

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.5 - phase: 0 /pseudo
(966 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
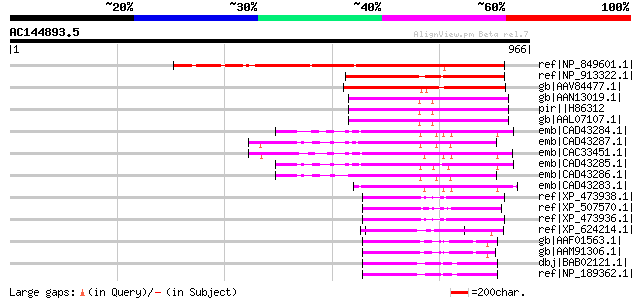
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_849601.1| DNA-binding bromodomain-containing protein [Ara... 494 e-138
ref|NP_913322.1| putative PSTVd RNA-biding protein [Oryza sativa... 288 5e-76
gb|AAV84477.1| At1g73150 [Arabidopsis thaliana] gi|15219397|ref|... 254 1e-65
gb|AAN13019.1| unknown protein [Arabidopsis thaliana] gi|1839453... 242 4e-62
pir||H86312 F2H15.2 protein - Arabidopsis thaliana gi|9665057|gb... 242 4e-62
gb|AAL07107.1| unknown protein [Arabidopsis thaliana] 240 2e-61
emb|CAD43284.1| bromodomain-containing RNA-binding protein 1 [Ni... 227 1e-57
emb|CAD43287.1| bromodomain-containing RNA-binding protein 2 [Ni... 227 2e-57
emb|CAC33451.1| PSTVd RNA-biding protein, Virp1 [Lycopersicon es... 227 2e-57
emb|CAD43285.1| bromodomain-containing RNA-binding protein 2 [Ni... 224 8e-57
emb|CAD43286.1| bromodomain-containing RNA-binding protein 1 [Ni... 224 1e-56
emb|CAD43283.1| bromodomain-containing RNA-binding protein 1 [So... 221 9e-56
ref|XP_473938.1| OSJNBb0085C12.15 [Oryza sativa (japonica cultiv... 182 5e-44
ref|XP_507570.1| PREDICTED OSJNBa0056O06.23 gene product [Oryza ... 173 2e-41
ref|XP_473936.1| OSJNBb0085C12.13 [Oryza sativa (japonica cultiv... 173 2e-41
ref|XP_624214.1| PREDICTED: similar to RING3 protein, partial [A... 151 1e-34
gb|AAF01563.1| hypothetical protein [Arabidopsis thaliana] gi|60... 144 1e-32
gb|AAM91306.1| unknown protein [Arabidopsis thaliana] gi|2046652... 144 2e-32
dbj|BAB02121.1| unnamed protein product [Arabidopsis thaliana] 143 2e-32
ref|NP_189362.1| DNA-binding bromodomain-containing protein [Ara... 143 2e-32
>ref|NP_849601.1| DNA-binding bromodomain-containing protein [Arabidopsis thaliana]
gi|15221424|ref|NP_172113.1| DNA-binding
bromodomain-containing protein [Arabidopsis thaliana]
gi|25406905|pir||A86198 hypothetical protein [imported]
- Arabidopsis thaliana gi|8844128|gb|AAF80220.1|
Contains similarity to a Ring3 protein from Homo sapiens
gi|133157 and contains a bromodomain PF|00439. EST
gb|F14211 comes from this gene. [Arabidopsis thaliana]
Length = 766
Score = 494 bits (1271), Expect = e-138
Identities = 294/632 (46%), Positives = 401/632 (62%), Gaps = 43/632 (6%)
Query: 306 DVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNSKLEDMTSQ 365
D N + DS ++++ E S QP+ +D S + + ++ +
Sbjct: 93 DKNLTEAPSENLPGDDSDKVIDKPLVEAFSQAQPQ-----DDASLAAMDKSEEVPSQIPK 147
Query: 366 QQD--NSILEDGNS-SQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDDLHINQQA 422
QD N+++ D NS +P +L E + V D N + SQ++S +
Sbjct: 148 AQDDVNTVVVDENSIKEPPKSLAQEDVTTVIV------DKNPIEAPSQTLSLEDGDTLVV 201
Query: 423 EPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLLEDDGPSSPIHRHKAVPS 482
+ + + V E+D +H A DNL ++ ++P +E D + S
Sbjct: 202 DKNPIEVSSEED-----VHVIDA--DNL-----IKEAHPENFVERDTTDAQQPAGLTSDS 249
Query: 483 TEYRHSENVTEEPNLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVG 542
+ ++ E + + RI+I++AST+KQ+K+E+ KLE +L+ VR +VK+IE K+G +G
Sbjct: 250 AHATAAGSMPMEEDADGRIRIHVASTTKQQKEEIRKKLEDQLNVVRGMVKKIEDKEGEIG 309
Query: 543 GYGNSNVVLGGGITNGGGAQRAHSEAGSVGVSRQ---PTRPLHQLSFPMFQNSRRASEGV 599
Y +S V++ GI NGGG R S S G+ R+ RP++QLS + +N++ +E V
Sbjct: 310 AYNDSRVLINTGINNGGG--RILSGFASAGLPREVIRAPRPVNQLSISVLENTQGVNEHV 367
Query: 600 EKEKRMPKANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSC 659
EKEKR PKANQFY NSEFLL DK PPAESNKKSK + KKQGG D+G G G+K FK+C
Sbjct: 368 EKEKRTPKANQFYRNSEFLLG-DKLPPAESNKKSKSSSKKQGG-DVGHGFGAGTKVFKNC 425
Query: 660 SSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAE 719
S+LLE+LM+H++ WVFN PVDV GLGL DY+TII +PMDLGT+K+ L KN YKSP+EFAE
Sbjct: 426 SALLERLMKHKHGWVFNAPVDVKGLGLLDYYTIIEHPMDLGTIKSALMKNLYKSPREFAE 485
Query: 720 DVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRY--GMDYGAPSPLS 777
DVRLTFHNAMTYNP+GQDVH MA L +IFE+RWA+IE+DYNREMR+ G + P+P
Sbjct: 486 DVRLTFHNAMTYNPEGQDVHLMAVTLLQIFEERWAVIEADYNREMRFVTGYEMNLPTPTM 545
Query: 778 R-RVPAFTPPPPLDMRRILDRQEPFARTPRS------MNNTPSSRTPAPKKPKAKDPNKR 830
R R+ PPPP+++R +DR + R P + + TPS RTPA KKPKA +PNKR
Sbjct: 546 RSRLGPTMPPPPINVRNTIDRADWSNRQPTTTPGRTPTSATPSGRTPALKKPKANEPNKR 605
Query: 831 DMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDR 890
DMTY+EKQKLS L +LP +KLDA+VQ++ K+N + +D+EIEVD D+VD E LWELDR
Sbjct: 606 DMTYEEKQKLSGHLQNLPPDKLDAIVQIVNKRNTAVKLRDEEIEVDIDSVDPETLWELDR 665
Query: 891 FVLNYKKSLSKNKRKAEQA-RERAEALQNSVQ 921
FV NYKK LSK KRKAE A + RAEA +NS Q
Sbjct: 666 FVTNYKKGLSKKKRKAELAIQARAEAERNSQQ 697
