

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145157.11 + phase: 0
(74 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
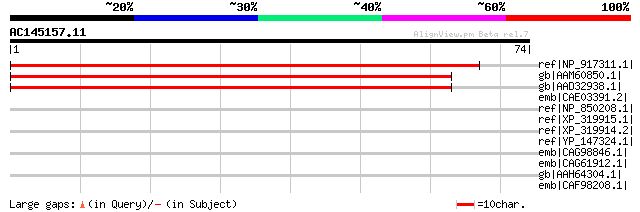
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_917311.1| OSJNBb0006H05.27 [Oryza sativa (japonica cultiv... 92 3e-18
gb|AAM60850.1| unknown [Arabidopsis thaliana] gi|26451978|dbj|BA... 90 2e-17
gb|AAD32938.1| T17H7.13 [Arabidopsis thaliana] gi|25403370|pir||... 90 2e-17
emb|CAE03391.2| OSJNBa0004N05.15 [Oryza sativa (japonica cultiva... 40 0.020
ref|NP_850208.1| actin-related protein 2/3 complex 34kDa subunit... 39 0.035
ref|XP_319915.1| ENSANGP00000024536 [Anopheles gambiae str. PEST... 32 4.2
ref|XP_319914.2| ENSANGP00000005273 [Anopheles gambiae str. PEST... 32 4.2
ref|YP_147324.1| 3-oxoacyl-[acyl-carrier-protein] reductase [Geo... 32 4.2
emb|CAG98846.1| unnamed protein product [Kluyveromyces lactis NR... 32 5.5
emb|CAG61912.1| unnamed protein product [Candida glabrata CBS138... 31 9.4
gb|AAH64304.1| Actin related protein 2/3 complex, subunit 2 [Dan... 31 9.4
emb|CAF98208.1| unnamed protein product [Tetraodon nigroviridis] 31 9.4
>ref|NP_917311.1| OSJNBb0006H05.27 [Oryza sativa (japonica cultivar-group)]
Length = 428
Score = 92.4 bits (228), Expect = 3e-18
Identities = 42/67 (62%), Positives = 55/67 (81%)
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINL 60
M+NP + +LSV+LP P E + GLP GAIEAIKAAYG +VQILDPP+DGF+LT+KINL
Sbjct: 135 MRNPKVMVLSVALPLPPPEAMLYDGLPLGAIEAIKAAYGPVVQILDPPKDGFDLTMKINL 194
Query: 61 SKVPANQ 67
+K+P ++
Sbjct: 195 TKLPPDE 201
>gb|AAM60850.1| unknown [Arabidopsis thaliana] gi|26451978|dbj|BAC43079.1| unknown
protein [Arabidopsis thaliana]
gi|28950897|gb|AAO63372.1| At1g30825 [Arabidopsis
thaliana] gi|18397679|ref|NP_564364.1| actin-related
protein 2/3 complex 34kDa subunit family / arp2/3
complex 34kDa subunit family [Arabidopsis thaliana]
Length = 318
Score = 89.7 bits (221), Expect = 2e-17
Identities = 44/63 (69%), Positives = 49/63 (76%)
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINL 60
MKNPN+ LLSVSLP P E + GLP GAIEAIK YG+ QILDPPRDGF+LTLK+N
Sbjct: 46 MKNPNLLLLSVSLPNPPPEAMSFDGLPLGAIEAIKTTYGTGFQILDPPRDGFSLTLKLNF 105
Query: 61 SKV 63
SKV
Sbjct: 106 SKV 108
>gb|AAD32938.1| T17H7.13 [Arabidopsis thaliana] gi|25403370|pir||B86434 protein
T17H7.13 [imported] - Arabidopsis thaliana
Length = 415
Score = 89.7 bits (221), Expect = 2e-17
Identities = 44/63 (69%), Positives = 49/63 (76%)
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINL 60
MKNPN+ LLSVSLP P E + GLP GAIEAIK YG+ QILDPPRDGF+LTLK+N
Sbjct: 163 MKNPNLLLLSVSLPNPPPEAMSFDGLPLGAIEAIKTTYGTGFQILDPPRDGFSLTLKLNF 222
Query: 61 SKV 63
SKV
Sbjct: 223 SKV 225
>emb|CAE03391.2| OSJNBa0004N05.15 [Oryza sativa (japonica cultivar-group)]
gi|50926408|ref|XP_473151.1| OSJNBa0004N05.15 [Oryza
sativa (japonica cultivar-group)]
Length = 368
Score = 39.7 bits (91), Expect = 0.020
Identities = 20/57 (35%), Positives = 34/57 (59%), Gaps = 1/57 (1%)
Query: 5 NIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINLS 61
NI+L S+S P+ S E GLP ++ + Y +I++P ++G+ LTL++N S
Sbjct: 50 NIYL-SISTPSLSYEASPSSGLPEITLQETRKMYHKFAEIIEPAKEGYTLTLRLNFS 105
>ref|NP_850208.1| actin-related protein 2/3 complex 34kDa subunit family / arp2/3
complex 34kDa subunit family [Arabidopsis thaliana]
Length = 365
Score = 38.9 bits (89), Expect = 0.035
Identities = 21/64 (32%), Positives = 33/64 (50%)
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINL 60
+ +PNI +S S + + + + E IK ++ I+DPPR GF LTL +NL
Sbjct: 45 VSDPNIVHVSTSTLLETQGAVTLKEISSYTYEVIKNIGVGVIDIVDPPRLGFQLTLGLNL 104
Query: 61 SKVP 64
+P
Sbjct: 105 DNIP 108
>ref|XP_319915.1| ENSANGP00000024536 [Anopheles gambiae str. PEST]
