

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
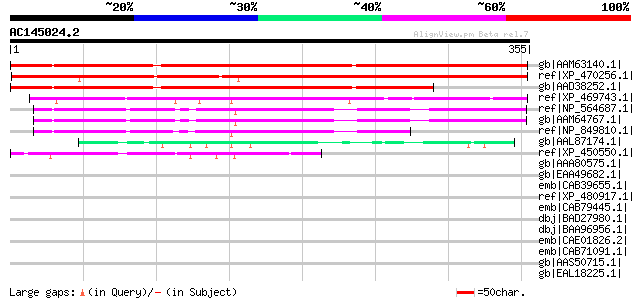
Query= AC145024.2 - phase: 0
(355 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63140.1| unknown [Arabidopsis thaliana] gi|21280797|gb|AAM... 410 e-113
ref|XP_470256.1| Hypothetical protein [Oryza sativa (japonica cu... 339 7e-92
gb|AAD38252.1| Hypothetical Protein [Arabidopsis thaliana] gi|25... 322 1e-86
ref|XP_469743.1| putative antifreeze glycoprotein precursor [Ory... 125 3e-27
ref|NP_564687.1| expressed protein [Arabidopsis thaliana] gi|123... 102 1e-20
gb|AAM64767.1| unknown [Arabidopsis thaliana] 98 4e-19
ref|NP_849810.1| expressed protein [Arabidopsis thaliana] 59 2e-07
gb|AAL87174.1| putative apospory-associated protein C [Oryza sat... 48 4e-04
ref|XP_450550.1| apospory-associated protein C-like [Oryza sativ... 47 9e-04
gb|AAA80575.1| possible apospory-associated protein gi|2501555|s... 46 0.001
gb|EAA49682.1| hypothetical protein MG08597.4 [Magnaporthe grise... 45 0.003
emb|CAB39655.1| possible apospory-associated like protein(fragme... 44 0.006
ref|XP_480917.1| putative Aldose 1-epimerase [Oryza sativa (japo... 44 0.006
emb|CAB79445.1| possible apospory-associated like protein [Arabi... 44 0.006
dbj|BAD27980.1| apospory-associated protein C-like [Oryza sativa... 44 0.007
dbj|BAA96956.1| apospory-associated protein C [Arabidopsis thali... 44 0.010
emb|CAE01826.2| OSJNBa0041A02.19 [Oryza sativa (japonica cultiva... 40 0.14
emb|CAB71091.1| putative protein [Arabidopsis thaliana] gi|15233... 39 0.18
gb|AAS50715.1| ABL056Cp [Ashbya gossypii ATCC 10895] gi|45185174... 39 0.18
gb|EAL18225.1| hypothetical protein CNBK2430 [Cryptococcus neofo... 39 0.24
>gb|AAM63140.1| unknown [Arabidopsis thaliana] gi|21280797|gb|AAM45001.1| unknown
protein [Arabidopsis thaliana]
gi|15028029|gb|AAK76545.1| unknown protein [Arabidopsis
thaliana] gi|18408126|ref|NP_564840.1| expressed protein
[Arabidopsis thaliana]
Length = 348
Score = 410 bits (1053), Expect = e-113
Identities = 213/356 (59%), Positives = 264/356 (73%), Gaps = 10/356 (2%)
Query: 1 MASFLSLSL-PKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRN 59
MAS +S SL PK +++S +A T ++T E L DKFGRKGIKF ESN+IP+VEL VRN
Sbjct: 1 MASLISFSLLPKPKAVRSSISAPQTQTINT-EKLEDKFGRKGIKFSESNNIPMVELKVRN 59
Query: 60 GSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGA 119
GSSL L L DAHV SYKPKV+WKD+G EE+LYT+ +E+ +GG+G+V+ +P
Sbjct: 60 GSSLKLSLSDAHVLSYKPKVYWKDEGFEEVLYTVDGDES-----RGGVGVVIVNGEEPKG 114
Query: 120 KELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKP 179
+ S +W+VKD D DAIDALQ+EL T D+TYIVSLYPVSMATA++VKN KP
Sbjct: 115 GSSVISGCDWSVKDTDSDAIDALQIELSCTAGVLDITYIVSLYPVSMATALVVKNNGRKP 174
Query: 180 VTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGA 239
VTL I+S+ RFK+R GA I+GL+ CSYC +PPL SPF++L+PSEAMK+ES FG+
Sbjct: 175 VTLKPGIMSYLRFKKRSGAGIQGLKGCSYCPNPPLSSPFELLSPSEAMKAESSGW--FGS 232
Query: 240 EPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVI 299
E KPG W + ITLLE KMSR++ APP ER KA YNTPPSK+E IDQGR +F+R+I
