

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
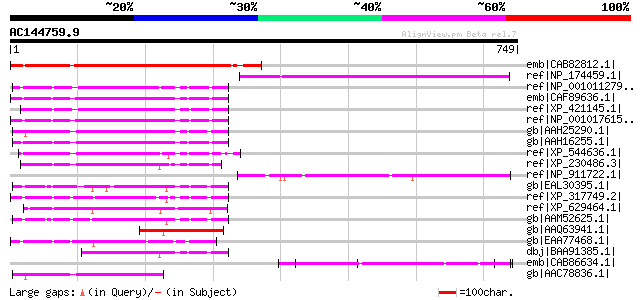
Query= AC144759.9 - phase: 0
(749 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB82812.1| phospholipase-like protein [Arabidopsis thaliana... 452 e-125
ref|NP_174459.1| pentatricopeptide (PPR) repeat-containing prote... 320 8e-86
ref|NP_001011279.1| hypothetical protein LOC496732 [Xenopus trop... 241 6e-62
emb|CAF89636.1| unnamed protein product [Tetraodon nigroviridis] 241 6e-62
ref|XP_421145.1| PREDICTED: similar to PLA2G4B protein [Gallus g... 239 3e-61
ref|NP_001017615.1| hypothetical protein LOC550278 [Danio rerio]... 236 2e-60
gb|AAH25290.1| PLA2G4B protein [Homo sapiens] 232 3e-59
gb|AAH16255.1| Pla2g4b protein [Mus musculus] 229 3e-58
ref|XP_544636.1| PREDICTED: similar to phospholipase A2, group I... 212 4e-53
ref|XP_230486.3| PREDICTED: similar to phospholipase A2, group I... 209 3e-52
ref|NP_911722.1| putative pentatricopeptide repeat-containing pr... 191 1e-46
gb|EAL30395.1| GA10099-PA [Drosophila pseudoobscura] 181 6e-44
ref|XP_317749.2| ENSANGP00000010328 [Anopheles gambiae str. PEST... 181 8e-44
ref|XP_629464.1| hypothetical protein DDB0184556 [Dictyostelium ... 181 8e-44
gb|AAM52625.1| GH14974p [Drosophila melanogaster] gi|7294469|gb|... 178 5e-43
gb|AAQ63941.1| phospholipase-like protein [Brachypodium sylvaticum] 171 8e-41
gb|EAA77468.1| hypothetical protein FG07451.1 [Gibberella zeae P... 165 4e-39
dbj|BAA91385.1| unnamed protein product [Homo sapiens] 162 5e-38
emb|CAB86634.1| putative protein [Arabidopsis thaliana] gi|15240... 157 9e-37
gb|AAC78836.1| cytosolic phospholipase A2 beta [Homo sapiens] gi... 154 8e-36
>emb|CAB82812.1| phospholipase-like protein [Arabidopsis thaliana]
gi|15231275|ref|NP_190174.1| hypothetical protein
[Arabidopsis thaliana] gi|11281171|pir||T47528
hypothetical protein T31B5.20 - Arabidopsis thaliana
Length = 431
Score = 452 bits (1163), Expect = e-125
Identities = 219/371 (59%), Positives = 271/371 (73%), Gaps = 16/371 (4%)
Query: 1 MNEKIEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSL 60
M ++IE LWREVRELSLG+K I+R +S P+P++F RN+++ +KPC+IS +I+HWP+L L
Sbjct: 1 MAKEIENLWREVRELSLGTK--IDRFDSQPSPVKFLRNYVSQSKPCVISKAITHWPALKL 58
Query: 61 WSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINS 120
WS P+YLT +LS VSLHLTP G AD++T S LCFASAHV+ + FPEAL+++ S
Sbjct: 59 WSDPAYLTGALSDDVVSLHLTPNGCADAVT----GDSDLCFASAHVEKVLFPEALKVVQS 114
Query: 121 SNPSQCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFH 180
S V Y QQQNDCFR+EY ++ DCD I WATEAFG PEAVNLWIG S T FH
Sbjct: 115 SCKGLKVGYLQQQNDCFRTEYSTVALDCDGDIEWATEAFGCSPEAVNLWIGTDDSVTSFH 174
Query: 181 KDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVPW 240
KDHYENLYAVV+G+KHFLL PPTDVHR YI YPAA Y Y+ +T F LE+++P R+VPW
Sbjct: 175 KDHYENLYAVVSGEKHFLLLPPTDVHRLYIEQYPAANYSYHRDTDAFKLEVEEPVRHVPW 234
Query: 241 CSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTI 300
SV+P+PSPE E KFPL+F+GP PF CTVKAGE+LYLPSMWFHHV Q+ DG TI
Sbjct: 235 SSVDPYPSPEKEASERLKFPLFFDGPKPFHCTVKAGEVLYLPSMWFHHVSQTPGDGGYTI 294
Query: 301 AVNYWYDMQFDIKYAYFNFLQSIDYRSPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPS 360
AVNYWYDMQFDIKYAYFNFLQS+ Y+S S + P++ + R + D+ S+ + S
Sbjct: 295 AVNYWYDMQFDIKYAYFNFLQSLLYKS--SSLNPVL--------SWREDEDSESSDAEGS 344
Query: 361 NHQPHLLRLPL 371
P L L
Sbjct: 345 KFTPSATNLYL 355
Score = 36.2 bits (82), Expect = 4.0
Identities = 17/39 (43%), Positives = 27/39 (68%)
Query: 648 EQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVF 686
++VH ++IKLG +S+ +Q LI MY + L+ +AE VF
Sbjct: 368 KRVHVNSIKLGFESNGLIQSGLIDMYRKYRLVINAEKVF 406
>ref|NP_174459.1| pentatricopeptide (PPR) repeat-containing protein [Arabidopsis
thaliana] gi|25513601|pir||E86441 46.3K hypothetical
protein F5M6.20 - Arabidopsis thaliana
gi|12321298|gb|AAG50719.1| hypothetical protein
[Arabidopsis thaliana]
Length = 409
Score = 320 bits (821), Expect = 8e-86
Identities = 176/402 (43%), Positives = 241/402 (59%), Gaps = 6/402 (1%)