>ref|NP_913322.1| putative PSTVd RNA-biding protein [Oryza sativa (japonica
cultivar-group)]
Length = 507
Score = 288 bits (738), Expect = 5e-76
Identities = 152/300 (50%), Positives = 196/300 (64%), Gaps = 12/300 (4%)
Query: 625 PPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFNTPVDVDGL 684
PP +K K N G ++ + FKSC +LL +LM+H+++WVFNTPVD L
Sbjct: 112 PPVSKHKSKKGNPSSNPGLSAEARRKLYAPVFKSCGALLARLMKHKHSWVFNTPVDASAL 171
Query: 685 GLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQ 744
GLHDY TIIT PMDLGTVK+RL YKSP+EFA DVRLTF NAM YNPKGQDVH MAEQ
Sbjct: 172 GLHDYHTIITKPMDLGTVKSRLAAGHYKSPREFAGDVRLTFQNAMRYNPKGQDVHFMAEQ 231
Query: 745 LSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLDMRRILDRQEPFART 804
L +FE++W IE++ + +P P + A P +D ++L+R +
Sbjct: 232 LLNMFEEKWPEIEAEVAQL--------SPQPPTPSSAAPRKPKEIDNSKVLERSDSTVHA 283
Query: 805 PRSMNNTP---SSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRK 861
M TP + R P KKPKA++PNKR+MT+ EKQ+LS +L LP EKLD VVQ+++K
Sbjct: 284 -AGMEATPKQNTGRPPVLKKPKAREPNKREMTFWEKQRLSNNLQELPPEKLDNVVQIIKK 342
Query: 862 KNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQARERAEALQNSVQ 921
+NL L+Q DDEIEVD D+ D E LWELDRFV NYKKS+SKNKRKAE + + + ++
Sbjct: 343 RNLSLSQHDDEIEVDIDSFDVETLWELDRFVTNYKKSISKNKRKAENPVAGQDEMNHDIE 402
>gb|AAV84477.1| At1g73150 [Arabidopsis thaliana] gi|15219397|ref|NP_177458.1|
DNA-binding bromodomain-containing protein [Arabidopsis
thaliana] gi|12324313|gb|AAG52122.1| hypothetical
protein; 61711-63380 [Arabidopsis thaliana]
gi|5903104|gb|AAD55662.1| Highly similar to non
intermediate filament IFA binding protein [Arabidopsis
thaliana] gi|25406254|pir||D96757 hypothetical protein
T18K17.19 [imported] - Arabidopsis thaliana
Length = 461
Score = 254 bits (649), Expect = 1e-65
Identities = 149/322 (46%), Positives = 195/322 (60%), Gaps = 27/322 (8%)
Query: 622 DKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFNTPVDV 681
+ F P + K N K+GG + + KSC++LL KLM+H+ W+FNTPVDV
Sbjct: 86 NNFAPVPNKKLKTANGGKKGGVHGAAADKGTVQILKSCNNLLTKLMKHKSGWIFNTPVDV 145
Query: 682 DGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAM 741
LGLHDY II PMDLGTVKTRL+K+ YKSP EFAEDVRLTF+NAM YNP G DV+ M
Sbjct: 146 VTLGLHDYHNIIKEPMDLGTVKTRLSKSLYKSPLEFAEDVRLTFNNAMLYNPVGHDVYHM 205
Query: 742 AEQLSKIFEDRWAIIESDYNREMR-----YGMDYGAP----------SPLSRRVPAFTPP 786
AE L +FE++W +E+ Y +R +D+ AP PL P+ +PP
Sbjct: 206 AEILLNLFEEKWVPLETQYELLIRKQQPVRDIDFHAPVSTNTHNVEALPLPAPTPSLSPP 265
Query: 787 PPLDM--RRILDRQEPFARTPRSMNN--TPSSRTPAPKKPKAKDPNKRDMTYDEKQKLST 842
PP + R L+R E SM N P+ P+K + RD+T+DEK++LS
Sbjct: 266 PPPKVVENRTLERAE-------SMTNPVKPAVLPVVPEKLVEEASANRDLTFDEKRQLSE 318
Query: 843 SLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKN 902
L LP +KL+AVVQ+++K+ L+QQDDEIE+D D++D E LWEL RFV YK+SLSK
Sbjct: 319 DLQDLPYDKLEAVVQIIKKRTPELSQQDDEIELDIDSLDLETLWELFRFVTEYKESLSKK 378
Query: 903 KRKAEQARER-AEALQNSVQSS 923
K + ER AE+ NSV S
Sbjct: 379 KEEQGLDSERDAESFHNSVHES 400
Score = 42.0 bits (97), Expect = 0.098
Identities = 30/91 (32%), Positives = 48/91 (51%), Gaps = 4/91 (4%)
Query: 497 LEDR---IKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNSNVVLGG 553
LED +KI+L+S SK E + + KL++EL+ VRSL+KR+E + N +
Sbjct: 41 LEDNHQMMKISLSSISKLEVRNLKRKLQAELEEVRSLIKRLEPQGNNFAPVPNKKLKTAN 100
Query: 554 GITNGGGAQRAHSEAGSVGVSRQPTRPLHQL 584
G GG A ++ G+V + + L +L
Sbjct: 101 G-GKKGGVHGAAADKGTVQILKSCNNLLTKL 130
>gb|AAN13019.1| unknown protein [Arabidopsis thaliana] gi|18394534|ref|NP_564037.1|
DNA-binding bromodomain-containing protein [Arabidopsis
thaliana]
Length = 487
Score = 242 bits (618), Expect = 4e-62
Identities = 136/321 (42%), Positives = 190/321 (58%), Gaps = 23/321 (7%)
Query: 631 KKSKLNWKKQGGGDMGXGLQMGS-KFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDY 689
+ K+ GG G G G+ + FK+C+SLL KLM+H+ AWVFN PVD GLGLHDY
Sbjct: 107 RSKKVKTGNGGGKKSGHGADKGTVQIFKNCNSLLTKLMKHKSAWVFNVPVDAKGLGLHDY 166
Query: 690 FTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
I+ PMDLGTVKT+L K+ YKSP +FAEDVRLTF+NA+ YNP G DV+ AE L +F
Sbjct: 167 HNIVKEPMDLGTVKTKLGKSLYKSPLDFAEDVRLTFNNAILYNPIGHDVYRFAELLLNMF 226
Query: 750 EDRWAIIESDYN---------REMRYGMDYGAPSPLSRRVPAFT----------PPPPLD 790
ED+W IE Y+ R++ + + +P+ +PA PPPP
Sbjct: 227 EDKWVSIEMQYDNLHRKFKPTRDIEFPAPAPSIAPIVEPLPAIVPSPSPSSPPPPPPPPV 286
Query: 791 MRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDP--NKRDMTYDEKQKLSTSLHSLP 848
+L+ + ++ P + AP+K + ++ N RD+T +EK++LS L LP
Sbjct: 287 AAPVLENRTWEREESMTIPVEPEAVITAPEKAEEEEAPVNNRDLTLEEKRRLSEELQDLP 346
Query: 849 VEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQ 908
+KL+ VVQ+++K N L+Q+DDEIE+D D++D LWEL RFV YK+SLSK