Sbjct: 233 EEGEKPGIWAVEDSVITLLEKKMSRIYGAPPAERLKAVYNTPPSKFETIDQGRGLFFRMI 292
Query: 300 RMGFEDIYVSSPGSMSDKYGRD-YFVCTGPASILEPVTVNPGEEFRGAQVIEHDNL 354
R+GFE++YV SPGSM DKYG+ YFVCTGP S+L PV V GE +RGA VIEHDNL
Sbjct: 293 RIGFEEMYVGSPGSMWDKYGKQHYFVCTGPTSMLVPVDVASGETWRGAMVIEHDNL 348
>ref|XP_470256.1| Hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|22773230|gb|AAN06836.1| Hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 399
Score = 339 bits (870), Expect = 7e-92
Identities = 185/399 (46%), Positives = 243/399 (60%), Gaps = 49/399 (12%)
Query: 2 ASFLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLE--SNSIPIVELTVRN 59
+S L L LP +SAA T TP++L + FGRKG++F + P EL+VRN
Sbjct: 4 SSLLPLHLPTRPSAVKASAAATAAAAPTPQSLEESFGRKGLRFAADPATGAPTAELSVRN 63
Query: 60 GSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGA 119
GSSL LRL D VTSY+PKV+WKDDG E+L+T+ G + KGG+GL L+EV GA
Sbjct: 64 GSSLQLRLADGLVTSYRPKVYWKDDGCREVLHTVAGAGAG-GEVKGGVGLALSEVSSSGA 122
Query: 120 KELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDM------------------------ 155
E L EW+V D D D+ DA+Q L++ N D+
Sbjct: 123 AESLLVGSEWSVVDADSDSYDAVQ--LVNDNWVLDLHPVLSRQAQTELDSLMTILASQQL 180
Query: 156 --------------------TYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRR 195
TY+V+LYP+SMATAV+VKN KPV+LT+A+LSH +F +R
Sbjct: 181 QHTEQDVELGCTKGSGTLEVTYVVTLYPLSMATAVMVKNNGKKPVSLTSAMLSHIKFDKR 240
Query: 196 GGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPI 255
G A++GL+ C YCSHPP + F +LTP+EAMK E G E + G WT +
Sbjct: 241 RGTAVEGLRGCPYCSHPPPAAGFALLTPAEAMKREDGGWFGGGGGEEPRQGVWTVEDNLY 300
Query: 256 TLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMS 315
T+L+ K+SRV+AAPP+ER K Y+T PSK+ IDQ + +RV+RMG+ED+Y+ SPG M
Sbjct: 301 TILKKKVSRVYAAPPEERKKRIYSTAPSKFTTIDQNSGLGFRVVRMGYEDMYLCSPGEMY 360
Query: 316 DKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNL 354
K+G+DYF+CTG AS+L PV VNPGEE+R AQVIEHDNL
Sbjct: 361 RKFGKDYFLCTGTASMLVPVVVNPGEEWRAAQVIEHDNL 399
>gb|AAD38252.1| Hypothetical Protein [Arabidopsis thaliana] gi|25373160|pir||H96670
hypothetical protein F13O11.8 [imported] - Arabidopsis
thaliana
Length = 338
Score = 322 bits (824), Expect = 1e-86
Identities = 169/291 (58%), Positives = 211/291 (72%), Gaps = 9/291 (3%)
Query: 1 MASFLSLSL-PKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRN 59
MAS +S SL PK +++S +A T ++T E L DKFGRKGIKF ESN+IP+VEL VRN
Sbjct: 1 MASLISFSLLPKPKAVRSSISAPQTQTINT-EKLEDKFGRKGIKFSESNNIPMVELKVRN 59
Query: 60 GSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGA 119
GSSL L L DAHV SYKPKV+WKD+G EE+LYT+ +E+ +GG+G+V+ +P
Sbjct: 60 GSSLKLSLSDAHVLSYKPKVYWKDEGFEEVLYTVDGDES-----RGGVGVVIVNGEEPKG 114
Query: 120 KELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKP 179
+ S +W+VKD D DAIDALQ+EL T D+TYIVSLYPVSMATA++VKN KP
Sbjct: 115 GSSVISGCDWSVKDTDSDAIDALQIELSCTAGVLDITYIVSLYPVSMATALVVKNNGRKP 174
Query: 180 VTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGA 239
VTL I+S+ RFK+R GA I+GL+ CSYC +PPL SPF++L+PSEAMK+ES FG+
Sbjct: 175 VTLKPGIMSYLRFKKRSGAGIQGLKGCSYCPNPPLSSPFELLSPSEAMKAESSGW--FGS 232
Query: 240 EPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQ 290