Query: 340 PPAPPTRRRNADTTSTTSPPSNHQPHLLRLPL-RRNPKPKNLSLIHPSSQPITPPKKSKR 398
PP+PP+ + + ST N + L R PK + + QP P+
Sbjct: 9 PPSPPSLVPSFNYNSTARSVGNDVRTNFDVQLFLRKPKHQKSEPVVVIQQPQIQPQNPSS 68
Query: 399 RRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLTL 458
R C +TS IL LMD+L P DIY+ L KE D A EL I+ I +T
Sbjct: 69 R--C-STSDILRLMDSLSLPGNEDIYSCLAKESARENDQRGAHELQVHIMKSSIRPTITF 125
Query: 459 LNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLD 518
+NR+L+M VSCG L+ R++FD M RDFHSWA +F+ E G+YE+A +FVSML
Sbjct: 126 INRLLLMHVSCGRLDITRQMFDRMPHRDFHSWAIVFLGCIEMGDYEDAAFLFVSMLKHSQ 185
Query: 519 VMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHV--LISSSLIRFYGRFKC 576
F P WI C+LKACA + LG QVH KLG D +S SLIRFYG F+C
Sbjct: 186 KGAFKIPSWILGCVLKACAMIRDFELGKQVHALCHKLGFIDEEDSYLSGSLIRFYGEFRC 245
Query: 577 LEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKA 636
LEDAN+V +++S NT+ W AK+ + RE F E + DF +MG G+KK+ FS+VLKA
Sbjct: 246 LEDANLVLHQLSNANTVAWAAKVTNDYREGEFQEVIRDFIEMGNHGIKKNVSVFSNVLKA 305
Query: 637 CGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVD 696
C + + G G+QVHA+AIKLG +SD ++C LI MYG+ G ++DAE VF+ +++E +V
Sbjct: 306 CSWVSDGGRSGQQVHANAIKLGFESDCLIRCRLIEMYGKYGKVKDAEKVFKSSKDETSVS 365
Query: 697 SLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRI 738
NAM+ Y+QNG+YIEA+K +YQMKA G++ H+ LL + +
Sbjct: 366 CWNAMVASYMQNGIYIEAIKLLYQMKATGIKAHDTLLNEAHL 407
>ref|NP_001011279.1| hypothetical protein LOC496732 [Xenopus tropicalis]
gi|56789252|gb|AAH87993.1| Hypothetical LOC496732
[Xenopus tropicalis]
Length = 313
Score = 241 bits (615), Expect = 6e-62
Identities = 127/319 (39%), Positives = 183/319 (56%), Gaps = 22/319 (6%)
Query: 5 IEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHP 64
+E EVREL S+ L++ P+PLQFHR++++PN+PCII N+ +HWP+L W+
Sbjct: 12 LESFSEEVRELH--GTDSVPYLDAPPSPLQFHRDWVSPNRPCIIRNAFTHWPALHKWTF- 68
Query: 65 SYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSSNPS 124
YL + S VS+ +TP G AD++ F + + + L ++ + +
Sbjct: 69 GYLRTHIGSKKVSVAVTPNGYADAVYKNR-------FVMPEERTMFLSDFLDIVEKKSNT 121
Query: 125 QCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFHKDHY 184
V Y Q+Q E+ +V+D + HI W +E G P+AVN W+G + T HKDHY
Sbjct: 122 PGVFYIQKQCSNLTEEFPELVEDVENHIPWMSETLGKSPDAVNFWLGESAAITSLHKDHY 181
Query: 185 ENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVPWCSVN 244
ENLY V++G+KHF+L PP+D + ATY Y E G F + + VPW ++
Sbjct: 182 ENLYCVISGEKHFILHPPSDRPFIPYEMFQPATYHVY-EDGSFKVVDHESAEKVPWIPLD 240
Query: 245 PFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTIAVNY 304
P LE ++ ++P Y P CTV+AGE+LYLPS+WFHHVRQS IAVN+
Sbjct: 241 P------LEPDLIRYPSY-KQTKPLHCTVRAGEMLYLPSLWFHHVRQSHG----CIAVNF 289
Query: 305 WYDMQFDIKYAYFNFLQSI 323
WYDM++DI+Y YF L SI
Sbjct: 290 WYDMEYDIRYHYFQLLDSI 308
>emb|CAF89636.1| unnamed protein product [Tetraodon nigroviridis]
Length = 308
Score = 241 bits (615), Expect = 6e-62
Identities = 131/324 (40%), Positives = 188/324 (57%), Gaps = 24/324 (7%)
Query: 1 MNEKIEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSL 60
+ +++ E E +L L RS+ LE AP PL+F+R+++ PN+PCII N++SHW +LS
Sbjct: 4 VKKRVTEFSVEAHDLYLN--RSVPHLEGAPDPLEFYRSWVAPNRPCIIRNALSHWAALSS 61
Query: 61 WSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINS 120
WS P+YL Q + S +S+ +TP G AD++ S F + + L ++
Sbjct: 62 WS-PAYLRQKVGSKVISVAVTPNGYADAV-------SGQHFVMPEERQMSLASVLDVMEG 113
Query: 121 SNPSQ-CVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWF 179
PS+ V Y Q+Q E +V D D HI+W + A G P+AVN W+G + T
Sbjct: 114 KEPSERAVFYVQKQCSNLLEELPELVGDVDPHISWMSAALGRLPDAVNFWLGEAGAVTSM 173
Query: 180 HKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVP 239
HKDHYENLY VV+G+KHF+L PPTD Y A Y+ + G F++ +++ VP
Sbjct: 174 HKDHYENLYCVVSGEKHFVLLPPTDRPFVPYGLYQPAVYR-QRDDGHFEV-VEQRGPKVP 231
Query: 240 WCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELT 299
W ++P L ++ K+P Y P C+VKAGE+LYLPS+WFHHV+QS
Sbjct: 232 WIPLDP------LNPDLEKYPQY-RRAQPLRCSVKAGEMLYLPSLWFHHVQQSHG----C 280
Query: 300 IAVNYWYDMQFDIKYAYFNFLQSI 323
AVN+WYDM +DIKY YF L+S+
