

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.14 - phase: 0
(113 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
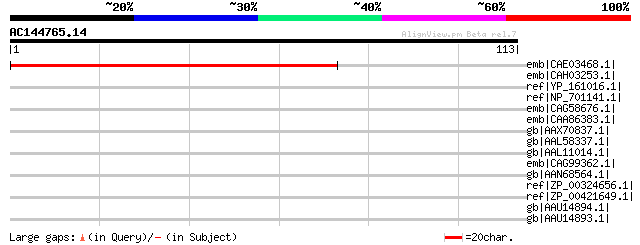
Score E
Sequences producing significant alignments: (bits) Value
emb|CAE03468.1| OSJNBa0083N12.5 [Oryza sativa (japonica cultivar... 76 2e-13
emb|CAH03253.1| WD-40 repeat protein, putative [Paramecium tetra... 36 0.25
ref|YP_161016.1| conserved hypothetical protein, possibly flavin... 33 1.6
ref|NP_701141.1| hypothetical protein PF11_0281 [Plasmodium falc... 32 2.8
emb|CAG58676.1| unnamed protein product [Candida glabrata CBS138... 32 2.8
emb|CAA86383.1| NO364 [Saccharomyces cerevisiae] gi|6324016|ref|... 32 3.7
gb|AAX70837.1| hypothetical protein, conserved [Trypanosoma brucei] 32 4.8
gb|AAL58337.1| TcpA [Vibrio cholerae] 32 4.8
gb|AAL11014.1| TcpA [Vibrio cholerae] 32 4.8
emb|CAG99362.1| unnamed protein product [Kluyveromyces lactis NR... 31 6.3
gb|AAN68564.1| ribosomal protein S6 modification protein-related... 31 6.3
ref|ZP_00324656.1| COG0457: FOG: TPR repeat [Trichodesmium eryth... 31 8.2
ref|ZP_00421649.1| Adenylosuccinate synthetase [Burkholderia vie... 31 8.2
gb|AAU14894.1| chloroplast Flu-like protein short form [Chlamydo... 31 8.2
gb|AAU14893.1| chloroplast Flu-like protein long form [Chlamydom... 31 8.2
>emb|CAE03468.1| OSJNBa0083N12.5 [Oryza sativa (japonica cultivar-group)]
gi|50928447|ref|XP_473751.1| OSJNBa0083N12.5 [Oryza
sativa (japonica cultivar-group)]
Length = 2646
Score = 76.3 bits (186), Expect = 2e-13
Identities = 37/73 (50%), Positives = 52/73 (70%)
Query: 1 MATMCFERAGDSYWGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQ 60
MATMCFE+AGD+Y K ++AA L ATA R+ N E + L++A+EIFE +G + AA
Sbjct: 1621 MATMCFEKAGDAYREKLARAAGLLATADRVISTNFEMGQSSLQKASEIFESIGKHEKAAT 1680
Query: 61 CFTDLGDYERAGI 73
C+ LGDY++AG+
Sbjct: 1681 CYMKLGDYKKAGM 1693
>emb|CAH03253.1| WD-40 repeat protein, putative [Paramecium tetraurelia]
gi|50404892|ref|YP_053984.1| WD-40 repeat protein,
putative [Paramecium tetraurelia]
Length = 1196
Score = 35.8 bits (81), Expect = 0.25
Identities = 16/38 (42%), Positives = 25/38 (65%)
Query: 34 NLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERA 71
+L++ NA+L + E F+ LGM SA +CF GD ++A
Sbjct: 840 SLQEGNALLCDLGERFQLLGMAKSAIKCFEKAGDIKKA 877
>ref|YP_161016.1| conserved hypothetical protein, possibly flavin-dependent
dehydrogenase [Azoarcus sp. EbN1]
gi|56315470|emb|CAI10115.1| conserved hypothetical,
possibly flavin-dependent dehydrogenase [Azoarcus sp.