E KPG W + ITLLE KMSR++ APP ER KA YNTPPSK+E IDQ
Sbjct: 233 EEGEKPGIWAVEDSVITLLEKKMSRIYGAPPAERLKAVYNTPPSKFETIDQ 283
>ref|XP_469743.1| putative antifreeze glycoprotein precursor [Oryza sativa]
gi|18087673|gb|AAL58965.1| putative antifreeze
glycoprotein precursor [Oryza sativa]
Length = 411
Score = 125 bits (313), Expect = 3e-27
Identities = 105/360 (29%), Positives = 166/360 (45%), Gaps = 24/360 (6%)
Query: 14 LIKASSAATTTTPLSTP-----EALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLP 68
L AS+AA + P + L +F G+ F V++ +RNGS+ + LP
Sbjct: 57 LAPASTAADAAAAAAAPAPPNVDYLAAEFAGHGVSFEAVGGSCAVKMELRNGSAAHVLLP 116
Query: 69 DAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLN--EVLQPGAKELLPST 126
VTSYKP + W E + T+ G +GG+ L L G + P +
Sbjct: 117 GGLVTSYKPAM-WHGAPTEVIHTTVAEGLGGRAVIRGGVSLDLRCGGAAGGGGDGMPPWS 175
Query: 127 LE--WTVKDVDYDAIDALQVELISTN----RFFDMTYIVSLYPVSMATAVIVKNK-SPKP 179
W+++DV +++VEL S + +V+L+P ++AT +N SP P
Sbjct: 176 PSGAWSLRDVRGSPTGSIEVELASAAPPEASGVEARCVVTLHPEALATEFTARNAASPSP 235
Query: 180 VTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSES-----QRL 234
V L+ A+ +H R GLQ Y + P+ S F I+ P +S S +R
Sbjct: 236 VALSAAVSTHLRVSTPDATYAVGLQGSDYRAIDPVLSEFAIVPPDFMSRSSSATTLARRW 295
Query: 235 ISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREI 294
+ G + + G G + + + +E + Y+ P ++ IID+GR
Sbjct: 296 ATKGFDAVLSGGGGGGAGAQEA--DGEEDDDYKRMTEEMCR-IYSHAPRQFTIIDRGRRN 352
Query: 295 FYRVIRMGFEDIYVSSPGSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNL 354
V R GFE++YV SPGS YG+ +VC GPA +LEP+ ++PG + GAQ + + NL
Sbjct: 353 SICVQRRGFEEVYVFSPGSKYQWYGKYAYVCVGPA-MLEPIVLSPGATWSGAQYLRNPNL 411
>ref|NP_564687.1| expressed protein [Arabidopsis thaliana] gi|12321610|gb|AAG50849.1|
hypothetical protein [Arabidopsis thaliana]
gi|12323162|gb|AAG51558.1| hypothetical protein;
3570-2351 [Arabidopsis thaliana] gi|25405825|pir||A96596
hypothetical protein T18I3.2 [imported] - Arabidopsis
thaliana
Length = 354
Score = 102 bits (255), Expect = 1e-20
Identities = 86/341 (25%), Positives = 152/341 (44%), Gaps = 41/341 (12%)
Query: 17 ASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYK 76
AS++A+ P+ E L +F G F + I L + NGSS ++ L +TSYK
Sbjct: 50 ASASASPAFPIDV-EYLRREFSGHGATFEDIGETCIARLKLDNGSSANVMLTRGMITSYK 108
Query: 77 PKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDY 136
+V+ G ELL+T E +GG+ + E+ +W ++ +
Sbjct: 109 VRVW--HGGKVELLHTWVEQEEEEVVIRGGVSSAFRS---SDSDEIS----DWRLQGISG 159
Query: 137 DAIDALQVELISTNRFF---DMTYIVSLYPVSMATAVIVKNKSPKPVTLTN-AILSHFRF 192
D+ D +Q+EL +++ ++ I+SL +++ + + NK P+ L +++S+
Sbjct: 160 DSKDCVQMELRRSDKKIKEIELKQIISLRENTLSIELSMTNKGISPIKLEGCSLVSYLTV 219
Query: 193 KRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQG 252
GL+ + P F ++ G + E KPG ++
Sbjct: 220 STPEATYAVGLEGSDFVETTPFLPRFGVVQ---------------GEKEEEKPGFGGEEE 264
Query: 253 VPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG 312
L +MSR++ PK T +ID+GR V R GFE++Y+ SPG
Sbjct: 265 SNYKQLNREMSRIYTCAPKSFT------------VIDRGRRNSVVVGREGFEEVYMYSPG 312
Query: 313 SMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDN 353