Sbjct: 281 TAVNFWYDMDYDIKYNYFRLLESL 304
>ref|XP_421145.1| PREDICTED: similar to PLA2G4B protein [Gallus gallus]
Length = 427
Score = 239 bits (609), Expect = 3e-61
Identities = 124/307 (40%), Positives = 177/307 (57%), Gaps = 20/307 (6%)
Query: 17 LGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHPSYLTQSLSSTTV 76
LG S+ L+ P+PL+F+R +++PNKPCII N+I HWP+L W+ +YL + + V
Sbjct: 134 LGCVESVPYLDRPPSPLEFYREWVSPNKPCIIRNAIGHWPALRKWTL-AYLREVVGHKVV 192
Query: 77 SLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSSNPSQCVAYAQQQNDC 136
S+ +TP G AD++ F + +PF + L ++ S V Y Q+Q
Sbjct: 193 SVAVTPNGYADAVFHDR-------FVMPEERQMPFMDFLDIVEKKVTSPSVFYVQKQCSN 245
Query: 137 FRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFHKDHYENLYAVVTGQKH 196
E+ ++ D I W +EA G +P+AVN W+G + T HKDHYENLY V++G+K
Sbjct: 246 LTEEFPELICDVQPDIPWMSEALGKKPDAVNFWLGESAAVTSLHKDHYENLYCVISGEKR 305
Query: 197 FLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVPWCSVNPFPSPENLEDEI 256
FLL PP+D Y AATYK E G F + +K VPW ++P L +
Sbjct: 306 FLLHPPSDRPFIPYELYQAATYK-VSEDGSFGIVDEKTAEKVPWIPLDP------LNPNL 358
Query: 257 SKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTIAVNYWYDMQFDIKYAY 316
++P Y P +CTVKAGE+LYLPS+WFHHV+QS IAVNYWYDM++D+KY+Y
Sbjct: 359 ERYPEYAQA-KPLQCTVKAGEMLYLPSLWFHHVQQSHG----CIAVNYWYDMEYDLKYSY 413
Query: 317 FNFLQSI 323
+ L +
Sbjct: 414 YQLLDCL 420
>ref|NP_001017615.1| hypothetical protein LOC550278 [Danio rerio]
gi|62202217|gb|AAH92834.1| Hypothetical protein
LOC550278 [Danio rerio]
Length = 311
Score = 236 bits (603), Expect = 2e-60
Identities = 126/323 (39%), Positives = 186/323 (57%), Gaps = 22/323 (6%)
Query: 1 MNEKIEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSL 60
+ E + + +E REL L ++ L+ +PLQF+R++I PNKPCII N+ + WP+LS
Sbjct: 4 VKECLRDFPKEARELYLND--AVPYLDEPLSPLQFYRDWIGPNKPCIIRNAFNDWPALSK 61
Query: 61 WSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINS 120
W+ P+YL + + S +S+ +TP G AD++ + F + + F L ++
Sbjct: 62 WN-PTYLREKVGSKVISVAVTPNGFADAV-------NGNRFVMPEERQMSFSSLLDIVEG 113
Query: 121 SNPSQCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFH 180
S V Y Q+Q E + D HI W +EA G P+AVN W+G + + T H
Sbjct: 114 KIKSSAVFYVQKQCSNLMEEIPELTGDVQTHIPWMSEALGKLPDAVNFWLGEESAVTSMH 173
Query: 181 KDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVPW 240
KDHYENLY V++GQK F+L PPTD Y ATY+ + G F++ ++ + VPW
Sbjct: 174 KDHYENLYCVISGQKEFILLPPTDRPFIPYELYQPATYR-QKDDGTFEIVDEENSPKVPW 232
Query: 241 CSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTI 300
++P L+ + +FP Y + CTVKAGE+LYLPS+WFHHVRQS I
Sbjct: 233 IPLDP------LKPDFERFPSYRHA-KALHCTVKAGEMLYLPSLWFHHVRQSHG----CI 281
Query: 301 AVNYWYDMQFDIKYAYFNFLQSI 323
AVN+WYDM++DIKY YF ++++
Sbjct: 282 AVNFWYDMEYDIKYNYFQLVEAL 304
>gb|AAH25290.1| PLA2G4B protein [Homo sapiens]
Length = 316
Score = 232 bits (592), Expect = 3e-59
Identities = 124/324 (38%), Positives = 185/324 (56%), Gaps = 25/324 (7%)
Query: 5 IEELWREVRELSLGSKR-----SIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLS 59
+E + E+RE ++ ++ L+ PTPL F+R+++ PN+PCII N++ HWP+L
Sbjct: 6 LEAVRSELREFPAAARELCVPLAVPYLDKPPTPLHFYRDWVCPNRPCIIRNALQHWPALQ 65
Query: 60 LWSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLIN 119
WS P Y ++ ST VS+ +TP G AD++ F + LP L ++
Sbjct: 66 KWSLP-YFRATVGSTEVSVAVTPDGYADAVR-------GDRFMMPAERRLPLSFVLDVLE 117
Query: 120 SSNPSQCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWF 179
V Y Q+Q SE ++ D + H+ WA+EA G P+AVN W+G + T
Sbjct: 118 GRAQHPGVLYVQKQCSNLPSELPQLLPDLESHVPWASEALGKMPDAVNFWLGEAAAVTSL 177
Query: 180 HKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVP 239
HKDHYENLY VV+G+KHFL PP+D Y ATY+ E G F + ++ VP
Sbjct: 178 HKDHYENLYCVVSGEKHFLFHPPSDRPFIPYELYTPATYQ-LTEEGTFKVVDEEAMEKVP 236
Query: 240 WCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELT 299
W ++P L +++++P Y + CTV+AGE+LYLP++WFHHV+QS +
Sbjct: 237 WIPLDP------LAPDLARYPSY-SQAQALRCTVRAGEMLYLPALWFHHVQQS----QGC 285
Query: 300 IAVNYWYDMQFDIKYAYFNFLQSI 323
IAVN+WYDM++D+KY+YF L S+