Sbjct: 347 YDKLETVVQIIKKSNPELSQKDDEIELDIDSLDINTLWELYRFVTGYKESLSKKNEAHGF 406
Query: 909 ARER-AEALQNSVQSSRLPIS 928
ER AE++ NS+Q +S
Sbjct: 407 GSERDAESVHNSIQEPTTLVS 427
Score = 45.1 bits (105), Expect = 0.012
Identities = 29/72 (40%), Positives = 39/72 (53%), Gaps = 9/72 (12%)
Query: 501 IKINLASTSKQEKQEMCLKLESELDAVRSLVKRIE---------VKQGYVGGYGNSNVVL 551
+KI+L+S SK E + + KL+SELD VRSL+KR + K G VG
Sbjct: 56 LKISLSSISKLEVRNLKRKLKSELDEVRSLIKRFDPEANPGGSMAKSGVVGRSKKVKTGN 115
Query: 552 GGGITNGGGAQR 563
GGG +G GA +
Sbjct: 116 GGGKKSGHGADK 127
>pir||H86312 F2H15.2 protein - Arabidopsis thaliana gi|9665057|gb|AAF97259.1|
Contains similarity to female sterile homeotic-related
protein Frg-1 from Mus musculus gb|AF045462 and contains
a bromodomain PF|00439. [Arabidopsis thaliana]
Length = 440
Score = 242 bits (618), Expect = 4e-62
Identities = 136/321 (42%), Positives = 190/321 (58%), Gaps = 23/321 (7%)
Query: 631 KKSKLNWKKQGGGDMGXGLQMGS-KFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDY 689
+ K+ GG G G G+ + FK+C+SLL KLM+H+ AWVFN PVD GLGLHDY
Sbjct: 107 RSKKVKTGNGGGKKSGHGADKGTVQIFKNCNSLLTKLMKHKSAWVFNVPVDAKGLGLHDY 166
Query: 690 FTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
I+ PMDLGTVKT+L K+ YKSP +FAEDVRLTF+NA+ YNP G DV+ AE L +F
Sbjct: 167 HNIVKEPMDLGTVKTKLGKSLYKSPLDFAEDVRLTFNNAILYNPIGHDVYRFAELLLNMF 226
Query: 750 EDRWAIIESDYN---------REMRYGMDYGAPSPLSRRVPAFT----------PPPPLD 790
ED+W IE Y+ R++ + + +P+ +PA PPPP
Sbjct: 227 EDKWVSIEMQYDNLHRKFKPTRDIEFPAPAPSIAPIVEPLPAIVPSPSPSSPPPPPPPPV 286
Query: 791 MRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDP--NKRDMTYDEKQKLSTSLHSLP 848
+L+ + ++ P + AP+K + ++ N RD+T +EK++LS L LP
Sbjct: 287 AAPVLENRTWEREESMTIPVEPEAVITAPEKAEEEEAPVNNRDLTLEEKRRLSEELQDLP 346
Query: 849 VEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQ 908
+KL+ VVQ+++K N L+Q+DDEIE+D D++D LWEL RFV YK+SLSK
Sbjct: 347 YDKLETVVQIIKKSNPELSQKDDEIELDIDSLDINTLWELYRFVTGYKESLSKKNEAHGF 406
Query: 909 ARER-AEALQNSVQSSRLPIS 928
ER AE++ NS+Q +S
Sbjct: 407 GSERDAESVHNSIQEPTTLVS 427
Score = 45.1 bits (105), Expect = 0.012
Identities = 29/72 (40%), Positives = 39/72 (53%), Gaps = 9/72 (12%)
Query: 501 IKINLASTSKQEKQEMCLKLESELDAVRSLVKRIE---------VKQGYVGGYGNSNVVL 551
+KI+L+S SK E + + KL+SELD VRSL+KR + K G VG
Sbjct: 56 LKISLSSISKLEVRNLKRKLKSELDEVRSLIKRFDPEANPGGSMAKSGVVGRSKKVKTGN 115
Query: 552 GGGITNGGGAQR 563
GGG +G GA +
Sbjct: 116 GGGKKSGHGADK 127
>gb|AAL07107.1| unknown protein [Arabidopsis thaliana]
Length = 487
Score = 240 bits (612), Expect = 2e-61
Identities = 135/321 (42%), Positives = 189/321 (58%), Gaps = 23/321 (7%)
Query: 631 KKSKLNWKKQGGGDMGXGLQMGS-KFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDY 689
+ K+ GG G G G+ + FK+C+SLL KLM+H+ AWVFN PVD GLGLHDY
Sbjct: 107 RSKKVKTGNGGGKKSGHGADKGTVQIFKNCNSLLTKLMKHKSAWVFNVPVDAKGLGLHDY 166
Query: 690 FTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
I+ PMDLGTVK +L K+ YKSP +FAEDVRLTF+NA+ YNP G DV+ AE L +F
Sbjct: 167 HNIVKEPMDLGTVKIKLGKSLYKSPLDFAEDVRLTFNNAILYNPIGHDVYRFAELLLNMF 226
Query: 750 EDRWAIIESDYN---------REMRYGMDYGAPSPLSRRVPAFT----------PPPPLD 790
ED+W IE Y+ R++ + + +P+ +PA PPPP
Sbjct: 227 EDKWVSIEMQYDNLHRKFKPTRDIEFPAPAPSIAPIVEPLPAIVPSPSPSSPPPPPPPPV 286
Query: 791 MRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDP--NKRDMTYDEKQKLSTSLHSLP 848
+L+ + ++ P + AP+K + ++ N RD+T +EK++LS L LP
Sbjct: 287 AAPVLENRTWEREESMTIPVEPEAVITAPEKAEEEEAPVNNRDLTLEEKRRLSEELQDLP 346
Query: 849 VEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQ 908
+KL+ VVQ+++K N L+Q+DDEIE+D D++D LWEL RFV YK+SLSK
Sbjct: 347 YDKLETVVQIIKKSNPELSQKDDEIELDIDSLDINTLWELYRFVTGYKESLSKKNEAHGF 406
Query: 909 ARER-AEALQNSVQSSRLPIS 928
ER AE++ NS+Q +S
Sbjct: 407 GSERDAESVHNSIQEPTTLVS 427
Score = 45.1 bits (105), Expect = 0.012
Identities = 29/72 (40%), Positives = 39/72 (53%), Gaps = 9/72 (12%)
Query: 501 IKINLASTSKQEKQEMCLKLESELDAVRSLVKRIE---------VKQGYVGGYGNSNVVL 551
+KI+L+S SK E + + KL+SELD VRSL+KR + K G VG
Sbjct: 56 LKISLSSISKLEVRNLKRKLKSELDEVRSLIKRFDPEANPGGSMAKSGVVGRSKKVKTGN 115
Query: 552 GGGITNGGGAQR 563
GGG +G GA +
Sbjct: 116 GGGKKSGHGADK 127
>emb|CAD43284.1| bromodomain-containing RNA-binding protein 1 [Nicotiana
benthamiana]
Length = 615
Score = 227 bits (579), Expect = 1e-57
Identities = 169/499 (33%), Positives = 234/499 (46%), Gaps = 114/499 (22%)
Query: 496 NLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNSNVVLGGGI 555
NL + N+AS +K E E+ +L +EL+ +RSL RIE
Sbjct: 91 NLGGYMTYNVASYNKTELHELRSRLVAELEQIRSLKDRIE-------------------- 130
Query: 556 TNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNS 615
Q + S S G S++ + N R G K+ +
