYQMDGKLLQNFQKSFVQVQDILDQNRLLINEINQNHESKQADHLGRNVGLIRE 69
Query: 96 LNGNISKVASLYSDLNSDF 114
LN NI VASLY DL+ F
Sbjct: 70 LNNNIRTVASLYGDLSHSF 88
>gb|AAR06305.1| expressed protein [Oryza sativa (japonica cultivar-group)]
gi|50916469|ref|XP_468631.1| expressed protein [Oryza
sativa (japonica cultivar-group)]
Length = 114
Score = 74.7 bits (182), Expect = 5e-13
Identities = 39/74 (52%), Positives = 56/74 (74%), Gaps = 4/74 (5%)
Query: 42 AEDGDP-EAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNI 100
AEDG A+QT SF QVQS+LD+NR +I ++N+N +S++P ++ +NVGLI+ELN NI
Sbjct: 11 AEDGKVLHAFQT---SFVQVQSLLDQNRVLINEINQNHESKVPGDLSRNVGLIRELNNNI 67
Query: 101 SKVASLYSDLNSDF 114
+V LY+DL+S F
Sbjct: 68 RRVVDLYADLSSLF 81
>ref|XP_481990.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|38637104|dbj|BAD03359.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 144
Score = 67.0 bits (162), Expect = 1e-10
Identities = 28/71 (39%), Positives = 51/71 (71%)
Query: 45 GDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVA 104
G + Q L ++F +VQ +L++NR +IQ++++N ++R D + +NV LI+ELN NI++V
Sbjct: 36 GGGKVVQVLQRNFGEVQGILEQNRVLIQEISQNHEARDADGLTRNVALIRELNTNIARVV 95
Query: 105 SLYSDLNSDFT 115
LY++L+ F+
Sbjct: 96 DLYANLSGSFS 106
>ref|XP_481991.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|38637105|dbj|BAD03360.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 357
Score = 53.1 bits (126), Expect = 1e-06
Identities = 21/55 (38%), Positives = 41/55 (74%)
Query: 56 SFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDL 110
S +V+ +L++N +IQ++++N ++R D + +NV LI++LN NI++V LY++L
Sbjct: 256 SLGEVRGILEQNHTLIQEISQNHKARDADRLTRNVALIRDLNTNIARVVDLYANL 310
>emb|CAG42895.1| putative cell division protein [Staphylococcus aureus subsp.
aureus MSSA476] gi|49486024|ref|YP_043245.1| putative
cell division protein [Staphylococcus aureus subsp.
aureus MSSA476]
Length = 440
Score = 37.4 bits (85), Expect = 0.081
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 2/77 (2%)
Query: 14 TNHRSKKNQRRRRPSATP-TNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQ 72
T + SKK ++R+R AT +N + D D D E + +NK F++ QS + N +
Sbjct: 24 TRNTSKKRRQRKRSKATHFSNQNKDDTSQQADFDEEIY-LINKDFKKEQSNDENNDSASS 82
Query: 73 QVNENQQSRMPDNMVKN 89
+ N N D+ ++N
Sbjct: 83 RANNNNIDDSTDSNIEN 99
>dbj|BAB94932.1| div1b [Staphylococcus aureus subsp. aureus MW2]
gi|21282796|ref|NP_645884.1| hypothetical protein
MW1067 [Staphylococcus aureus subsp. aureus MW2]
Length = 439
Score = 37.4 bits (85), Expect = 0.081
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 2/77 (2%)
Query: 14 TNHRSKKNQRRRRPSATP-TNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQ 72
T + SKK ++R+R AT +N + D D D E + +NK F++ QS + N +
Sbjct: 23 TRNTSKKRRQRKRSKATHFSNQNKDDTSQQADFDEEIY-LINKDFKKEQSNDENNDSASS 81
Query: 73 QVNENQQSRMPDNMVKN 89
+ N N D+ ++N
Sbjct: 82 RANNNNIDDSTDSNIEN 98
>emb|CAA43434.3| endonuclease [Escherichia coli] gi|52001476|sp|P25239|T257_ECOLI
Type IIS restriction enzyme Eco57I (Endonuclease Eco57I)
[Includes: Adenine-specific methyltransferase activity
Eco57IA (M.Eco57IA)]
Length = 998
Score = 37.0 bits (84), Expect = 0.11
Identities = 24/78 (30%), Positives = 41/78 (51%), Gaps = 3/78 (3%)
Query: 35 DYSDVEDAEDGDP--EAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGL 92
D+ D + E D Q LNK + Q Q DR++ I ++ E ++ +M D ++KN+
Sbjct: 922 DFDDPKQKEIHDTISSKQQYLNKLYSQTQKSADRDKIIFERQFEQEKIQM-DYLIKNLFD 980
Query: 93 IQELNGNISKVASLYSDL 110
+ +L+ I V LY +L
Sbjct: 981 LGDLDSEIPTVEDLYKNL 998
>pir||S26426 type IV site-specific deoxyribonuclease (EC 3.1.21.-) Eco57I -
Escherichia coli gi|145139|gb|AAA23389.1| restriction
endonuclease
Length = 997
Score = 37.0 bits (84), Expect = 0.11
Identities = 24/78 (30%), Positives = 41/78 (51%), Gaps = 3/78 (3%)
Query: 35 DYSDVEDAEDGDP--EAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGL 92
D+ D + E D Q LNK + Q Q DR++ I ++ E ++ +M D ++KN+
Sbjct: 921 DFDDPKQKEIHDTISSKQQYLNKLYSQTQKSADRDKIIFERQFEQEKIQM-DYLIKNLFD 979
Query: 93 IQELNGNISKVASLYSDL 110
+ +L+ I V LY +L
Sbjct: 980 LGDLDSEIPTVEDLYKNL 997
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.309 0.123 0.340
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 223,253,657
Number of Sequences: 2540612
Number of extensions: 8618142
Number of successful extensions: 30351
Number of sequences better than 10.0: 221
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 167
Number of HSP's that attempted gapping in prelim test: 30226
Number of HSP's gapped (non-prelim): 279
length of query: 130
length of database: 863,360,394
effective HSP length: 106
effective length of query: 24
effective length of database: 594,055,522
effective search space: 14257332528
effective search space used: 14257332528
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC145219.5