S + Y + +VC GP+S+L P+++ G +RG + + N
Sbjct: 313 SRLESYTKSAYVCIGPSSLLSPISLESGCVWRGVLHLHNPN 353
>gb|AAM64767.1| unknown [Arabidopsis thaliana]
Length = 354
Score = 97.8 bits (242), Expect = 4e-19
Identities = 84/341 (24%), Positives = 151/341 (43%), Gaps = 41/341 (12%)
Query: 17 ASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYK 76
AS++A+ P+ E L +F G F + I L + +GSS ++ L +TSYK
Sbjct: 50 ASASASPAFPIDV-EYLRREFSGHGATFEDIGETCIARLKLDDGSSANVMLTRGMITSYK 108
Query: 77 PKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDY 136
+V+ G ELL+T E +GG+ + E+ +W ++ +
Sbjct: 109 VRVW--HGGKVELLHTWVEQEEEEVVIRGGVSSAFRS---SDSDEIS----DWRLQGISG 159
Query: 137 DAIDALQVELISTNRFF---DMTYIVSLYPVSMATAVIVKNKSPKPVTLTN-AILSHFRF 192
D+ D +Q+EL +++ ++ I+SL +++ + + NK P+ L +++S+
Sbjct: 160 DSKDCVQMELRRSDKKIKEIELKQIISLRENTLSIELSMTNKGISPIKLEGCSLVSYLTV 219
Query: 193 KRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQG 252
GL+ + P F ++ G + E KPG ++
Sbjct: 220 STPEATYAVGLEGSDFVETTPFLPRFGVVQ---------------GEKEEEKPGFGGEEE 264
Query: 253 VPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG 312
L +MSR++ PK T +ID+GR V R GFE++Y+ S G
Sbjct: 265 SNYKQLNREMSRIYTCAPKSFT------------VIDRGRRNSVVVGREGFEEVYMYSSG 312
Query: 313 SMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDN 353
S + Y + +VC GP+S+L P+++ G +RG + + N
Sbjct: 313 SRLESYTKSAYVCIGPSSLLSPISLESGCVWRGVLHLHNPN 353
>ref|NP_849810.1| expressed protein [Arabidopsis thaliana]
Length = 308
Score = 58.9 bits (141), Expect = 2e-07
Identities = 60/262 (22%), Positives = 110/262 (41%), Gaps = 29/262 (11%)
Query: 17 ASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYK 76
AS++A+ P+ E L +F G F + I L + NGSS ++ L +TSYK
Sbjct: 50 ASASASPAFPIDV-EYLRREFSGHGATFEDIGETCIARLKLDNGSSANVMLTRGMITSYK 108
Query: 77 PKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDY 136
+V+ G ELL+T E +GG+ + E+ +W ++ +
Sbjct: 109 VRVW--HGGKVELLHTWVEQEEEEVVIRGGVSSAFR---SSDSDEI----SDWRLQGISG 159
Query: 137 DAIDALQVELISTN---RFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTN-AILSHFRF 192
D+ D +Q+EL ++ + ++ I+SL +++ + + NK P+ L +++S+
Sbjct: 160 DSKDCVQMELRRSDKKIKEIELKQIISLRENTLSIELSMTNKGISPIKLEGCSLVSYLTV 219
Query: 193 KRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQG 252
GL+ + P F ++ G + E KPG ++
Sbjct: 220 STPEATYAVGLEGSDFVETTPFLPRFGVVQ---------------GEKEEEKPGFGGEEE 264
Query: 253 VPITLLENKMSRVFAAPPKERT 274
L +MSR++ PK T
Sbjct: 265 SNYKQLNREMSRIYTCAPKSFT 286
>gb|AAL87174.1| putative apospory-associated protein C [Oryza sativa (japonica
cultivar-group)] gi|50929321|ref|XP_474188.1|
OSJNBa0011F23.4 [Oryza sativa (japonica cultivar-group)]
gi|38346398|emb|CAE04231.2| OSJNBa0011F23.4 [Oryza
sativa (japonica cultivar-group)]
Length = 344
Score = 48.1 bits (113), Expect = 4e-04
Identities = 77/328 (23%), Positives = 129/328 (38%), Gaps = 83/328 (25%)
Query: 48 NSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKA---- 103
N + V L GSS + L HVTS WKD+ EELL+ + ++K
Sbjct: 37 NGLDKVVLREVRGSSAEVYLYGGHVTS------WKDEHGEELLFV---SNKAIFKPPKAI 87