Sbjct: 131 ----SGQLSTSNPRSHGKSKK-----------LSGNKRATPSGSSKDPK----------- 164
Query: 616 EFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVF 675
K P N+ N+ GG + + M K C +L KLM+H+ W+F
Sbjct: 165 -------KLPNGVDNR----NFGNPGGVGVKGIIGM-ENMMKECRQVLGKLMKHKSGWIF 212
Query: 676 NTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKG 735
NTPVD + LGLHDY II PMDLGTVK+ L+ +Y +P EFA DVRLTF+NA+ YNPK
Sbjct: 213 NTPVDAEALGLHDYHQIIKRPMDLGTVKSNLSNCFYPTPFEFAADVRLTFNNALLYNPKT 272
Query: 736 QDVHAMAEQLSKIFEDRWAIIESDYNR-------------EMRYGMDYG-APSPLSRRVP 781
VH AEQL FED + I+ N+ + G+ + P+P + P
Sbjct: 273 DQVHGFAEQLLARFEDMFRPIQDKLNKLDGGSDRRDFHPTDELQGISWNHIPTPERVKKP 332
Query: 782 AFTPPPPLDMRR-----------ILDRQEPFARTP----RSMNNTPS--SRTPAPK---- 820
TP P + ++ L Q+P TP +S+ +TPS PAPK
Sbjct: 333 KPTPAPHISKKQERMMQNHSSALTLPVQQPPDNTPVVRQQSLLSTPSPVRAPPAPKPQSS 392
Query: 821 ------------KPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQ 868
KP+AKDPNKR+M+ +EK KL L SLP EK+ +VQ++RK+N L Q
Sbjct: 393 VAAKVPPMGKQPKPRAKDPNKREMSMEEKHKLGVGLQSLPQEKMPQLVQIIRKRNEHLAQ 452
Query: 869 QDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKA---------EQARERAEALQNS 919
DEIE+D +A+D E LWELDRFV N+KK +SK KR+A + + A S
Sbjct: 453 DGDEIELDIEALDTETLWELDRFVTNWKKMVSKTKRQALINNLGQPPSASAAASAATTTS 512
Query: 920 VQSSRLPISVQMKEMFPHP 938
V + P + + + F P
Sbjct: 513 VAEADAPSTSEKNDSFKKP 531
>emb|CAD43287.1| bromodomain-containing RNA-binding protein 2 [Nicotiana tabacum]
Length = 616
Score = 227 bits (578), Expect = 2e-57
Identities = 174/529 (32%), Positives = 241/529 (44%), Gaps = 125/529 (23%)
Query: 444 GAISDNLHSHQQVEP-SNPNVL-------------LEDDGPSSPIHRHKAVPSTEYRHSE 489
G + +SH Q+ P SNPN L D+ P+ S R E
Sbjct: 23 GFMGKTPYSHTQLNPNSNPNPKKKQKQFHHASNGRLTDESPAVTQTASDDAYSFNQRPIE 82
Query: 490 NVTEEP--NLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNS 547
+ T NL + N AS +K E E+ +L +EL+ +R+L RIE G S
Sbjct: 83 SSTNVDGLNLGGYMTYNXASYNKTELHELRSRLVAELEQIRNLKDRIES-----GQLSTS 137
Query: 548 NVVLGGGITNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPK 607
N R+H ++ + +++PT G K+ +
Sbjct: 138 N-------------PRSHGKSKKLSGNKRPT-----------------PSGSSKDPK--- 164
Query: 608 ANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLM 667
K P N+ N+ GGG + G+ K C+ +L KLM
Sbjct: 165 ---------------KLPNGVDNR----NFGNPGGGGV-KGIIGKENMMKECTQVLGKLM 204
Query: 668 EHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHN 727
+H+ W+FNTPVD + LGLHDY II P DLGT K+ L+ N+Y +P EFA DVRLTF+N
Sbjct: 205 KHKSGWIFNTPVDAEALGLHDYHQIIKRPXDLGTXKSNLSNNFYPTPFEFAADVRLTFNN 264
Query: 728 AMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNR--------------EMRYGMDYGAP 773
A+ YNPK VH AEQL FED + I+ N+ E++ P
Sbjct: 265 ALLYNPKTDQVHGFAEQLLARFEDMFRPIQDKLNKLDGGSGRRDFHAIDELQGSSWNHIP 324
Query: 774 SPLSRRVPAFTPPPPLDMRR-------------ILDRQEPFARTPRSMNNTPSSRTPAP- 819
+P + P TP P + ++ L Q+P P +P S TP+P
Sbjct: 325 TPERVKKPKPTPAPHISKKQERMMQNHSSASTPSLPVQQPPDNPPVVRQQSPLS-TPSPV 383
Query: 820 ----------------------KKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQ 857
KP+AKDPNKR+M+ +EK KL L SLP EK+ +VQ
Sbjct: 384 RAPPAAKPQSSVAAKVPPMGKQPKPRAKDPNKREMSMEEKHKLGVGLQSLPQEKMPQLVQ 443
Query: 858 MMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKA 906
++RK+N L Q DEIE+D +A+D E LWELDRFV N+KK +SK KR+A
Sbjct: 444 IIRKRNEHLAQDGDEIELDIEALDTETLWELDRFVTNWKKMVSKTKRQA 492
>emb|CAC33451.1| PSTVd RNA-biding protein, Virp1 [Lycopersicon esculentum]
gi|13186136|emb|CAC33450.1| PSTVd RNA-biding protein,
Virp1 [Lycopersicon esculentum]
gi|13186134|emb|CAC33449.1| PSTVd RNA-biding protein,
Virp1 [Lycopersicon esculentum]
gi|13186132|emb|CAC33448.1| PSTVd RNA-biding protein,
Virp1 [Lycopersicon esculentum]
gi|10179608|gb|AAG13813.1| PSTVd RNA-binding protein
Virp1d [Lycopersicon esculentum]
gi|10179606|gb|AAG13812.1| PSTVd RNA-binding protein
Virp1c [Lycopersicon esculentum]
gi|10179604|gb|AAG13811.1| PSTVd RNA-binding protein
Virp1b [Lycopersicon esculentum]
gi|10179602|gb|AAG13810.1| PSTVd RNA-binding protein
Virp1a [Lycopersicon esculentum]
Length = 602
Score = 227 bits (578), Expect = 2e-57
Identities = 181/555 (32%), Positives = 247/555 (43%), Gaps = 121/555 (21%)
Query: 444 GAISDNLHSHQQVEPS-NPNVLLE-------------DDGPSSPIHRHKAVPSTEYRHSE 489
G + +SH Q+ P+ NPN + DD P+ S R E
Sbjct: 23 GFMGKTPYSHTQLNPNHNPNPKKKQKQFHHTSNGRHMDDSPAVTQTASDDAYSFNQRPIE 82
Query: 490 NVTEEPNLE--DRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNS 547
+ T L + N+ S +K E E+ +L +E++ +R+L RIE Q
Sbjct: 83 STTNVDGLNFGGYLTFNVVSYNKAEVNELRSRLMAEVEQIRNLKDRIESGQL-------- 134
Query: 548 NVVLGGGITNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPK 607