gi|30174343|gb|EAA43392.1| ENSANGP00000024536 [Anopheles
gambiae str. PEST]
Length = 296
Score = 32.0 bits (71), Expect = 4.2
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 3/38 (7%)
Query: 29 GAIEAIKAAYGSLVQILDPPRDGFNLTLKINLSKVPAN 66
GA E +K YG L L P DG+N+++ ++L +P N
Sbjct: 71 GADELLKREYGDL---LVAPEDGYNVSVLVDLENIPEN 105
>ref|XP_319914.2| ENSANGP00000005273 [Anopheles gambiae str. PEST]
gi|62899868|sp|Q7PVX8|ARPC2_ANOGA Probable actin-related
protein 2/3 complex subunit 2 (ARP2/3 complex 34 kDa
subunit) (p34-ARC) gi|55235491|gb|EAA14760.3|
ENSANGP00000005273 [Anopheles gambiae str. PEST]
Length = 304
Score = 32.0 bits (71), Expect = 4.2
Identities = 15/38 (39%), Positives = 23/38 (60%), Gaps = 3/38 (7%)
Query: 29 GAIEAIKAAYGSLVQILDPPRDGFNLTLKINLSKVPAN 66
GA E +K YG L L P DG+N+++ ++L +P N
Sbjct: 71 GADELLKREYGDL---LVAPEDGYNVSVLVDLENIPEN 105
>ref|YP_147324.1| 3-oxoacyl-[acyl-carrier-protein] reductase [Geobacillus
kaustophilus HTA426] gi|56379848|dbj|BAD75756.1|
3-oxoacyl-[acyl-carrier-protein] reductase [Geobacillus
kaustophilus HTA426]
Length = 239
Score = 32.0 bits (71), Expect = 4.2
Identities = 18/60 (30%), Positives = 30/60 (50%), Gaps = 4/60 (6%)
Query: 5 NIFLLSVSLPTPSSETIFVCGL--PFGAIEAI--KAAYGSLVQILDPPRDGFNLTLKINL 60
N+FL V + S FVC + G+++A+ A G+ +LD D F+ + +NL
Sbjct: 56 NVFLAEVDVADERSVKDFVCDVVSTLGSVDALINSAGVGTFANVLDSQTDEFDTMISVNL 115
>emb|CAG98846.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50312213|ref|XP_456138.1| unnamed protein product
[Kluyveromyces lactis]
Length = 127
Score = 31.6 bits (70), Expect = 5.5
Identities = 17/60 (28%), Positives = 29/60 (48%), Gaps = 3/60 (5%)
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAY---GSLVQILDPPRDGFNLTLK 57
MKNP + S+P PS + + +G I A++A + G L+ + D D F ++
Sbjct: 66 MKNPEVEFCGYSIPHPSENFLHIRIQTYGKITAVEALHKGLGDLMDMCDAVEDRFTQRIR 125
>emb|CAG61912.1| unnamed protein product [Candida glabrata CBS138]
gi|50293019|ref|XP_448942.1| unnamed protein product
[Candida glabrata]
Length = 137
Score = 30.8 bits (68), Expect = 9.4
Identities = 16/55 (29%), Positives = 28/55 (50%), Gaps = 3/55 (5%)
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFG---AIEAIKAAYGSLVQILDPPRDGF 52
MKNP + S+P PS E + + +G A++A++ L+ + D +D F
Sbjct: 77 MKNPEVEFCGYSIPHPSEELLNLRIQTYGNITAVDALQKGLDDLIDLCDAVKDKF 131
>gb|AAH64304.1| Actin related protein 2/3 complex, subunit 2 [Danio rerio]
gi|47086645|ref|NP_997864.1| actin related protein 2/3
complex, subunit 2 [Danio rerio]
Length = 299
Score = 30.8 bits (68), Expect = 9.4
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 3/39 (7%)
Query: 29 GAIEAIKAAYGSLVQILDPPRDGFNLTLKINLSKVPANQ 67
GA E +K YG+ L P DG+N++L +L +P N+
Sbjct: 71 GADELLKRVYGNF---LVAPEDGYNISLLFDLDALPPNK 106
>emb|CAF98208.1| unnamed protein product [Tetraodon nigroviridis]
Length = 299
Score = 30.8 bits (68), Expect = 9.4
Identities = 16/39 (41%), Positives = 23/39 (58%), Gaps = 3/39 (7%)
Query: 29 GAIEAIKAAYGSLVQILDPPRDGFNLTLKINLSKVPANQ 67
GA E +K YG+ L P G+N++L NL +PAN+
Sbjct: 71 GADELLKRVYGNF---LVPAEAGYNVSLLYNLEALPANK 106
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.141 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 129,291,442
Number of Sequences: 2540612
Number of extensions: 4165014
Number of successful extensions: 9721
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 9715
Number of HSP's gapped (non-prelim): 12
length of query: 74
length of database: 863,360,394
effective HSP length: 50
effective length of query: 24
effective length of database: 736,329,794
effective search space: 17671915056
effective search space used: 17671915056
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC145157.11


