

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145218.1 - phase: 0
(63 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
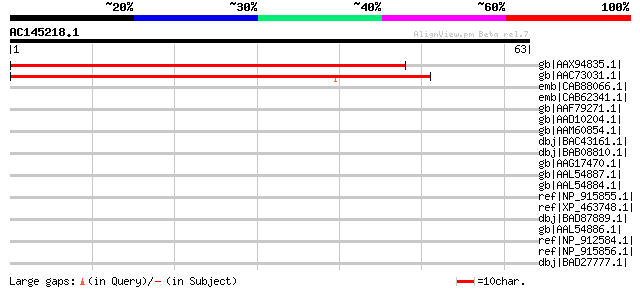
Score E
Sequences producing significant alignments: (bits) Value
gb|AAX94835.1| Cytochrome P450 [Oryza sativa (japonica cultivar-... 47 1e-04
gb|AAC73031.1| putative cytochrome P450 [Arabidopsis thaliana] g... 47 1e-04
emb|CAB88066.1| cytochrome P450-like protein [Arabidopsis thalia... 37 0.10
emb|CAB62341.1| cytochrome P450-like protein [Arabidopsis thalia... 37 0.14
gb|AAF79271.1| F12K21.15 [Arabidopsis thaliana] gi|15218671|ref|... 37 0.14
gb|AAD10204.1| CYP94A1 [Vicia sativa] gi|17366212|sp|O81117|C94A... 35 0.68
gb|AAM60854.1| cytochrome P450-like protein [Arabidopsis thaliana] 34 1.2
dbj|BAC43161.1| putative cytochrome P450 [Arabidopsis thaliana] ... 34 1.2
dbj|BAB08810.1| cytochrome P450-like protein [Arabidopsis thalia... 34 1.2
gb|AAG17470.1| cytochrome P450 [Triticum aestivum] 33 2.6
gb|AAL54887.1| cytochrome P450-dependent fatty acid hydroxylase ... 32 3.4
gb|AAL54884.1| cytochrome P450-dependent fatty acid hydroxylase ... 32 3.4
ref|NP_915855.1| cytochrome P450-like protein [Oryza sativa (jap... 32 4.4
ref|XP_463748.1| putative cytochrome P450-like protein [Oryza sa... 32 4.4
dbj|BAD87889.1| putative cytochrome P450-dependent fatty acid hy... 32 4.4
gb|AAL54886.1| cytochrome P450-dependent fatty acid hydroxylase ... 32 4.4
ref|NP_912584.1| Putative cytochrome P450 [Oryza sativa (japonic... 32 5.7
ref|NP_915856.1| cytochrome P450-like protein [Oryza sativa (jap... 31 7.5
dbj|BAD27777.1| putative cytochrome P450 [Oryza sativa (japonica... 31 7.5
>gb|AAX94835.1| Cytochrome P450 [Oryza sativa (japonica cultivar-group)]
gi|62701694|gb|AAX92767.1| Cytochrome P450 [Oryza sativa
(japonica cultivar-group)]
Length = 520
Score = 47.4 bits (111), Expect = 1e-04
Identities = 24/48 (50%), Positives = 32/48 (66%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
MASLEL SV VRSYA +II +E+ RL+P + A + DL+D+ R
Sbjct: 142 MASLELGSVAVRSYAYKIIAQEVEARLMPVLADAADRGAVLDLQDVFR 189
>gb|AAC73031.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|18087602|gb|AAL58931.1| At2g27690/F15K20.21
[Arabidopsis thaliana] gi|13877669|gb|AAK43912.1|
putative cytochrome P450 [Arabidopsis thaliana]
gi|15226228|ref|NP_180337.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|25282628|pir||G84675 probable
cytochrome P450 [imported] - Arabidopsis thaliana
Length = 495
Score = 47.0 bits (110), Expect = 1e-04
Identities = 25/52 (48%), Positives = 36/52 (69%), Gaps = 1/52 (1%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDE-KIFDLRDIMRSSS 51
+ASLEL SV+VR +A EI+ EI TRL+P + SF+ + + DL+D+ R S
Sbjct: 127 LASLELGSVSVRVFAHEIVKTEIETRLLPILTSFSDNPGSVLDLQDVFRRFS 178
>emb|CAB88066.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|15228981|ref|NP_191222.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|11249683|pir||T49064
cytochrome P450-like protein - Arabidopsis thaliana
Length = 499
Score = 37.4 bits (85), Expect = 0.10
Identities = 16/46 (34%), Positives = 30/46 (64%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++ ++R + + + EI+TRL+P + A + K+ DL+DI+
Sbjct: 129 ASYEFSTKSLRDFVMSNVTVEINTRLVPVLAEAATNGKLIDLQDIL 174