TN ++ ++G N R G K+ +
Sbjct: 135 ------STTNPRSQGKSKKQSG---------------------NKRPTPSGSSKDLK--- 164
Query: 608 ANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLM 667
K P N+ N+ GG D + S K C +L KLM
Sbjct: 165 ---------------KLPNGVENR----NFGNPGGVDGVKAIGTES-MMKECRQILAKLM 204
Query: 668 EHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHN 727
+H+ W+FN PVD + LGLHDY II PMDLGTVK+ L KN+Y SP EFA DVRLTF+N
Sbjct: 205 KHKNGWIFNIPVDAEALGLHDYHQIIKRPMDLGTVKSNLAKNFYPSPFEFAADVRLTFNN 264
Query: 728 AMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDY------------GAPSP 775
A+ YNPK V+A AEQL FED + ++ N+ DY P+P
Sbjct: 265 ALLYNPKTDQVNAFAEQLLGRFEDMFRPLQDKMNKLEGGRRDYHPVDELQGSSWNHIPTP 324
Query: 776 LSRRVPAFTPPPPLDMRRILDRQEPFARTP-------------RSMNNTPSS-RTPAPK- 820
+ P TP P + ++ + A TP +S +TPS R PA K
Sbjct: 325 ERVKKPKPTPVPNISKKQERMQNHSSASTPSLPVPPPNPPARQQSPLSTPSPVRAPAAKP 384
Query: 821 -------------KPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LN 867
KP+AKDPNKR+M +EK KL L SLP EK+ +VQ++RK+N L
Sbjct: 385 QSAAKVPTMGKQPKPRAKDPNKREMNMEEKHKLGVGLQSLPQEKMPQLVQIIRKRNEHLA 444
Query: 868 QQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKA-------EQARERAEALQNSV 920
Q DEIE+D +A+D E LWELDRFV N+KK +SK KR+A A A A SV
Sbjct: 445 QDGDEIELDIEALDTETLWELDRFVTNWKKMVSKTKRQALMNNLGPPSASAAASAATTSV 504
Query: 921 QSSRLPISVQMKEMF 935
+ P + + + F
Sbjct: 505 AEADGPTTSEKNDSF 519
>emb|CAD43285.1| bromodomain-containing RNA-binding protein 2 [Nicotiana
benthamiana]
Length = 613
Score = 224 bits (572), Expect = 8e-57
Identities = 166/501 (33%), Positives = 232/501 (46%), Gaps = 117/501 (23%)
Query: 496 NLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNSNVVLGGGI 555
NL + N+AS SK E ++ +L +EL+ +R+L RIE G SN
Sbjct: 91 NLGGYMTYNVASYSKTELHDLRSRLVAELEQIRNLKDRIES-----GQLSTSN------- 138
Query: 556 TNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNS 615
R+H ++ + +++PT G K+ +
Sbjct: 139 ------PRSHGKSKKLSGNKRPT-----------------PSGSSKDPK----------- 164
Query: 616 EFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVF 675
K P N+ N+ GGG + + M K C +L KLM+H+ W+F
Sbjct: 165 -------KLPNGVDNR----NFGNPGGGGVKGIIGM-ENMMKECRQVLGKLMKHKSGWIF 212
Query: 676 NTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKG 735
NTPVD + LGLHDY II PMDLGTVK+ L+ N Y +P EFA DVRLTF+NA+ YNPK
Sbjct: 213 NTPVDAEALGLHDYRQIIKRPMDLGTVKSNLSNNLYPTPFEFAADVRLTFNNALLYNPKT 272
Query: 736 QDVHAMAEQLSKIFEDRWAIIESDYNR--------------EMRYGMDYGAPSPLSRRVP 781
VH AE L FED + + N+ E++ P+P + P
Sbjct: 273 DQVHVFAELLLTRFEDMFRPFQDKLNKLDGGSGRRDFHAIDELQGSSWNHIPTPERVKKP 332
Query: 782 AFTPPP-------------------------PLDMRRILDRQEPFARTPRSMNNTPSSR- 815
TP P P D ++ +Q P + TP + P+++
Sbjct: 333 KPTPAPHISKKQERMMQNHSSASTPSLPVQQPPDNPPVVQQQSPLS-TPSPVRAPPAAKP 391
Query: 816 --------TPAPK--KPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL* 865
P K KP+AKDPNKR+M+ +EK KL L SLP EK+ +VQ++RK+N
Sbjct: 392 QSSVAAKVPPMEKQPKPRAKDPNKREMSMEEKHKLGVGLQSLPQEKMPQLVQIIRKRNEH 451
Query: 866 LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKA--------EQARERAEALQ 917
L Q DEIE+D +A+D E LWELDRFV N+KK +SK KR+A A A A
Sbjct: 452 LAQDGDEIELDIEALDTETLWELDRFVTNWKKMVSKTKRQALINNLGQPPSASAAASAAT 511
Query: 918 NSVQSSRLPISVQMKEMFPHP 938
SV + P + + + F P
Sbjct: 512 TSVAEADGPSTSEKNDSFKKP 532
>emb|CAD43286.1| bromodomain-containing RNA-binding protein 1 [Nicotiana tabacum]
Length = 617
Score = 224 bits (571), Expect = 1e-56
Identities = 159/461 (34%), Positives = 218/461 (46%), Gaps = 108/461 (23%)
Query: 496 NLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGNSNVVLGGGI 555
NL + N+AS +K E E+ +L +EL+ +R+L RIE G SN
Sbjct: 91 NLGGYMTYNVASYNKTELHELRSRLVAELEQIRNLKDRIES-----GQLSTSN------- 138
Query: 556 TNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNS 615
R+H ++ + +++PT G K+++
Sbjct: 139 ------PRSHGKSKKLSGNKRPT-----------------PSGSSKDRKK---------- 165
Query: 616 EFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVF 675
LL N N+ GGG G+ K C +L KLM+H+ W+F
Sbjct: 166 --LL----------NGVDNRNFGNPGGGGGVKGIIGMENMMKECRQVLAKLMKHKSGWIF 213
Query: 676 NTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKG 735
NTPVD +GLHDY II PMDLGTVK+ L N+Y +P EFA DVRLTF+NA+ YNPK
Sbjct: 214 NTPVDAKAMGLHDYHQIIKRPMDLGTVKSNLINNFYPTPFEFAADVRLTFNNALLYNPKT 273
Query: 736 QDVHAMAEQLSKIFEDRWAIIESDYNR--------------EMRYGMDYGAPSPLSRRVP 781
VH AEQL FED + I+ N+ E++ P+P + P
Sbjct: 274 DQVHGFAEQLLARFEDMFRPIQDKLNKLDGGTGRRDFRAIDELQGSSWNHIPTPERVKKP 333
Query: 782 AFTPPPPLDMRR-------------ILDRQEPFARTPRSMNNTPSSRTPAP--------- 819
TP P + ++ L Q+P P +P S TP+P
Sbjct: 334 KPTPAPHISKKQERMMQNHSSASTPSLPVQQPPDNPPVVRQQSPLS-TPSPVRAPPAAKP 392