Sbjct: 286 IAVNFWYDMEYDLKYSYFQLLDSL 309
>gb|AAH16255.1| Pla2g4b protein [Mus musculus]
Length = 316
Score = 229 bits (583), Expect = 3e-58
Identities = 121/319 (37%), Positives = 186/319 (57%), Gaps = 22/319 (6%)
Query: 5 IEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHP 64
++E R+L++ R + L+ P+PL F+R+++ PN+PCII N++ HWP+L WS
Sbjct: 13 LQEFPAAARDLNV--PRVVPYLDEPPSPLCFYRDWVCPNRPCIIRNALQHWPALQKWSL- 69
Query: 65 SYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSSNPS 124
SYL ++ ST VS+ +TP G AD++ F + LP L ++
Sbjct: 70 SYLRATVGSTEVSVAVTPDGYADAVR-------GDRFVMPAERRLPISHVLDVLEGRAQH 122
Query: 125 QCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFHKDHY 184
V Y Q+Q +E ++ D + H+ WA+E+ G P+AVN W+G+ + T HKDHY
Sbjct: 123 PGVLYVQKQCSNLPTELPQLLSDIESHVPWASESLGKMPDAVNFWLGDASAVTSLHKDHY 182
Query: 185 ENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPTRYVPWCSVN 244
ENLY VV+G+KHFLL PP+D Y ATY+ E G F + ++ VPW ++
Sbjct: 183 ENLYCVVSGEKHFLLHPPSDRPFIPYNLYTPATYQ-LTEEGTFRVVDEEAMEKVPWIPLD 241
Query: 245 PFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTIAVNY 304
P L +++++P Y + CTV+AGE+LYLP++WFHHV+QS IAVN+
Sbjct: 242 P------LAPDLTQYPSY-SQAQALHCTVRAGEMLYLPALWFHHVQQSHG----CIAVNF 290
Query: 305 WYDMQFDIKYAYFNFLQSI 323
WYDM++D+KY+YF + ++
Sbjct: 291 WYDMEYDLKYSYFQLMDTL 309
>ref|XP_544636.1| PREDICTED: similar to phospholipase A2, group IVB [Canis
familiaris]
Length = 1082
Score = 212 bits (539), Expect = 4e-53
Identities = 127/337 (37%), Positives = 183/337 (53%), Gaps = 42/337 (12%)
Query: 13 RELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHPSYLTQSLS 72
RELS+ ++ L+ PTPL F+R+++ PN+PCII N++ HWP+L WS P YL ++
Sbjct: 11 RELSV--PLAVPYLDEPPTPLHFYRDWVCPNRPCIIRNALQHWPALQKWSFP-YLRATVG 67
Query: 73 STTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSSNPSQCVAYAQQ 132
ST VS+ +TP G AD++ F + LP L ++ V Y Q+
Sbjct: 68 STEVSVAVTPDGYADAVR-------GNRFVMPAERRLPLSCVLDVLEGQAQHPGVLYVQK 120
Query: 133 QNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFHKDHYENLYAVVT 192
Q +E ++ D + H+ WA+EA G P+AVN W+G + T HKDHYENLY VV+
Sbjct: 121 QCSNLPTELPQLLSDLEPHVPWASEALGKMPDAVNFWLGEAAAVTSLHKDHYENLYCVVS 180
Query: 193 GQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDK---------PTRYVPWCSV 243
G+K FLL PP+D Y ATY+ E G F + ++ P VPW +
Sbjct: 181 GEKRFLLHPPSDRPFIPYELYTPATYQLTQE-GSFKMVDEEAMEKVSASFPGPGVPWIPL 239
Query: 244 NPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTIAVN 303
+P L +++++P Y + CTV+AGE+LYLP++WFHHV+QS IAVN
Sbjct: 240 DP------LAPDLARYPNY-SQARALCCTVQAGEMLYLPALWFHHVQQSHG----CIAVN 288
Query: 304 YWYDMQFDIKYAYFNFLQSIDYRSPTSPMMPIVAPPP 340
YWYDM++D+ N LQ P+ P+ A P
Sbjct: 289 YWYDMEYDL-----NSLQQ------ACPLPPLQAKVP 314
>ref|XP_230486.3| PREDICTED: similar to phospholipase A2, group IVB [Rattus
norvegicus]
Length = 1186
Score = 209 bits (532), Expect = 3e-52
Identities = 117/317 (36%), Positives = 174/317 (53%), Gaps = 40/317 (12%)
Query: 17 LGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHPSYLTQSLSSTTV 76
L R + L+ P+PL F+R+++ PN+PCII N++ HWP+L WS SYL ++ ST V
Sbjct: 56 LNVPRVVPYLDEPPSPLCFYRDWVCPNRPCIIRNALQHWPALQKWSF-SYLRATVGSTEV 114
Query: 77 SLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSSNPSQCVAYAQQQNDC 136
S+ +TP G AD++ F + LP L ++ V Y Q+Q
Sbjct: 115 SVAVTPDGYADAVR-------GDRFVMPAERRLPVSHVLDVLEGQAQHPGVLYVQKQCSN 167
Query: 137 FRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWFHKDHYENLYAVVTGQKH 196
+E ++ D + H+ WA+E+ G P+AVN W+G+ + T HKDHYENLY VV+G+KH
Sbjct: 168 LPTELPQLLSDIESHVPWASESLGKMPDAVNFWLGDAAAVTSLHKDHYENLYCVVSGEKH 227
Query: 197 FLLFPPTDVHRFYIRNYPAATYK--------------------YYMETGEFDLELDKPTR 236
FLL PP+D Y ATY+ ++ +GE+ TR
Sbjct: 228 FLLHPPSDRPFIPYNLYTPATYQLTEEGTFRVVDEEAMEKVPVLFLGSGEWADPQGIGTR 287
Query: 237 YVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDG 296
VPW ++P L +++++P Y + CTV+AGE+LYLP++WFHHV+QS
Sbjct: 288 -VPWIPLDP------LAPDLARYPSY-SQARALHCTVRAGELLYLPALWFHHVQQSHG-- 337
Query: 297 ELTIAVNYWYDMQFDIK 313
IAVN+WYDM++D+K
Sbjct: 338 --CIAVNFWYDMEYDLK 352