>emb|CAB62341.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|53828589|gb|AAU94404.1| At3g48520 [Arabidopsis
thaliana] gi|51536434|gb|AAU05455.1| At3g48520
[Arabidopsis thaliana] gi|15228396|ref|NP_190421.1|
cytochrome P450 family protein [Arabidopsis thaliana]
gi|11249685|pir||T46196 cytochrome P450-like protein -
Arabidopsis thaliana
Length = 506
Score = 37.0 bits (84), Expect = 0.14
Identities = 15/48 (31%), Positives = 32/48 (66%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E ++ ++RS+A E++ +E+ RL+P + + A DL+D+++
Sbjct: 133 LASHEFSTRSLRSFAFEVLKDEVENRLVPVLSTAADVGTTVDLQDVLK 180
>gb|AAF79271.1| F12K21.15 [Arabidopsis thaliana] gi|15218671|ref|NP_174713.1|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 498
Score = 37.0 bits (84), Expect = 0.14
Identities = 16/46 (34%), Positives = 29/46 (62%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++ ++R + + + EI+TRL+P + A K+ DL+DI+
Sbjct: 129 ASYEFSTKSLRDFVMSNVTVEINTRLVPVLVEAATTGKLIDLQDIL 174
>gb|AAD10204.1| CYP94A1 [Vicia sativa] gi|17366212|sp|O81117|C94A1_VICSA Cytochrome
P450 94A1 (P450-dependent fatty acid omega-hydroxylase)
Length = 514
Score = 34.7 bits (78), Expect = 0.68
Identities = 15/48 (31%), Positives = 29/48 (60%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E + ++R++ I++ E+ RLIP + S + I D +DI++
Sbjct: 145 VASHEFNTKSIRNFVEHIVDTELTNRLIPILTSSTQTNNILDFQDILQ 192
>gb|AAM60854.1| cytochrome P450-like protein [Arabidopsis thaliana]
Length = 508
Score = 33.9 bits (76), Expect = 1.2
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 133 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 180
>dbj|BAC43161.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|29028924|gb|AAO64841.1| At5g63450 [Arabidopsis
thaliana]
Length = 510
Score = 33.9 bits (76), Expect = 1.2
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 135 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 182
>dbj|BAB08810.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|15242759|ref|NP_201150.1| cytochrome P450, putative
[Arabidopsis thaliana]
Length = 510
Score = 33.9 bits (76), Expect = 1.2
Identities = 15/48 (31%), Positives = 28/48 (58%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E T ++R + EI+ EE+ RLIP + S + D +++++
Sbjct: 135 LASHEFTMRSLREFTFEILREEVQNRLIPVLSSAVDCGETVDFQEVLK 182
>gb|AAG17470.1| cytochrome P450 [Triticum aestivum]
Length = 541
Score = 32.7 bits (73), Expect = 2.6
Identities = 15/46 (32%), Positives = 28/46 (60%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
A+LE T+ T+R+ ++ IH RL+P + A+ + DL+D++
Sbjct: 134 AALEFTTRTLRTAMSRWVSRSIHGRLLPILADAAKGKAQVDLQDLL 179
>gb|AAL54887.1| cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 511
Score = 32.3 bits (72), Expect = 3.4
Identities = 13/48 (27%), Positives = 30/48 (62%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E + ++R + +++ E+ RLIP + + A ++ + D +DI++
Sbjct: 142 VASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQDILQ 189
>gb|AAL54884.1| cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 510
Score = 32.3 bits (72), Expect = 3.4
Identities = 13/48 (27%), Positives = 30/48 (62%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
+AS E + ++R + +++ E+ RLIP + + A ++ + D +DI++
Sbjct: 141 VASHEFNTRSLRKFVETVVDTELSERLIPILATAAANKTVLDFQDILQ 188