Query: 820 --------------KKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL* 865
KP+AKDPNKR+M+ +EK KL L SLP EK+ +VQ++RK+N
Sbjct: 393 QSSVAAKVPPMGKQPKPRAKDPNKREMSMEEKHKLGVGLQSLPQEKMPQLVQIIRKRNEH 452
Query: 866 LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKA 906
L Q DEIE+D +A+D E LWELDRFV N+KK +SK KR+A
Sbjct: 453 LAQDGDEIELDIEALDTETLWELDRFVTNWKKMVSKTKRQA 493
>emb|CAD43283.1| bromodomain-containing RNA-binding protein 1 [Solanum tuberosum]
Length = 602
Score = 221 bits (563), Expect = 9e-56
Identities = 142/353 (40%), Positives = 184/353 (51%), Gaps = 53/353 (15%)
Query: 641 GGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLG 700
GGG G + K C +L KLM+H+ W+FN PVD + LGLHDY II P+DLG
Sbjct: 181 GGGVKAIGTE---SMMKECRQILAKLMKHKNGWIFNIPVDAEALGLHDYHQIIKRPIDLG 237
Query: 701 TVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDY 760
TVK+ L KN+Y SP EFA DVRLTF+NA+ YNPK V+ AEQL FED + ++
Sbjct: 238 TVKSNLAKNFYPSPFEFAADVRLTFNNALLYNPKTDQVNGFAEQLLGRFEDMFRPLQDKM 297
Query: 761 NREMRYGMDY------------GAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTP--- 805
N+ DY P+P + P TP P + ++ + A TP
Sbjct: 298 NKLEGGRRDYHPVDELQGSSWNHIPTPERVKKPKATPVPHISKKQERMQNHSSASTPSLP 357
Query: 806 ----------RSMNNTPSS-RTPAPK--------------KPKAKDPNKRDMTYDEKQKL 840
+S +TPS R P K KP+AKDPNKR M +EK KL
Sbjct: 358 VPPPNPPARQQSPLSTPSPVRAPPSKPESAAKVPAMGKQPKPRAKDPNKRVMNMEEKHKL 417
Query: 841 STSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLS 900
L SLP EK+ +VQ++RK+N L Q DEIE+D +A+D E LWELDRFV N+KK +S
Sbjct: 418 GVGLQSLPQEKMPQLVQIIRKRNEHLAQDGDEIELDIEALDTETLWELDRFVTNWKKMVS 477
Query: 901 KNKRKA--------EQARERAEALQNSVQSSRLPISVQMKEMFPHPCLCKGEV 945
K KR+A A A A SV + P + + + F P KG+V
Sbjct: 478 KTKRQALMINNLGPPSASAAASAATTSVAEADGPTTSEKNDSFKKP--KKGDV 528
>ref|XP_473938.1| OSJNBb0085C12.15 [Oryza sativa (japonica cultivar-group)]
gi|38345713|emb|CAD41835.2| OSJNBb0085C12.15 [Oryza
sativa (japonica cultivar-group)]
gi|38344165|emb|CAE03496.2| OSJNBa0053K19.4 [Oryza
sativa (japonica cultivar-group)]
Length = 456
Score = 182 bits (462), Expect = 5e-44
Identities = 110/264 (41%), Positives = 146/264 (54%), Gaps = 23/264 (8%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +L KL + + + FN PV+VD LGLHDY +I PMDLGTV+ L Y S +
Sbjct: 122 KRCEQILAKLRKDKRSIWFNAPVEVDRLGLHDYHAVIKCPMDLGTVRANLAAGRYPSHDD 181
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA DVRLTF NA+ YNP G +VH A L FE + S + +E++ P P+
Sbjct: 182 FAADVRLTFSNALRYNPAGHEVHTFAGDLLASFEKMYKASMSWFEQELKL---LEPPMPV 238
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
PPP L P A P + P + +KPKA++PNKRDMT +E
Sbjct: 239 --------PPPEL----------PPATAPVQVK--PRAANVKMRKPKAREPNKRDMTLEE 278
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYK 896
K L L SLP EK+ V+Q++RK+N EIE+D D +D E WELDRFV N+K
Sbjct: 279 KNLLRVGLESLPEEKMHNVLQIVRKRNGNPELVGGEIELDIDEMDVETQWELDRFVNNFK 338
Query: 897 KSLSKNKRKAEQARERAEALQNSV 920
K+L+K++R A E A+ + SV
Sbjct: 339 KALNKSRRAAIVNGENADVIDASV 362
>ref|XP_507570.1| PREDICTED OSJNBa0056O06.23 gene product [Oryza sativa (japonica
cultivar-group)] gi|50941927|ref|XP_480491.1| putative
RNA-binding protein Virp1a [Oryza sativa (japonica
cultivar-group)] gi|51964744|ref|XP_507156.1| PREDICTED
OSJNBa0056O06.23 gene product [Oryza sativa (japonica
cultivar-group)] gi|40253661|dbj|BAD05604.1| putative
RNA-binding protein Virp1a [Oryza sativa (japonica
cultivar-group)]
Length = 481
Score = 173 bits (439), Expect = 2e-41
Identities = 106/259 (40%), Positives = 145/259 (55%), Gaps = 20/259 (7%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C+ +L +L + + + FN+PVDV+ L LHDY II NPMDLGTVK L Y S +
Sbjct: 139 KRCTQILTRLRKQKISVWFNSPVDVERLKLHDYHAIIRNPMDLGTVKENLAFGRYPSHEA 198
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA DVRLTF NA+ YNP VH A L FE + S + +E + P PL
Sbjct: 199 FATDVRLTFSNALRYNPADHHVHRYASNLLATFEGLYKEALSWFEQECQ---RLEPPMPL 255
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ P P PP+ M P PR R P KPKA++PNKR+M+ +E
Sbjct: 256 ALPPP---PQPPVPM--------PMQAPPRIGG---GGRRP---KPKAREPNKREMSDEE 298
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYK 896
K KL + +LP EK+ V+Q+++K+N + +E+DFD +D E LWELDRFV+N K
Sbjct: 299 KHKLRVEIGNLPEEKMGNVLQIVQKRNTDPALMGEVVELDFDEMDVETLWELDRFVVNCK 358
Query: 897 KSLSKNKRKAEQARERAEA 915
K+LSK++R + +A
Sbjct: 359 KALSKSRRTVAMNGDAVDA 377
>ref|XP_473936.1| OSJNBb0085C12.13 [Oryza sativa (japonica cultivar-group)]
gi|38345711|emb|CAD41833.2| OSJNBb0085C12.13 [Oryza
sativa (japonica cultivar-group)]
gi|38344163|emb|CAE03494.2| OSJNBa0053K19.2 [Oryza
sativa (japonica cultivar-group)]