>ref|NP_911722.1| putative pentatricopeptide repeat-containing protein [Oryza sativa
(japonica cultivar-group)] gi|24417179|dbj|BAC22540.1|
putative pentatricopeptide repeat-containing protein
[Oryza sativa (japonica cultivar-group)]
gi|50508329|dbj|BAD30147.1| putative pentatricopeptide
repeat-containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 435
Score = 191 bits (484), Expect = 1e-46
Identities = 136/426 (31%), Positives = 206/426 (47%), Gaps = 31/426 (7%)
Query: 337 APPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPPKKS 396
A P +P RRR + T+ LR P + P P +HP P S
Sbjct: 17 AAPRLSPRRRRRRLLAANATTARGALPLPALRKPTKPPPPPP----LHPRPSLPVPTTSS 72
Query: 397 KR----RRKCDT----------TSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIE 442
RRK T +L L+DAL P D+Y SL+++C D
Sbjct: 73 DDDGDIRRKPATGATASLCSSGAGDVLRLLDALRLPPDEDVYVSLLRDCA---DAAEVAS 129
Query: 443 LHTQIITRGIE--LPLTLLNRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYEN 500
+H I + LPL L NR+++ + +CG + AR+VFD M V++ +WAT+ +Y +
Sbjct: 130 VHAHIAGKFAVSGLPLPLANRLVLAYAACGDIGAARQVFDEMPVKNGITWATMVSAYSDG 189
Query: 501 GEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKL-GACD 559
+ +A+ +FV M Q+ + +L++CA + G QVH ++K G C
Sbjct: 190 CFHHDALQLFVQMCHQVRGITGDHYTHAIVAVLRSCARVNELQFGEQVHAFVVKKNGVCG 249
Query: 560 HVLISSSLIRFYGRFKCLEDANMVFN--RVSRHNTL---TWTAKIVSSCRERHFSEALGD 614
V SSL++ Y L A V R S + WT+ I + R+ +A+
Sbjct: 250 DV--GSSLLQLYCDSGQLSSARHVLEMMRFSCQEPVPEAAWTSLITAYHRDGILDDAIDV 307
Query: 615 FKKMGRVGVKKDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYG 674
F+ M G+ + SF+ SS+L C +N+G G+QVHADAIK GLD + +V L+ MY
Sbjct: 308 FRGMASSGIARSSFSLSSILAVCAEAKNKGCYGQQVHADAIKRGLDMNQFVGSGLLHMYA 367
Query: 675 RSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLE 734
+ G L DA FE + + NAM M Y + G+Y EA + VYQMKAAG+ P + +
Sbjct: 368 KEGQLADAARAFEAIDGKPDAVCWNAMAMAYARGGMYREATRVVYQMKAAGMNPSKLTMN 427
Query: 735 KLRIAC 740
++++AC
Sbjct: 428 EVKLAC 433
>gb|EAL30395.1| GA10099-PA [Drosophila pseudoobscura]
Length = 314
Score = 181 bits (460), Expect = 6e-44
Identities = 110/332 (33%), Positives = 173/332 (51%), Gaps = 42/332 (12%)
Query: 5 IEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHP 64
I L +E +L +GS SI L+ PT L+F R++ N P II N++S WP++ W+ P
Sbjct: 8 INLLLQEAEDLRIGS--SISELDHLPTALEFCRDYFAKNSPVIIRNALS-WPAIGKWT-P 63
Query: 65 SYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINS---- 120
YL + L+ V + +TP G AD L + LP + ++L +
Sbjct: 64 DYLIKKLNDKIVDVAVTPNGYADGLATQKGREYFV---------LPLEKQMKLSDLVQRL 114
Query: 121 SNPSQCVAYAQQQNDCFRSEY-----DSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHS 175
+P + Y Q+QN F ++ D ++ D D +A ++F P+AVN W+G++ +
Sbjct: 115 DDPMGAIHYVQKQNSNFSQDFPELGSDLVISDLD----FAQQSFNKPPDAVNFWLGDERA 170
Query: 176 STWFHKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDL----EL 231
T HKD YEN+Y V++G K F+L PP + YP YK ++G+F + +
Sbjct: 171 ITSMHKDPYENMYCVISGYKDFILIPPYQLSCVPRSTYPTGIYK-TSDSGQFYIDPLTDE 229
Query: 232 DKPTRYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQ 291
D W S++P L +++K+P Y P + V AG++LYLP+ WFHHVRQ
Sbjct: 230 DGVELLTEWVSIDP------LAPDLAKYPEYARA-KPLKVRVHAGDVLYLPNYWFHHVRQ 282
Query: 292 SGDDGELTIAVNYWYDMQFDIKYAYFNFLQSI 323
S IAVN+WYD+ +D +Y Y+ L+ +
Sbjct: 283 S----HKCIAVNFWYDLDYDSRYCYYRMLEEL 310
>ref|XP_317749.2| ENSANGP00000010328 [Anopheles gambiae str. PEST]
gi|55237953|gb|EAA12162.2| ENSANGP00000010328 [Anopheles
gambiae str. PEST]
Length = 315
Score = 181 bits (459), Expect = 8e-44
Identities = 101/331 (30%), Positives = 176/331 (52%), Gaps = 30/331 (9%)
Query: 1 MNEKIEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSL 60
+ + + L E ++L L S +I P+ L+F R+++ N P I+ N+++ WP++
Sbjct: 3 LRDAFQFLTSEAKDLFLPS--NIPETYGIPSSLEFVRDYVAKNLPLIMRNAVNDWPAVDK 60
Query: 61 WSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINS 120
W+ Y ++ V++ +TP G AD L F Q + + L ++
Sbjct: 61 WNS-KYFRDTIPDKEVTVAITPNGYADGLA---FHEDEEYFVLPLEQTMRMEDFLSALDH 116
Query: 121 SNPSQCVAYAQQQNDCFRSEYDSIVKDCDQH-IAWATEAFGLEPEAVNLWIGNKHSSTWF 179