Query: 104 KGGIGLVLNEVLQPGAKEL--LPSTLEWTVKD----VDYDAIDALQVELISTN------- 150
+GGI + + G + W+V++ + V+LI T+
Sbjct: 88 RGGIPICFPQFSNFGNLDPHGFARNRTWSVENDPPPFPVPTSNKAYVDLILTHTEEDLKI 147
Query: 151 --RFFDMTYIVSLYP---VSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQT 205
++ V+L P + + + + N KP + T A ++F ++GL+T
Sbjct: 148 WPHSYEFRLRVALGPGGDLMLTSRIRNTNADGKPFSFTFAYHTYFSISDISEVRVEGLET 207
Query: 206 CSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRV 265
Y + E R +T+QG I + E+++ R+
Sbjct: 208 LDYLDN----------------LQERNR--------------YTEQGDAI-VFESELDRI 236
Query: 266 FAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG-----SMSDKYGR 320
+ P SK IID ++ + V + G D V +P +MSD +G
Sbjct: 237 YLGTP------------SKIAIIDHEKKRTFVVRKGGLPDAVVWNPWDKKAKAMSD-FGD 283
Query: 321 DYF---VCTGPASILEPVTVNPGEEFRG 345
D + VC A+I +P+T+ PGEE+ G
Sbjct: 284 DEYKRMVCVEAAAIEKPITLKPGEEWTG 311
>ref|XP_450550.1| apospory-associated protein C-like [Oryza sativa (japonica
cultivar-group)] gi|48716905|dbj|BAD23600.1|
apospory-associated protein C-like [Oryza sativa
(japonica cultivar-group)]
Length = 319
Score = 47.0 bits (110), Expect = 9e-04
Identities = 57/233 (24%), Positives = 102/233 (43%), Gaps = 30/233 (12%)
Query: 1 MASFLSLSLPKLNLIKASSAATTTTP--------LSTPEALNDKFGRKGIKFLESNS-IP 51
MA+ +L+LP + I S + ++ +P + ++ K G+ E N +P
Sbjct: 1 MAASCALTLP--SAISVSVSVSSNSPRRFRRSRRVVAMASVGQKVYAPGVAVSEGNGGLP 58
Query: 52 IVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIP-ANETGLYKAKGGIGLV 110
++L +GS + L A VTS WK ++LL+ P A G GGI
Sbjct: 59 KIDLKSPHGSEAEIYLFGACVTS------WKVPSGKDLLFVRPDAVFNGQKPISGGIPHC 112
Query: 111 LNEVLQPGAKEL--LPSTLEWTVKDVDYDAID---ALQVELISTNRF-----FDMTYIVS 160
+ PG + + W++ D + + D L+++ S +R F Y V+
Sbjct: 113 FPQ-FGPGTMQQHGFARNMNWSITDSEANEGDPAVTLELKDDSYSRSMWDFSFQALYKVA 171
Query: 161 LYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPP 213
L+ S++T + + N KP + +A+ S+F G ++KGL+ C + P
Sbjct: 172 LHSTSLSTTLKITNTDDKPFSFNSALHSYFS-ASISGVSVKGLKGCKTLNKDP 223
>gb|AAA80575.1| possible apospory-associated protein
gi|2501555|sp|Q40784|AAPC_PENCL POSSIBLE
APOSPORY-ASSOCIATED PROTEIN C
Length = 329
Score = 46.2 bits (108), Expect = 0.001
Identities = 31/105 (29%), Positives = 53/105 (49%), Gaps = 20/105 (19%)
Query: 248 WTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIY 307
+T+QG I + E+++ +V+ A P SK IID ++ + V + G D
Sbjct: 214 FTEQGDAI-VFESEVDKVYLAAP------------SKIAIIDHEKKKTFVVTKEGLPDAV 260
Query: 308 VSSPGSMSDKYGRDY-------FVCTGPASILEPVTVNPGEEFRG 345
V +P K +D+ +C PA++ +P+T+ PGEE+RG
Sbjct: 261 VWNPWDKKAKAMQDFGDAEYKNMLCVEPAAVEKPITLKPGEEWRG 305
>gb|EAA49682.1| hypothetical protein MG08597.4 [Magnaporthe grisea 70-15]
gi|39946606|ref|XP_362840.1| hypothetical protein
MG08597.4 [Magnaporthe grisea 70-15]
Length = 326
Score = 45.1 bits (105), Expect = 0.003
Identities = 77/339 (22%), Positives = 118/339 (34%), Gaps = 89/339 (26%)
Query: 53 VELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGIGLVL- 111
VE + G S+ + L A V S WK +G E+L + A G +GGI LV
Sbjct: 32 VEAVLPTGESVEVLLYGATVLS------WKKNGQEKLWLSEGAKLDGSKAVRGGIPLVFP 85
