

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.1 + phase: 0 /pseudo
(52 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
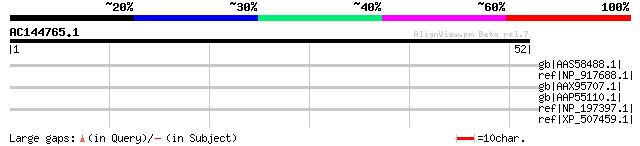
gb|AAS58488.1| putative transposase [Triticum monococcum] 44 0.002
ref|NP_917688.1| P0686E09.10 [Oryza sativa (japonica cultivar-gr... 42 0.003
gb|AAX95707.1| FAR1 family, putative [Oryza sativa (japonica cul... 40 0.013
gb|AAP55110.1| putative transposase [Oryza sativa (japonica cult... 38 0.064
ref|NP_197397.1| far-red impaired responsive protein, putative [... 33 2.7
ref|XP_507459.1| PREDICTED OJ1135_F06.10-1 gene product [Oryza s... 32 4.6
>gb|AAS58488.1| putative transposase [Triticum monococcum]
Length = 751
Score = 43.5 bits (101), Expect = 0.002
Identities = 22/49 (44%), Positives = 29/49 (58%), Gaps = 1/49 (2%)
Query: 2 EEANKFWLAYSFRVGFGVRVRFANKR-EDGSVSSCRMLYEGSVNTWYVK 49
+EA FWL YS + GF VR R+ N+R DG V+SCR + + W K
Sbjct: 7 DEAWAFWLTYSGQKGFEVRKRYTNRRPTDGKVTSCRFVCANEGHRWQDK 55
>ref|NP_917688.1| P0686E09.10 [Oryza sativa (japonica cultivar-group)]
Length = 1066
Score = 42.4 bits (98), Expect = 0.003
Identities = 20/38 (52%), Positives = 28/38 (73%), Gaps = 1/38 (2%)
Query: 2 EEANKFWLAYSFRVGFGVRVRFANK-REDGSVSSCRML 38
+EA FWL+YS + GF VR R+ N+ + DGSV+SCR +
Sbjct: 7 DEAWAFWLSYSGQKGFEVRKRYTNRSKSDGSVTSCRFV 44
>gb|AAX95707.1| FAR1 family, putative [Oryza sativa (japonica cultivar-group)]
Length = 1113
Score = 40.4 bits (93), Expect = 0.013
Identities = 19/37 (51%), Positives = 27/37 (72%), Gaps = 1/37 (2%)
Query: 1 MEEANKFWLAYSFRVGFGVRVRFANKRE-DGSVSSCR 36
++EA FW++Y + GF VR R++NKR+ DG V SCR
Sbjct: 260 VDEAWMFWVSYGGQKGFEVRKRYSNKRKSDGKVRSCR 296
>gb|AAP55110.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|37537042|ref|NP_922823.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|19224996|gb|AAL86472.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|49614722|dbj|BAD26733.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
Length = 486
Score = 38.1 bits (87), Expect = 0.064
Identities = 19/36 (52%), Positives = 24/36 (65%), Gaps = 1/36 (2%)
Query: 2 EEANKFWLAYSFRVGFGVRVRFANKRE-DGSVSSCR 36
+EA FW +Y + GF VR R+ NKR+ DG V SCR
Sbjct: 65 DEAWMFWNSYGGQKGFEVRKRYTNKRKSDGKVRSCR 100
>ref|NP_197397.1| far-red impaired responsive protein, putative [Arabidopsis
thaliana]
Length = 788
Score = 32.7 bits (73), Expect = 2.7
Identities = 18/38 (47%), Positives = 24/38 (62%), Gaps = 1/38 (2%)
Query: 2 EEANKFWLAYSFRVGFGVRV-RFANKREDGSVSSCRML 38
EEA +F+ AY+ R GF VR + R DG+VSS R +
Sbjct: 53 EEAREFYNAYAARTGFKVRTGQLYRSRTDGTVSSRRFV 90
>ref|XP_507459.1| PREDICTED OJ1135_F06.10-1 gene product [Oryza sativa (japonica
cultivar-group)] gi|51963940|ref|XP_506753.1| PREDICTED
OJ1135_F06.10-1 gene product [Oryza sativa (japonica
cultivar-group)]
Length = 721
Score = 32.0 bits (71), Expect = 4.6
Identities = 15/32 (46%), Positives = 20/32 (61%)
Query: 3 EANKFWLAYSFRVGFGVRVRFANKREDGSVSS 34
EA F+ +Y+ VGF +R AN R DGS+ S
Sbjct: 85 EAYDFYNSYARNVGFSIRKNHANTRADGSLCS 116
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.134 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 86,398,940
Number of Sequences: 2540612
Number of extensions: 2199719
Number of successful extensions: 3772
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 3766
Number of HSP's gapped (non-prelim): 7
length of query: 52
length of database: 863,360,394
effective HSP length: 28
effective length of query: 24
effective length of database: 792,223,258
effective search space: 19013358192
effective search space used: 19013358192
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 69 (31.2 bits)
Medicago: description of AC144765.1