+P + Y Q+QN ++ + D ++ + +A+EAF +P+A+N W+G++ + T
Sbjct: 117 KDPD-VIPYIQRQNSNLTEDFQELWIDVNESSLDFASEAFNKQPDAINFWMGDERAITSM 175
Query: 180 HKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDL-------ELD 232
HKD YEN+Y V++G K F+L PP D+H R YP YM+ + + E+
Sbjct: 176 HKDPYENIYCVISGYKDFILIPPIDLHNVPRRQYPMG---IYMQENDDSIVIEPILDEIG 232
Query: 233 KPTRYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQS 292
KP R + W V+P L+ ++ +FP Y + +E + AG++LYLPS+W+HHVRQS
Sbjct: 233 KP-RMIEWVGVDP------LQPDLERFPCYADA-TTYEIRLNAGDLLYLPSLWYHHVRQS 284
Query: 293 GDDGELTIAVNYWYDMQFDIKYAYFNFLQSI 323
IAVN+WYDM +D +Y ++ ++ +
Sbjct: 285 ----HKCIAVNFWYDMDYDARYCFYKMMEKL 311
>ref|XP_629464.1| hypothetical protein DDB0184556 [Dictyostelium discoideum]
gi|60462860|gb|EAL61059.1| hypothetical protein
DDB0184556 [Dictyostelium discoideum]
Length = 388
Score = 181 bits (459), Expect = 8e-44
Identities = 107/316 (33%), Positives = 166/316 (51%), Gaps = 20/316 (6%)
Query: 21 RSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHPSYLTQSLSSTTVSLHL 80
+ IER+E PT L+F+R +++ NKP II+ + +W + WS YL + V++ +
Sbjct: 63 KDIERIEK-PTALEFYREYVSQNKPVIITGLLENWKAYKEWSD-DYLENVMKDVEVTVSI 120
Query: 81 TPTGSADSLTPLPSSP--SSLCFASAHVQNLPFPEALRLINSS---NPSQCVAYAQQQND 135
T G AD++ P+ + S F + + F E ++ S N ++ Y Q QN+
Sbjct: 121 TNDGLADAVKPINENDPKSERVFCKPFEKKIKFQEYIKHSKKSSKENKNKLAYYIQYQNN 180
Query: 136 CFRSEYDSIVKDCDQHIA-WATEAFGLEPEAVNLWIGNKHSSTWFHKDHYENLYAVVTGQ 194
EYD ++ D D+ + +A EAFG +A N W+G S + H+D YEN+Y VV G
Sbjct: 181 SLNVEYDKLLNDIDESVIDFAKEAFGSNIDATNFWMGQDKSVSSLHQDPYENMYCVVRGT 240
Query: 195 KHFLLFPPTDVHRFYIRNYPAATYKYY-----METGEFDLELDK-PTRYVPWCSVNPFPS 248
K F L PP D Y +P+A++ E + ++++D P +PW V+P
Sbjct: 241 KIFTLLPPIDYPFLYKSEFPSASFVNVGCDDNDENIKLEIQIDNDPKMNIPWIPVDP--- 297
Query: 249 PENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGD---DGELTIAVNYW 305
E LE+ I P V+AGE+LYLPS++FH V Q + + TIA+NYW
Sbjct: 298 TETLENNIKLGYPLIERAHPITIRVEAGEVLYLPSLYFHRVAQESNKTSNSLSTIAINYW 357
Query: 306 YDMQFDIKYAYFNFLQ 321
+DM++ I Y YF FL+
Sbjct: 358 FDMKYGINYVYFQFLK 373
>gb|AAM52625.1| GH14974p [Drosophila melanogaster] gi|7294469|gb|AAF49813.1|
CG10133-PA [Drosophila melanogaster]
gi|24663831|ref|NP_648651.1| CG10133-PA [Drosophila
melanogaster]
Length = 316
Score = 178 bits (452), Expect = 5e-43
Identities = 106/324 (32%), Positives = 173/324 (52%), Gaps = 26/324 (8%)
Query: 5 IEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLWSHP 64
++ L +E EL +GS S+ L+ PT L+F R F + N+P +I +++ WP++ W+ P
Sbjct: 8 LDVLLQEAEELCIGS--SVVELDRIPTALEFCREFYSKNQPVVIRKALN-WPAIGKWT-P 63
Query: 65 SYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSSNPS 124
YL ++L +V + +TP G AD L + F + E +R ++ +P+
Sbjct: 64 KYLIEALGDRSVDVAITPNGYADGLA---TQNGQEYFVLPLETKMKLSEVVRRLD--DPT 118
Query: 125 QCVAYAQQQNDCFRSEYDSIVKDCD-QHIAWATEAFGLEPEAVNLWIGNKHSSTWFHKDH 183
V Y Q+QN + + D + +A ++F P+AVN W+G++ + T HKD
Sbjct: 119 GAVHYIQKQNSNLSVDLPELAADLRVSDLDFAQQSFNKPPDAVNFWLGDERAVTSMHKDP 178
Query: 184 YENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLE----LDKPTRYVP 239
YEN+Y V++G K F+L PP + YP YK ++G+F +E + ++
Sbjct: 179 YENVYCVISGHKDFVLIPPHQLSCVPRGIYPTGVYK-TSDSGQFYIEPLRDEEGSDQFTE 237
Query: 240 WCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELT 299
W SV+P L +++K+P Y P + V AG+ILYLP+ WFHHV QS
Sbjct: 238 WVSVDP------LSPDLAKYPEYARA-KPLKVRVHAGDILYLPNYWFHHVSQS----HKC 286
Query: 300 IAVNYWYDMQFDIKYAYFNFLQSI 323
IAVN+WYD+ +D +Y Y+ L+ +
Sbjct: 287 IAVNFWYDLDYDSRYCYYRMLEQM 310
>gb|AAQ63941.1| phospholipase-like protein [Brachypodium sylvaticum]
Length = 130
Score = 171 bits (433), Expect = 8e-41
Identities = 81/130 (62%), Positives = 95/130 (72%), Gaps = 5/130 (3%)
Query: 192 TGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGE----FDLELDKPTRYVPWCSVNPFP 247
+G+KHFLL PPT+ HR Y+R+YPAA Y E E LE+++P R VPW SV+P P
Sbjct: 1 SGEKHFLLLPPTEHHRLYVRDYPAARYVTENEGEEELTGLKLEMEEPERIVPWSSVDPNP 60
Query: 248 S-PENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQSGDDGELTIAVNYWY 306
S PE + ++S FPLYF GP P CTV+AGE+LYLPSMWFHHV QS LTIAVNYWY
Sbjct: 61 SSPEEMAAQVSSFPLYFEGPRPIRCTVRAGEVLYLPSMWFHHVSQSPGPNGLTIAVNYWY 120
Query: 307 DMQFDIKYAY 316
DMQFDIKYAY
Sbjct: 121 DMQFDIKYAY 130
>gb|EAA77468.1| hypothetical protein FG07451.1 [Gibberella zeae PH-1]
gi|46126147|ref|XP_387627.1| hypothetical protein
FG07451.1 [Gibberella zeae PH-1]
Length = 314
Score = 165 bits (418), Expect = 4e-39
Identities = 106/312 (33%), Positives = 155/312 (48%), Gaps = 23/312 (7%)
Query: 2 NEKIEELWREVRELSLGSKRSIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLSLW 61
++ IE L EL+ +I+ + P+PL+F R ++ N P ++ S W + W
Sbjct: 18 DKAIENLISTFNELN---SNAIDEFQDEPSPLEFMR-YVARNTPFVVRGGASSWKACQEW 73
Query: 62 SHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLINSS 121
+ +YL +L V++ +TP G+AD T +P SL FA H ++ PF E L +
Sbjct: 74 NS-AYLLSALKGQYVNVAVTPHGNADMPT-VPPGEESLVFAKPHYEDQPFEELLEYVARQ 131
Query: 122 N-----PSQC-VAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHS 175
P+ V YAQ QND R EY S+ D + + +A A +P+AVNLWIGN S
Sbjct: 132 ETDPDFPADAEVRYAQTQNDNLREEYISLFSDVQKDVPFARIALAKDPDAVNLWIGNSKS 191
Query: 176 STWFHKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEFDLELDKPT 235
T HKD+YEN+Y V G+KHF+L P + + P Y LE+D
Sbjct: 192 VTAMHKDNYENIYVQVLGRKHFVLLP--SLCHPCVNEQPLKPATYKRGENGMQLEMDSDA 249
Query: 236 RYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFHHVRQS--G 293
VP+ P+ E +KF + P T+ G++LYLP+MW+H V QS
Sbjct: 250 ESVPFA----IWDPDRPEQNATKFS---HLARPLRVTLNPGDMLYLPAMWYHKVLQSCAE 302
Query: 294 DDGELTIAVNYW 305
+D +AVNYW
Sbjct: 303 EDEGFVLAVNYW 314
>dbj|BAA91385.1| unnamed protein product [Homo sapiens]
Length = 235
Score = 162 bits (409), Expect = 5e-38
Identities = 86/234 (36%), Positives = 125/234 (52%), Gaps = 28/234 (11%)
Query: 107 QNLPFPEALRLINSSNPSQCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAV 166
+ LP L ++ V Y Q+Q SE ++ D + H+ WA+EA G P+AV
Sbjct: 6 RRLPLSFVLDVLEGRAQHPGVLYVQKQCSNLPSELPQLLPDLESHVPWASEALGKMPDAV 65
Query: 167 NLWIGNKHSSTWFHKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYK------- 219
N W+G + T HKDHYENLY VV+G+KHFL PP+D Y TY+
Sbjct: 66 NFWLGEAAAVTSLHKDHYENLYCVVSGEKHFLFHPPSDRPFIPYELYTPGTYQPSDRPFI 125
Query: 220 ----------YYMETGEFDLELDKPTRYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPF 269
E G F + ++ VPW ++P L +++++P Y +
Sbjct: 126 PYELYTPATYQLTEEGTFKVVDEEAMEKVPWIPLDP------LAPDLARYPSY-SQAQAL 178
Query: 270 ECTVKAGEILYLPSMWFHHVRQSGDDGELTIAVNYWYDMQFDIKYAYFNFLQSI 323
C V+AGE+LYLP++WFHHV+QS + IAVN+WYDM++D+KY+YF L S+
Sbjct: 179 RCMVRAGEMLYLPALWFHHVQQS----QGCIAVNFWYDMEYDLKYSYFQLLDSL 228
>emb|CAB86634.1| putative protein [Arabidopsis thaliana]
gi|15240591|ref|NP_196831.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|11281369|pir||T48574 hypothetical protein T31B5.90 -
Arabidopsis thaliana
Length = 752
Score = 157 bits (398), Expect = 9e-37
Identities = 98/319 (30%), Positives = 164/319 (50%), Gaps = 7/319 (2%)
Query: 423 IYTSLVKECTLSTDPETAIELHTQIITRGIELPLTLLNRILIMFVSCGLLENARRVFDVM 482
+YT+L+K + ++H +I G+ ++ I+ M+V CG L A+RVFD M
Sbjct: 186 MYTTLLKSLVNPRALDFGRQIHAHVIRAGLCSNTSIETGIVNMYVKCGWLVGAKRVFDQM 245
Query: 483 SVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQLDVMGFSFPPWIWSCLLKACACTMNV 542
+V+ + L V Y + G +A+ +FV ++ + G + +++S +LKACA +
Sbjct: 246 AVKKPVACTGLMVGYTQAGRARDALKLFVDLVTE----GVEWDSFVFSVVLKACASLEEL 301
Query: 543 PLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKCLEDANMVFNRVSRHNTLTWTAKIVSS 602
LG Q+H C+ KLG V + + L+ FY + E A F + N ++W+A I
Sbjct: 302 NLGKQIHACVAKLGLESEVSVGTPLVDFYIKCSSFESACRAFQEIREPNDVSWSAIISGY 361
Query: 603 CRERHFSEALGDFKKMGRVGVK-KDSFTFSSVLKACGRMQNRGSCGEQVHADAIKLGLDS 661
C+ F EA+ FK + +SFT++S+ +AC + + + G QVHADAIK L
Sbjct: 362 CQMSQFEEAVKTFKSLRSKNASILNSFTYTSIFQACSVLAD-CNIGGQVHADAIKRSLIG 420
Query: 662 DSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVDSLNAMLMGYIQNGLYIEAVKFVYQM 721
Y + +LI MY + G L DA VFE N ++ + A + G+ G EA++ +M
Sbjct: 421 SQYGESALITMYSKCGCLDDANEVFESMDNP-DIVAWTAFISGHAYYGNASEALRLFEKM 479