>ref|NP_915855.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 508
Score = 32.0 bits (71), Expect = 4.4
Identities = 12/46 (26%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++R++ ++ + E+ RL+P + RD + D++D++
Sbjct: 136 ASYEFNKRSLRNFVVDTVRSEVVERLLPLLERAERDGRTLDVQDVL 181
>ref|XP_463748.1| putative cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 544
Score = 32.0 bits (71), Expect = 4.4
Identities = 12/47 (25%), Positives = 28/47 (59%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
+AS +S ++R ++ ++ +H RL+P + + A + DL+D++
Sbjct: 137 LASYSFSSRSLRRFSARVLRAHLHRRLVPLLDAAAGSGEAVDLQDVL 183
>dbj|BAD87889.1| putative cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa (japonica cultivar-group)]
Length = 546
Score = 32.0 bits (71), Expect = 4.4
Identities = 12/47 (25%), Positives = 28/47 (59%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
+AS +S ++R ++ ++ +H RL+P + + A + DL+D++
Sbjct: 139 LASYSFSSRSLRRFSARVLRAHLHRRLVPLLDAAAGSGEAVDLQDVL 185
>gb|AAL54886.1| cytochrome P450-dependent fatty acid hydroxylase [Nicotiana
tabacum]
Length = 511
Score = 32.0 bits (71), Expect = 4.4
Identities = 12/48 (25%), Positives = 31/48 (64%)
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIMR 48
++S E + ++R + +++ E++ RLIP + + A ++ + D +DI++
Sbjct: 139 VSSHEFNTKSLRKFVETVVDTELNERLIPILATAAVEKTVLDFQDILQ 186
>ref|NP_912584.1| Putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
gi|22748335|gb|AAN05337.1| Putative cytochrome P450
[Oryza sativa (japonica cultivar-group)]
Length = 523
Score = 31.6 bits (70), Expect = 5.7
Identities = 13/46 (28%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E + ++R++ ++ E+H RL+P + A + + DL+D +
Sbjct: 137 ASYEFNTRSLRAFVARCVHGELHGRLLPLLRRAAAEGRAIDLQDAL 182
>ref|NP_915856.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20804564|dbj|BAB92256.1| putative
cytochrome P450-dependent fatty acid hydroxylase [Oryza
sativa (japonica cultivar-group)]
Length = 512
Score = 31.2 bits (69), Expect = 7.5
Identities = 13/46 (28%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
AS E ++RS+ ++ + E+ RL+P + RD + D++D++
Sbjct: 138 ASYEFNQRSLRSFVVDTVRFEVVERLLPLLEWARRDGRTLDVQDVL 183
>dbj|BAD27777.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
gi|50251373|dbj|BAD28400.1| putative cytochrome P450
[Oryza sativa (japonica cultivar-group)]
Length = 532
Score = 31.2 bits (69), Expect = 7.5
Identities = 15/46 (32%), Positives = 27/46 (58%)
Query: 2 ASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEKIFDLRDIM 47
A+LE T+ T+R+ ++ IH RL+P + A + DL+D++
Sbjct: 134 AALEFTTRTLRTAMSRWVSRSIHHRLLPILDDAAAGKAHVDLQDLL 179
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.135 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 76,401,158
Number of Sequences: 2540612
Number of extensions: 1957848
Number of successful extensions: 6331
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 6312
Number of HSP's gapped (non-prelim): 19
length of query: 63
length of database: 863,360,394
effective HSP length: 39
effective length of query: 24
effective length of database: 764,276,526
effective search space: 18342636624
effective search space used: 18342636624
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC145218.1