Query: 112 --NEVLQPGAKELLP-------STLEWTVKDVDYDAIDA-------------LQVELIST 149
A LP ST E+ K A + L L S
Sbjct: 86 VFGTAPDHEATSALPQHGFARTSTWEFLGKSTSESAGEGDGAPRSMADLSVKLDFGLSSA 145
Query: 150 N----------RFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAA 199
N F + Y V+L ++ T++IV N+ KP+ + + ++ R K
Sbjct: 146 NLDEKTRALWPHPFGLLYSVTLDRDALTTSLIVTNEGDKPIEVQVLLHTYLRVKDINNVT 205
Query: 200 IKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLE 259
I+GL+ SY S T A+ E+ R+ + P P+T+LE
Sbjct: 206 IEGLENASYIDKVDNASTKTQDTAPLAINGETDRIYTPVGGPR----------APVTVLE 255
Query: 260 NKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMG-FEDIYVSSP------- 311
G + + V+R G +D+ V +P
Sbjct: 256 -------------------------------GGQKTFEVVRDGNLDDVVVWNPWTEKANS 284
Query: 312 -GSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVI 349
G K G + +C P ++ T+ G+ GAQ I
Sbjct: 285 IGDFEPKVGFNNMLCIEPGAVRAWQTLEAGDALEGAQTI 323
>emb|CAB39655.1| possible apospory-associated like protein(fragment) [Arabidopsis
thaliana]
Length = 239
Score = 44.3 bits (103), Expect = 0.006
Identities = 43/178 (24%), Positives = 73/178 (40%), Gaps = 35/178 (19%)
Query: 172 VKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSES 231
VKN KP T A+ +F ++GL Y + F ++
Sbjct: 80 VKNTDTKPFNFTFALHPYFAVSNISEIHVEGLHNLDYLDQQKNRTRF----------TDH 129
Query: 232 QRLISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQG 291
+++I+F A+ K L+ K+ R+ Y + P + I+D
Sbjct: 130 EKVITFNAQSITKKKH---------TLQKKLDRL------------YLSTPDQLRIVDHK 168
Query: 292 REIFYRVIRMGFEDIYVSSPGS--MSDKYGRDY--FVCTGPASILEPVTVNPGEEFRG 345
++ V + G D V +P +SD DY FV A++ +P+TVNPG+E++G
Sbjct: 169 KKKTIVVHKEGQVDAVVWNPWDKKVSDLGVEDYKRFVTVESAAVAKPITVNPGKEWKG 226
>ref|XP_480917.1| putative Aldose 1-epimerase [Oryza sativa (japonica
cultivar-group)] gi|40253263|dbj|BAD05401.1| putative
Aldose 1-epimerase [Oryza sativa (japonica
cultivar-group)] gi|40253632|dbj|BAD05576.1| putative
Aldose 1-epimerase [Oryza sativa (japonica
cultivar-group)]
Length = 337
Score = 44.3 bits (103), Expect = 0.006
Identities = 72/329 (21%), Positives = 127/329 (37%), Gaps = 83/329 (25%)
Query: 47 SNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKA--- 103
++ I V L G S + L A VTS WK+D EELL+ + ++K
Sbjct: 38 NSGIEKVLLRGTRGFSAEVYLYGAQVTS------WKNDHAEELLFV---SSKAIFKPPKA 88
Query: 104 -KGGIGLVLNEVLQPGAKEL--LPSTLEWTVKD--------------VDYDAIDALQVEL 146
+GGI + + G E WT+ D VD + +L
Sbjct: 89 IRGGIPICFPQFGTHGNLEQHGFARNRLWTIDDNPPPLPVNPAIKSFVDL-ILRPSDEDL 147
Query: 147 ISTNRFFDMTYIVSLYP---VSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGL 203
F+ V+L P +S+ + + N + + T A ++F ++GL
Sbjct: 148 KIWPHSFEFRLRVALGPNGDLSLTSRIRNTNTDGRSFSYTFAYHTYFSVSDISEVRVEGL 207
Query: 204 QTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMS 263
+T Y + +K + + +T+QG I + E+++
Sbjct: 208 ETMDYLDN---------------LKGKER---------------FTEQGDAI-VFESEID 236
Query: 264 RVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDY- 322
+V+ A P SK IID ++ + + + G D V +P K +D+
Sbjct: 237 KVYLAAP------------SKIAIIDHEKKRTFVLTKEGLPDAVVWNPWDKKAKAMQDFG 284
Query: 323 ------FVCTGPASILEPVTVNPGEEFRG 345