EbN1]
Length = 368
Score = 33.1 bits (74), Expect = 1.6
Identities = 29/86 (33%), Positives = 35/86 (39%), Gaps = 12/86 (13%)
Query: 16 KKSKAASLRATAIRLHDLNLEDANAILREAAEIFE------------GLGMVDSAAQCFT 63
+ +AA+LR R + D + R A FE G+VDS A F
Sbjct: 95 RAEQAAALRQIRRRAEANGVGDLRELDRAAVAAFEPALDVHAALLSPSTGIVDSHALMFA 154
Query: 64 DLGDYERAGINFNFGIHVYSFMSFSI 89
LGD ER G F FG V S SI
Sbjct: 155 LLGDAERDGAMFAFGSPVLDGYSDSI 180
>ref|NP_701141.1| hypothetical protein PF11_0281 [Plasmodium falciparum 3D7]
gi|23496206|gb|AAN35865.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 247
Score = 32.3 bits (72), Expect = 2.8
Identities = 17/73 (23%), Positives = 33/73 (44%)
Query: 19 KAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGINFNFG 78
K S + + + D+N E+ N ILR + +F G+ + L + + N
Sbjct: 36 KFPSFKPERLHIVDVNTENNNYILRSSIPLFNGVYSEEKLLSYIKKLFEENNLSYSDNLT 95
Query: 79 IHVYSFMSFSIKK 91
+H+ SF+ +K+
Sbjct: 96 LHIISFLRNDLKE 108
>emb|CAG58676.1| unnamed protein product [Candida glabrata CBS138]
gi|50286655|ref|XP_445757.1| unnamed protein product
[Candida glabrata]
Length = 904
Score = 32.3 bits (72), Expect = 2.8
Identities = 14/31 (45%), Positives = 21/31 (67%)
Query: 41 ILREAAEIFEGLGMVDSAAQCFTDLGDYERA 71
+L+ A EI+E LGM AA C+ +GD ++A
Sbjct: 521 VLKSAVEIYERLGMSCEAALCYAAVGDEKKA 551
>emb|CAA86383.1| NO364 [Saccharomyces cerevisiae] gi|6324016|ref|NP_014086.1|
Protein required for cell viability; Ynl313cp
[Saccharomyces cerevisiae] gi|1302419|emb|CAA96243.1|
unnamed protein product [Saccharomyces cerevisiae]
gi|1077429|pir||S51299 hypothetical protein YNL313c -
yeast (Saccharomyces cerevisiae)
gi|1176586|sp|P42842|YN53_YEAST Hypothetical 102.3 kDa
protein in DAL82-RFA2 intergenic region
Length = 904
Score = 32.0 bits (71), Expect = 3.7
Identities = 14/31 (45%), Positives = 20/31 (64%)
Query: 41 ILREAAEIFEGLGMVDSAAQCFTDLGDYERA 71
IL+ A EI+E LGM A C+ +GD ++A
Sbjct: 525 ILKSAVEIYERLGMACETALCYAAVGDEKKA 555
>gb|AAX70837.1| hypothetical protein, conserved [Trypanosoma brucei]
Length = 575
Score = 31.6 bits (70), Expect = 4.8
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 1/71 (1%)
Query: 10 GDSYWGKKSKAASLRATAIRLHDLNLED-ANAILREAAEIFEGLGMVDSAAQCFTDLGDY 68
GD +G+ + SL A +D+ ED A + +A++ + G+V + Q DLGD
Sbjct: 463 GDQLYGRFKRKKSLAAGGSLKNDVTSEDVAEGKDQSSAKVVDAEGLVKAQRQTSMDLGDE 522
Query: 69 ERAGINFNFGI 79
+ A + F +
Sbjct: 523 DCADVASRFRV 533
>gb|AAL58337.1| TcpA [Vibrio cholerae]
Length = 199
Score = 31.6 bits (70), Expect = 4.8
Identities = 16/61 (26%), Positives = 26/61 (42%)
Query: 14 WGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGI 73
W AA +A AI + L ++ ++F + + + AA DLGD+E
Sbjct: 96 WSFPRNAAGNKAFAISVDGLTQAQCKTLVTSVGDMFPYINVQEKAAVALADLGDFETGPA 155
Query: 74 N 74
N
Sbjct: 156 N 156
>gb|AAL11014.1| TcpA [Vibrio cholerae]
Length = 198
Score = 31.6 bits (70), Expect = 4.8
Identities = 16/61 (26%), Positives = 26/61 (42%)
Query: 14 WGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGI 73
W AA +A AI + L ++ ++F + + + AA DLGD+E
Sbjct: 95 WSFPRNAAGNKAFAISVDGLTQAQCKTLVTSVGDMFPYINVQEKAAVALADLGDFETGPA 154
Query: 74 N 74
N
Sbjct: 155 N 155
>emb|CAG99362.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50308545|ref|XP_454275.1| unnamed protein product
[Kluyveromyces lactis]