Length = 299
Score = 173 bits (439), Expect = 2e-41
Identities = 105/264 (39%), Positives = 144/264 (53%), Gaps = 23/264 (8%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +L KL + + + FN PV+VD LGL DY +I PMDLGTV+ L Y S +
Sbjct: 3 KRCDQILAKLRKDKRSIWFNAPVEVDRLGLQDYHAVIKCPMDLGTVRANLAAGRYPSHDD 62
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA D+RLTF NA+ YNP G +VH A L FE + S + +E++ P P+
Sbjct: 63 FAADIRLTFSNALRYNPAGHEVHTFAGDLLASFEKMYKASVSWFEQELKI---LEPPMPV 119
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
PPP L P A+ P + P + +K KA++PNKR+MT +E
Sbjct: 120 --------PPPEL----------PPAKAPAQVK--PRAGNVKMRKTKAREPNKREMTLEE 159
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYK 896
K L L SLP EK+ V+Q++RK+N EIE+D D +D E WELDRFV +K
Sbjct: 160 KNLLRVGLESLPEEKMHNVLQIVRKRNGNPELVGGEIELDIDEMDVETQWELDRFVNKFK 219
Query: 897 KSLSKNKRKAEQARERAEALQNSV 920
K+L+K++R A E A+ + SV
Sbjct: 220 KALNKSRRAAIVNGENADVIDASV 243
>ref|XP_624214.1| PREDICTED: similar to RING3 protein, partial [Apis mellifera]
Length = 614
Score = 151 bits (381), Expect = 1e-34
Identities = 100/282 (35%), Positives = 151/282 (53%), Gaps = 35/282 (12%)
Query: 653 SKFFKSCSSLLEKLMEHQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKN 709
S+ KSC+ +L++L + YAW F PVD + LGLHDY II PMDLGTVKT+++
Sbjct: 288 SEALKSCNEILKELFSKKHSGYAWPFYKPVDAELLGLHDYHDIIKKPMDLGTVKTKMDNR 347
Query: 710 WYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMD 769
YK+ +EFA DVRL F N YNP DV AMA +L +FE R+A I +
Sbjct: 348 EYKTAQEFASDVRLIFTNCYKYNPPDHDVVAMARKLQDVFEMRYAKIPDE---------- 397
Query: 770 YGAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDP-N 828
P+ + A MR++++ + + T SS P P ++D N
Sbjct: 398 -----PMGSILKAMQE----QMRKLVEESGKKKSKKKKPDKTKSSSQPPPITFDSEDEDN 448
Query: 829 KRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQD-DEIEVDFDAVDAEILWE 887
+ M+YDEK++LS ++ LP +KL VV +++ + L + DEIE+DF+ + L E
Sbjct: 449 AKPMSYDEKRQLSLDINKLPGDKLGRVVHIIQSREPSLRDSNPDEIEIDFETLKPSTLRE 508
Query: 888 LDRFVLN---------YKK--SLSKNKRKAEQARERAEALQN 918
L+ +V + YKK SK+++ AE+ +E + LQ+
Sbjct: 509 LESYVASCLRKKPRKLYKKVSGKSKDEQMAEKKQELEKRLQD 550
Score = 89.4 bits (220), Expect = 5e-16
Identities = 62/186 (33%), Positives = 87/186 (46%), Gaps = 28/186 (15%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+L+ + +HQ+AW F PVD L L DY II PMDLGT+K RL +Y S KE +D
Sbjct: 135 VLKPVWKHQFAWPFQQPVDAKKLNLPDYHKIIKQPMDLGTIKKRLENTYYWSGKECIQDF 194
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVP 781
F N YN G+DV MA+ L K+F + A + + ++ P P +VP
Sbjct: 195 NTMFTNCYVYNKPGEDVVVMAQALEKLFLTKVAQMPKE-------EVELEPPVPKGPKVP 247
Query: 782 -AFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKL 840
A T PP +P A+ + + + TP P+ K K+KL
Sbjct: 248 TAPTVMPP---------SQP-AKLKKGVKRKADTTTPTPQHTTGK----------SKEKL 287
Query: 841 STSLHS 846
S +L S
Sbjct: 288 SEALKS 293
>gb|AAF01563.1| hypothetical protein [Arabidopsis thaliana]
gi|6091741|gb|AAF03453.1| hypothetical protein
[Arabidopsis thaliana]
Length = 601
Score = 144 bits (363), Expect = 1e-32
Identities = 99/258 (38%), Positives = 134/258 (51%), Gaps = 36/258 (13%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C SLL++LM Q+ W+FNTPVDV L + DYFTII +PMDLGTVK++L Y SP E
Sbjct: 131 KQCESLLKRLMSQQHCWLFNTPVDVVKLNIPDYFTIIKHPMDLGTVKSKLTSGTYSSPSE 190
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
F+ DVRLTF NAMTYNP +V+ A+ LSK FE RW IE + PS L
Sbjct: 191 FSADVRLTFRNAMTYNPSDNNVYRFADTLSKFFEVRWKTIEKKSSGTK------SEPSNL 244
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ P EP A+ R MN A K+ +P KR MT ++
Sbjct: 245 ATLAHKDIAIP-----------EPVAKK-RKMN--------AVKRNSLLEPAKRVMTDED 284
Query: 837 KQKLSTSLHSL---PVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWEL----D 889
+ KL L SL PV+ ++ + K+ DDEIE+D + + + L++L D
Sbjct: 285 RVKLGRDLGSLTEFPVQIINFLRDHSSKEE---RSGDDEIEIDINDLSHDALFQLRDLFD 341
Query: 890 RFVLNYKKSLSKNKRKAE 907
F+ +K S + +E
Sbjct: 342 EFLRENQKKDSNGEPWSE 359
>gb|AAM91306.1| unknown protein [Arabidopsis thaliana] gi|20466526|gb|AAM20580.1|
unknown protein [Arabidopsis thaliana]
gi|16323117|gb|AAL15293.1| AT3g01770/F28J7_10
[Arabidopsis thaliana] gi|18395937|ref|NP_566151.1|
DNA-binding bromodomain-containing protein [Arabidopsis
thaliana]
Length = 620
Score = 144 bits (362), Expect = 2e-32
Identities = 98/254 (38%), Positives = 132/254 (51%), Gaps = 36/254 (14%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C SLL++LM Q+ W+FNTPVDV L + DYFTII +PMDLGTVK++L Y SP E
Sbjct: 131 KQCESLLKRLMSQQHCWLFNTPVDVVKLNIPDYFTIIKHPMDLGTVKSKLTSGTYSSPSE 190
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
F+ DVRLTF NAMTYNP +V+ A+ LSK FE RW IE + PS L