+C PA++ +P+T+ PGEE++G
Sbjct: 285 DGEYKHMLCVEPAAVEKPITLKPGEEWKG 313
>emb|CAB79445.1| possible apospory-associated like protein [Arabidopsis thaliana]
gi|4539308|emb|CAB39611.1| possible apospory-associated
like protein [Arabidopsis thaliana]
gi|7485483|pir||T04244 hypothetical protein F14M19.180 -
Arabidopsis thaliana
Length = 329
Score = 44.3 bits (103), Expect = 0.006
Identities = 43/178 (24%), Positives = 73/178 (40%), Gaps = 35/178 (19%)
Query: 172 VKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSES 231
VKN KP T A+ +F ++GL Y + F ++
Sbjct: 170 VKNTDTKPFNFTFALHPYFAVSNISEIHVEGLHNLDYLDQQKNRTRF----------TDH 219
Query: 232 QRLISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQG 291
+++I+F A+ K L+ K+ R+ Y + P + I+D
Sbjct: 220 EKVITFNAQSITKKKH---------TLQKKLDRL------------YLSTPDQLRIVDHK 258
Query: 292 REIFYRVIRMGFEDIYVSSPGS--MSDKYGRDY--FVCTGPASILEPVTVNPGEEFRG 345
++ V + G D V +P +SD DY FV A++ +P+TVNPG+E++G
Sbjct: 259 KKKTIVVHKEGQVDAVVWNPWDKKVSDLGVEDYKRFVTVESAAVAKPITVNPGKEWKG 316
>dbj|BAD27980.1| apospory-associated protein C-like [Oryza sativa (japonica
cultivar-group)]
Length = 319
Score = 43.9 bits (102), Expect = 0.007
Identities = 26/85 (30%), Positives = 43/85 (50%), Gaps = 8/85 (9%)
Query: 272 ERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDY-------FV 324
E K + + PP K IID ++ + + + G D+ V +P K D+ +
Sbjct: 213 EAEKVYLSAPP-KIVIIDHDKKRTFELRKEGLPDVVVWNPWDRKAKTILDFGEEEYKCML 271
Query: 325 CTGPASILEPVTVNPGEEFRGAQVI 349
C G A+ +P+T+ PGEE++G Q I
Sbjct: 272 CVGAANAEKPITLRPGEEWQGRQEI 296
>dbj|BAA96956.1| apospory-associated protein C [Arabidopsis thaliana]
gi|28059286|gb|AAO30044.1| unknown protein [Arabidopsis
thaliana] gi|20260440|gb|AAM13118.1| unknown protein
[Arabidopsis thaliana] gi|15242099|ref|NP_200543.1|
aldose 1-epimerase family protein [Arabidopsis thaliana]
Length = 312
Score = 43.5 bits (101), Expect = 0.010
Identities = 70/327 (21%), Positives = 126/327 (38%), Gaps = 81/327 (24%)
Query: 48 NSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKA---- 103
N + + L G S + L +HVTS WK++ EELL+ + ++K
Sbjct: 15 NGLDKIVLRESRGRSAEVYLYGSHVTS------WKNENGEELLHL---SSKAIFKPPKPI 65
Query: 104 KGGIGLVLNEVLQPGAKEL--LPSTLEWTVK----DVDYDAIDALQVELISTNRFFDMTY 157
+GGI L + G E W V+ + ++ + V+LI D+
Sbjct: 66 RGGIPLCFPQFSNFGTLESHGFARNRIWEVEANPPPLPLNSCSSAFVDLILRPTEDDLKI 125
Query: 158 IVSLYPVSMATAVIVK------------NKSPKPVTLTNAILSHFRFKRRGGAAIKGLQT 205
+ + + A+ + N KP T T A ++F ++GL+T
Sbjct: 126 WPNNFEFRLRIALGTEGELTLTSRIRNTNSDGKPFTFTFAYHTYFSVSDISEVRVEGLET 185
Query: 206 CSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRV 265
Y + +K + +T+QG IT E+++ ++
Sbjct: 186 LDYLDN---------------LKDRER---------------FTEQGDAITF-ESEVDKI 214
Query: 266 FAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPG-----SMSDKYGR 320
Y + P+K I+D ++ + + + G D V +P ++SD
Sbjct: 215 ------------YLSTPTKIAILDHEKKRTFVIRKDGLADAVVWNPWDKKSKTISDLGDE 262
Query: 321 DY--FVCTGPASILEPVTVNPGEEFRG 345
DY +C A+I P+T+ PGEE++G
Sbjct: 263 DYKHMLCVEAAAIERPITLKPGEEWKG 289
>emb|CAE01826.2| OSJNBa0041A02.19 [Oryza sativa (japonica cultivar-group)]