Length = 896
Score = 31.2 bits (69), Expect = 6.3
Identities = 15/31 (48%), Positives = 20/31 (64%)
Query: 41 ILREAAEIFEGLGMVDSAAQCFTDLGDYERA 71
ILR A EI+E L M AA C+ +GD ++A
Sbjct: 516 ILRSAVEIYERLHMACEAALCYAAVGDEKQA 546
>gb|AAN68564.1| ribosomal protein S6 modification protein-related protein
[Pseudomonas putida KT2440] gi|26989675|ref|NP_745100.1|
ribosomal protein S6 modification protein-related
protein [Pseudomonas putida KT2440]
Length = 526
Score = 31.2 bits (69), Expect = 6.3
Identities = 20/69 (28%), Positives = 34/69 (48%), Gaps = 4/69 (5%)
Query: 33 LNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGINFNFGIHVYSFMSFSIKKH 92
+ + DA A+ A E+FE ++ + C+T+ D+ +N G +Y+ F K H
Sbjct: 370 IKVADAQALASAAQELFEHSVLLLAQEYCYTEY-DWRIGVLN---GEALYACQYFMSKGH 425
Query: 93 YDILEHKYQ 101
+ I HK Q
Sbjct: 426 WQIYNHKAQ 434
>ref|ZP_00324656.1| COG0457: FOG: TPR repeat [Trichodesmium erythraeum IMS101]
Length = 887
Score = 30.8 bits (68), Expect = 8.2
Identities = 21/73 (28%), Positives = 34/73 (45%), Gaps = 1/73 (1%)
Query: 28 IRLHDLNLEDANAILREAAEIFEGLGMVD-SAAQCFTDLGDYERAGINFNFGIHVYSFMS 86
I L + +A ++A E+ L +V + F ++G+ + A I FN I +
Sbjct: 188 ISLKTKDFNEAINYYQKAIELKPDLWIVHYKLGKLFQEIGELDTATIEFNLAIELNPSFI 247
Query: 87 FSIKKHYDILEHK 99
+S K DIL HK
Sbjct: 248 YSYKNLGDILHHK 260
>ref|ZP_00421649.1| Adenylosuccinate synthetase [Burkholderia vietnamiensis G4]
gi|67534886|gb|EAM31624.1| Adenylosuccinate synthetase
[Burkholderia vietnamiensis G4]
Length = 448
Score = 30.8 bits (68), Expect = 8.2
Identities = 14/36 (38%), Positives = 21/36 (57%)
Query: 39 NAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGIN 74
+ I+RE F G G+V S F ++G+ E AG+N
Sbjct: 67 SGIMREGVACFIGNGVVLSPEALFKEIGELEEAGVN 102
>gb|AAU14894.1| chloroplast Flu-like protein short form [Chlamydomonas reinhardtii]
Length = 373
Score = 30.8 bits (68), Expect = 8.2
Identities = 19/62 (30%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 16 KKSKAASLRAT-AIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGIN 74
++ AAS R T R +LE I RE E + A C+TD+G++E+A +
Sbjct: 300 QRGLAASCRITHQYRQAIKHLERVLEISREIGEYTGDADAFGTMADCYTDMGEFEKAAVY 359
Query: 75 FN 76
++
Sbjct: 360 YD 361
>gb|AAU14893.1| chloroplast Flu-like protein long form [Chlamydomonas reinhardtii]
Length = 385
Score = 30.8 bits (68), Expect = 8.2
Identities = 19/62 (30%), Positives = 31/62 (49%), Gaps = 1/62 (1%)
Query: 16 KKSKAASLRAT-AIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGIN 74
++ AAS R T R +LE I RE E + A C+TD+G++E+A +
Sbjct: 312 QRGLAASCRITHQYRQAIKHLERVLEISREIGEYTGDADAFGTMADCYTDMGEFEKAAVY 371
Query: 75 FN 76
++
Sbjct: 372 YD 373
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.325 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 174,522,200
Number of Sequences: 2540612
Number of extensions: 6107104
Number of successful extensions: 17787
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 17775
Number of HSP's gapped (non-prelim): 16
length of query: 113
length of database: 863,360,394
effective HSP length: 89
effective length of query: 24
effective length of database: 637,245,926
effective search space: 15293902224
effective search space used: 15293902224
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144765.14