Sbjct: 191 FSADVRLTFRNAMTYNPSDNNVYRFADTLSKFFEVRWKTIEKKSSGTK------SEPSNL 244
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ P EP A+ R MN A K+ +P KR MT ++
Sbjct: 245 ATLAHKDIAIP-----------EPVAKK-RKMN--------AVKRNSLLEPAKRVMTDED 284
Query: 837 KQKLSTSLHSL---PVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWEL----D 889
+ KL L SL PV+ ++ + K+ DDEIE+D + + + L++L D
Sbjct: 285 RVKLGRDLGSLTEFPVQIINFLRDHSSKEE---RSGDDEIEIDINDLSHDALFQLRDLFD 341
Query: 890 RFVLNYKKSLSKNK 903
F+ +K S +
Sbjct: 342 EFLRENQKKDSNGE 355
>dbj|BAB02121.1| unnamed protein product [Arabidopsis thaliana]
Length = 818
Score = 143 bits (361), Expect = 2e-32
Identities = 95/252 (37%), Positives = 123/252 (48%), Gaps = 24/252 (9%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +LL KL H ++WVF PVDV L + DY T I +PMDLGTVK L Y SP E
Sbjct: 178 KQCDTLLRKLWSHPHSWVFQAPVDVVKLNIPDYLTTIKHPMDLGTVKKNLASGVYSSPHE 237
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA DVRLTF NAMTYNP G DVH M + LSK+FE RW I+ P
Sbjct: 238 FAADVRLTFTNAMTYNPPGHDVHIMGDILSKLFEARWKTIKKK------------LPPCS 285
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ +PA T P D R+ P + R M + P P KP MT E
Sbjct: 286 MQTLPAVTLEPN-DERKAAISVPPAKK--RKMASPVRESVPEPVKPL--------MTEVE 334
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQ-QDDEIEVDFDAVDAEILWELDRFVLNY 895
+ +L L SL E ++ ++K N + +DEIE+D D + E+L L + Y
Sbjct: 335 RHRLGRQLESLLDELPAHIIDFLKKHNSNGGEIAEDEIEIDIDVLSDEVLVTLRNLLDEY 394
Query: 896 KKSLSKNKRKAE 907
++ + E
Sbjct: 395 IQNKEAKQTNVE 406
>ref|NP_189362.1| DNA-binding bromodomain-containing protein [Arabidopsis thaliana]
Length = 813
Score = 143 bits (361), Expect = 2e-32
Identities = 95/252 (37%), Positives = 123/252 (48%), Gaps = 24/252 (9%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +LL KL H ++WVF PVDV L + DY T I +PMDLGTVK L Y SP E
Sbjct: 178 KQCDTLLRKLWSHPHSWVFQAPVDVVKLNIPDYLTTIKHPMDLGTVKKNLASGVYSSPHE 237
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPL 776
FA DVRLTF NAMTYNP G DVH M + LSK+FE RW I+ P
Sbjct: 238 FAADVRLTFTNAMTYNPPGHDVHIMGDILSKLFEARWKTIKKK------------LPPCS 285
Query: 777 SRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDE 836
+ +PA T P D R+ P + R M + P P KP MT E
Sbjct: 286 MQTLPAVTLEPN-DERKAAISVPPAKK--RKMASPVRESVPEPVKPL--------MTEVE 334
Query: 837 KQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQ-QDDEIEVDFDAVDAEILWELDRFVLNY 895
+ +L L SL E ++ ++K N + +DEIE+D D + E+L L + Y
Sbjct: 335 RHRLGRQLESLLDELPAHIIDFLKKHNSNGGEIAEDEIEIDIDVLSDEVLVTLRNLLDEY 394
Query: 896 KKSLSKNKRKAE 907
++ + E
Sbjct: 395 IQNKEAKQTNVE 406
Score = 36.6 bits (83), Expect = 4.1
Identities = 71/337 (21%), Positives = 123/337 (36%), Gaps = 63/337 (18%)
Query: 193 QLEEENSAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGD---GNSLRQKVEGHNLVQ 249
++E N ++P SS + + ++ +PP S D G+S Q + +VQ
Sbjct: 409 EIELINGSRPSNSSLQRGNEMADEYVDGN---EPPISRSSSDSDSGSSEDQSDDAKPMVQ 465
Query: 250 PQASSIIEDGNSPRQKTEDQ-------------SLAQPPVSSRTDDGDSAQQQVEDQNLA 296
+S + E NS Q+ E+ +L Q + S+ + N+
Sbjct: 466 GDSSKMPETANSEAQRDENTRIDDLFVGSQSTGALEQMDICSQQKLSSDESDGQHEGNIL 525
Query: 297 QTHVSS-----------RTEDV-----NSPRPQNSTQKDGDSPQILENSRPEDASLTQPE 340
+T SS R D+ P PQN + D P+ L R E+ L + +
Sbjct: 526 ETPASSEKRYRAALLKNRFADIILKAREKPLPQNGIKGD---PERLRKER-EELVLQKKK 581
Query: 341 VNSRLEDRSFPQPELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSE 400
+RL+ + ED Q + + E ++ + L E + + +
Sbjct: 582 EKARLQAEA-------EAAEDARRQAEAEAAAEAAAEAKRKRELEREAARQALLKMEKTV 634
Query: 401 DLNLAQPLSQSVSDDLHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSN 460
++N + +DL + + P L E+ P P+ G S NL +E
Sbjct: 635 EIN----ENSRFLEDLEMLSSSAPEQLPSSAEETSPERPLDALG--SFNLRGSNPLEQLG 688
Query: 461 PNVLLEDD--GPSSPIHRHKAVPSTEYRHSENVTEEP 495
+ +DD P +P AVP + E TE P
Sbjct: 689 LYMKQDDDEEEPEAP-----AVPKPD----ETSTERP 716
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.310 0.128 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,716,551,735
Number of Sequences: 2540612
Number of extensions: 80786917
Number of successful extensions: 295304
Number of sequences better than 10.0: 4153
Number of HSP's better than 10.0 without gapping: 1036
Number of HSP's successfully gapped in prelim test: 3306
Number of HSP's that attempted gapping in prelim test: 271112
Number of HSP's gapped (non-prelim): 14061
length of query: 966
length of database: 863,360,394
effective HSP length: 138
effective length of query: 828
effective length of database: 512,755,938
effective search space: 424561916664
effective search space used: 424561916664
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC144893.5