gi|50928507|ref|XP_473781.1| OSJNBa0041A02.19 [Oryza
sativa (japonica cultivar-group)]
Length = 325
Score = 39.7 bits (91), Expect = 0.14
Identities = 21/75 (28%), Positives = 39/75 (52%), Gaps = 7/75 (9%)
Query: 278 YNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDY-------FVCTGPAS 330
Y P+K IID ++ + + + G D + +P K +D+ +C PA+
Sbjct: 227 YLDAPAKIAIIDHEKKRTFVLRKDGLPDAVLWNPWDKRTKNMQDFGDEEYKHMLCVEPAA 286
Query: 331 ILEPVTVNPGEEFRG 345
+ +P+T+ PGEE++G
Sbjct: 287 VEKPITLKPGEEWKG 301
>emb|CAB71091.1| putative protein [Arabidopsis thaliana]
gi|15233165|ref|NP_191720.1| aldose 1-epimerase family
protein [Arabidopsis thaliana] gi|11281675|pir||T47953
hypothetical protein F2A19.210 - Arabidopsis thaliana
Length = 317
Score = 39.3 bits (90), Expect = 0.18
Identities = 28/108 (25%), Positives = 47/108 (42%), Gaps = 20/108 (18%)
Query: 249 TQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYV 308
T+QG IT E++M R + PK ++D R+ Y + + G D V
Sbjct: 204 TEQGDAITF-ESEMDRTYLRSPKV------------VAVLDHERKRTYVIGKEGLPDTVV 250
Query: 309 SSPGSMSDKYGRDY-------FVCTGPASILEPVTVNPGEEFRGAQVI 349
+P K D+ +C A++ P+T+ PGEE+ G ++
Sbjct: 251 WNPWEKKSKTMADFGDEEYKSMLCVDGAAVERPITLKPGEEWTGRLIL 298
>gb|AAS50715.1| ABL056Cp [Ashbya gossypii ATCC 10895] gi|45185174|ref|NP_982891.1|
ABL056Cp [Eremothecium gossypii]
Length = 296
Score = 39.3 bits (90), Expect = 0.18
Identities = 28/109 (25%), Positives = 49/109 (44%), Gaps = 19/109 (17%)
Query: 254 PITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSP-- 311
P+ ++ R++ +R +I+D+GR I + V R D+ V +P
Sbjct: 194 PVVAFHEEVDRIYQQVADDRV----------IQIVDKGRPI-HSVSRGNLPDVVVWNPWI 242
Query: 312 ---GSMSD---KYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNL 354
+M D K G +C P + + VT+ PGE++ G Q++ D L
Sbjct: 243 EKSANMQDFQPKEGYQKMICIEPGHVHDFVTLQPGEKWSGYQLLSKDTL 291
>gb|EAL18225.1| hypothetical protein CNBK2430 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57229876|gb|AAW46278.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58260770|ref|XP_567795.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 283
Score = 38.9 bits (89), Expect = 0.24
Identities = 27/80 (33%), Positives = 37/80 (45%), Gaps = 9/80 (11%)
Query: 278 YNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSP--------GSMSDKYGRDYFVCTGPA 329
Y PS+ +D G Y+V GFED + +P M DK G D +VC P
Sbjct: 202 YQKVPSREITVDDGAGSGYKVRFRGFEDCTIWNPQEATGSKMADMEDK-GWDKYVCIEPG 260
Query: 330 SILEPVTVNPGEEFRGAQVI 349
+ E T++P +E AQ I
Sbjct: 261 YVREFKTLSPSQELILAQTI 280
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 624,225,885
Number of Sequences: 2540612
Number of extensions: 26391558
Number of successful extensions: 56857
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 56811
Number of HSP's gapped (non-prelim): 47
length of query: 355
length of database: 863,360,394
effective HSP length: 129
effective length of query: 226
effective length of database: 535,621,446
effective search space: 121050446796
effective search space used: 121050446796
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC145024.2