Query: 722 KAAGVQPHEPLLEKLRIAC 740
+ G++P+ + AC
Sbjct: 480 VSCGMKPNSVTFIAVLTAC 498
Score = 121 bits (304), Expect = 7e-26
Identities = 94/320 (29%), Positives = 151/320 (46%), Gaps = 8/320 (2%)
Query: 398 RRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDPETAIELHTQIITRGIELPLT 457
+ RK + L MD ++ Y L + C LH ++ GIE P
Sbjct: 60 KHRKLNEAFEFLQEMDKAGVSVSSYSYQCLFEACRELRSLSHGRLLHDRM-RMGIENPSV 118
Query: 458 LL-NRILIMFVSCGLLENARRVFDVMSVRDFHSWATLFVSYYENGEYENAIDVFVSMLCQ 516
LL N +L M+ C LE+A ++FD MS + S T+ +Y E G + A+ +F ML
Sbjct: 119 LLQNCVLQMYCECRSLEDADKLFDEMSELNAVSRTTMISAYAEQGILDKAVGLFSGMLAS 178
Query: 517 LDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGRFKC 576
G P +++ LLK+ + G Q+H +++ G C + I + ++ Y +
Sbjct: 179 ----GDKPPSSMYTTLLKSLVNPRALDFGRQIHAHVIRAGLCSNTSIETGIVNMYVKCGW 234
Query: 577 LEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSVLKA 636
L A VF++++ + T +V + +AL F + GV+ DSF FS VLKA
Sbjct: 235 LVGAKRVFDQMAVKKPVACTGLMVGYTQAGRARDALKLFVDLVTEGVEWDSFVFSVVLKA 294
Query: 637 CGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNERNVD 696
C ++ + G+Q+HA KLGL+S+ V L+ Y + A F+ R +V
Sbjct: 295 CASLEEL-NLGKQIHACVAKLGLESEVSVGTPLVDFYIKCSSFESACRAFQEIREPNDV- 352
Query: 697 SLNAMLMGYIQNGLYIEAVK 716
S +A++ GY Q + EAVK
Sbjct: 353 SWSAIISGYCQMSQFEEAVK 372
Score = 80.9 bits (198), Expect = 1e-13
Identities = 58/229 (25%), Positives = 112/229 (48%), Gaps = 2/229 (0%)
Query: 514 LCQLDVMGFSFPPWIWSCLLKACACTMNVPLGMQVHGCLLKLGACDHVLISSSLIRFYGR 573
L ++D G S + + CL +AC ++ G +H + VL+ + +++ Y
Sbjct: 71 LQEMDKAGVSVSSYSYQCLFEACRELRSLSHGRLLHDRMRMGIENPSVLLQNCVLQMYCE 130
Query: 574 FKCLEDANMVFNRVSRHNTLTWTAKIVSSCRERHFSEALGDFKKMGRVGVKKDSFTFSSV 633
+ LEDA+ +F+ +S N ++ T I + + +A+G F M G K S ++++
Sbjct: 131 CRSLEDADKLFDEMSELNAVSRTTMISAYAEQGILDKAVGLFSGMLASGDKPPSSMYTTL 190
Query: 634 LKACGRMQNRGSCGEQVHADAIKLGLDSDSYVQCSLIAMYGRSGLLRDAELVFEMTRNER 693
LK+ + G Q+HA I+ GL S++ ++ ++ MY + G L A+ VF+ ++
Sbjct: 191 LKSLVNPRAL-DFGRQIHAHVIRAGLCSNTSIETGIVNMYVKCGWLVGAKRVFDQMAVKK 249
Query: 694 NVDSLNAMLMGYIQNGLYIEAVKFVYQMKAAGVQPHEPLLEKLRIACGS 742
V + +++GY Q G +A+K + GV+ + + AC S
Sbjct: 250 PV-ACTGLMVGYTQAGRARDALKLFVDLVTEGVEWDSFVFSVVLKACAS 297
>gb|AAC78836.1| cytosolic phospholipase A2 beta [Homo sapiens]
gi|4826914|ref|NP_005081.1| phospholipase A2, group IVB
[Homo sapiens]
Length = 1012
Score = 154 bits (390), Expect = 8e-36
Identities = 85/228 (37%), Positives = 123/228 (53%), Gaps = 14/228 (6%)
Query: 5 IEELWREVRELSLGSKR-----SIERLESAPTPLQFHRNFITPNKPCIISNSISHWPSLS 59
+E + E+RE ++ ++ L+ PTPL F+R+++ PN+PCII N++ HWP+L
Sbjct: 6 LEAVRSELREFPAAARELCVPLAVPYLDKPPTPLHFYRDWVCPNRPCIIRNALQHWPALQ 65
Query: 60 LWSHPSYLTQSLSSTTVSLHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEALRLIN 119
WS P Y ++ ST VS+ +TP G AD++ F + LP L ++
Sbjct: 66 KWSLP-YFRATVGSTEVSVAVTPDGYADAVR-------GDRFMMPAERRLPLSFVLDVLE 117
Query: 120 SSNPSQCVAYAQQQNDCFRSEYDSIVKDCDQHIAWATEAFGLEPEAVNLWIGNKHSSTWF 179
V Y Q+Q SE ++ D + H+ WA+EA G P+AVN W+G + T
Sbjct: 118 GRAQHPGVLYVQKQCSNLPSELPQLLPDLESHVPWASEALGKMPDAVNFWLGEAAAVTSL 177
Query: 180 HKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEF 227
HKDHYENLY VV+G+KHFL PP+D Y ATY+ E G F
Sbjct: 178 HKDHYENLYCVVSGEKHFLFHPPSDRPFIPYELYTPATYQ-LTEEGTF 224
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,371,892,974
Number of Sequences: 2540612
Number of extensions: 62639379
Number of successful extensions: 268400
Number of sequences better than 10.0: 1413
Number of HSP's better than 10.0 without gapping: 662
Number of HSP's successfully gapped in prelim test: 778
Number of HSP's that attempted gapping in prelim test: 253883
Number of HSP's gapped (non-prelim): 7050
length of query: 749
length of database: 863,360,394
effective HSP length: 136
effective length of query: 613
effective length of database: 517,837,162
effective search space: 317434180306
effective search space used: 317434180306
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 79 (35.0 bits)
Medicago: description of AC144759.9


