

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144731.10 + phase: 0 /pseudo
(222 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
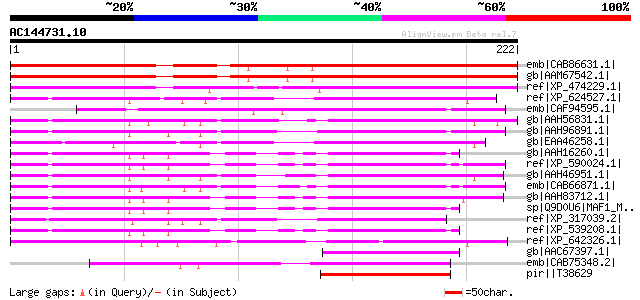
Sequences producing significant alignments: (bits) Value
emb|CAB86631.1| putative protein [Arabidopsis thaliana] gi|11358... 201 2e-50
gb|AAM67542.1| unknown protein [Arabidopsis thaliana] gi|1934794... 201 2e-50
ref|XP_474229.1| OSJNBa0084K01.13 [Oryza sativa (japonica cultiv... 162 5e-39
ref|XP_624527.1| PREDICTED: similar to ENSANGP00000019317 [Apis ... 87 3e-16
emb|CAF94595.1| unnamed protein product [Tetraodon nigroviridis] 87 4e-16
gb|AAH56831.1| Zgc:63803 [Danio rerio] gi|51701652|sp|Q6PGU2|MAF... 85 1e-15
gb|AAH96891.1| Unknown (protein for MGC:112356) [Danio rerio] 84 3e-15
gb|EAA46258.1| CG40196-PB.3 [Drosophila melanogaster] gi|3092378... 82 1e-14
gb|AAH16260.1| Repressor of RNA polymerase III transcription MAF... 80 3e-14
ref|XP_590024.1| PREDICTED: similar to Repressor of RNA polymera... 80 5e-14
gb|AAH46951.1| Maf1-pending-prov protein [Xenopus laevis] 79 7e-14
emb|CAB66871.1| hypothetical protein [Homo sapiens] gi|51701694|... 79 7e-14
gb|AAH83712.1| Repressor of RNA polymerase III transcription MAF... 79 7e-14
sp|Q9D0U6|MAF1_MOUSE Repressor of RNA polymerase III transcripti... 79 9e-14
ref|XP_317039.2| ENSANGP00000019317 [Anopheles gambiae str. PEST... 79 1e-13
ref|XP_539208.1| PREDICTED: similar to MAF1 homolog [Canis famil... 78 2e-13
ref|XP_642326.1| repressor of RNA polymerase III transcription [... 78 2e-13
gb|AAC67397.1| Hypothetical protein C43H8.2 [Caenorhabditis eleg... 57 5e-07
emb|CAB75348.2| n150 [Schizosaccharomyces pombe] gi|19114924|ref... 55 1e-06
pir||T38629 hypothetical protein SPAC31G5.12c - fission yeast (... 55 2e-06
>emb|CAB86631.1| putative protein [Arabidopsis thaliana] gi|11358307|pir||T48571
hypothetical protein T31B5.60 - Arabidopsis thaliana
Length = 283
Score = 201 bits (510), Expect = 2e-50
Identities = 120/235 (51%), Positives = 147/235 (62%), Gaps = 25/235 (10%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLE LDRL+ FL +LNLGERTIKGC+EAYSCKH+G+DK+LS+SL NE+LDYLGKSS
Sbjct: 60 MKFLEYTNLDRLNVFLGHLNLGERTIKGCLEAYSCKHAGSDKRLSLSLENEMLDYLGKSS 119
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAGRHWFT*FLPSITCIQIM----ISAQLKLTSFSQR 116
D DS SSPVD L +R+S + +T Q+ SA FS+
Sbjct: 120 DTDS-------SSPVDLLLSRSSRKALIYLV-----LTLYQMYPDYDFSAVKAHQFFSEE 167
Query: 117 NHGT---VLSRFLKPTCL------KPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPL 167
+ T + + ++ + SLL+ ++KA+DEVV L +CEIY Y P+ ADP
Sbjct: 168 SWDTFKQIFNNYMFEASKEWTERNEDGSLLEVIYKALDEVVKLAECEIYVYNPNPNADPF 227
Query: 168 PERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
E GAIWSF FLFYNRKLKR+ FR SC SNL D F D E DEEIFADMD
Sbjct: 228 LEEGAIWSFCFLFYNRKLKRVAGFRFSCTSNLANDAFLTDSPPYEEDEEIFADMD 282
>gb|AAM67542.1| unknown protein [Arabidopsis thaliana] gi|19347946|gb|AAL86308.1|
unknown protein [Arabidopsis thaliana]
gi|22326767|ref|NP_196828.2| expressed protein
[Arabidopsis thaliana]
Length = 224
Score = 201 bits (510), Expect = 2e-50
Identities = 120/235 (51%), Positives = 147/235 (62%), Gaps = 25/235 (10%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLE LDRL+ FL +LNLGERTIKGC+EAYSCKH+G+DK+LS+SL NE+LDYLGKSS
Sbjct: 1 MKFLEYTNLDRLNVFLGHLNLGERTIKGCLEAYSCKHAGSDKRLSLSLENEMLDYLGKSS 60
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAGRHWFT*FLPSITCIQIM----ISAQLKLTSFSQR 116
D DS SSPVD L +R+S + +T Q+ SA FS+
Sbjct: 61 DTDS-------SSPVDLLLSRSSRKALIYLV-----LTLYQMYPDYDFSAVKAHQFFSEE 108
Query: 117 NHGT---VLSRFLKPTCL------KPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPL 167
+ T + + ++ + SLL+ ++KA+DEVV L +CEIY Y P+ ADP
Sbjct: 109 SWDTFKQIFNNYMFEASKEWTERNEDGSLLEVIYKALDEVVKLAECEIYVYNPNPNADPF 168
Query: 168 PERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
E GAIWSF FLFYNRKLKR+ FR SC SNL D F D E DEEIFADMD
Sbjct: 169 LEEGAIWSFCFLFYNRKLKRVAGFRFSCTSNLANDAFLTDSPPYEEDEEIFADMD 223
>ref|XP_474229.1| OSJNBa0084K01.13 [Oryza sativa (japonica cultivar-group)]
gi|38346073|emb|CAE04841.2| OSJNBa0084K01.13 [Oryza
sativa (japonica cultivar-group)]
Length = 224
Score = 162 bits (411), Expect = 5e-39
Identities = 97/232 (41%), Positives = 132/232 (56%), Gaps = 19/232 (8%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLE P D ++ FL NL+LG+ TI+G +EA+SCKH+G D++LSISL +EILD L KSS
Sbjct: 1 MKFLEYTPFDSINLFLDNLDLGDCTIRGNLEAFSCKHTGNDRRLSISLEHEILDCLSKSS 60
Query: 61 DNDSPSPVDSLSSPVDSLSARTSPAG--------RHWFT*FLPSITCIQIMISAQLKLTS 112
D+D SSPV+ LS R+S H + + S + + + S
Sbjct: 61 DSDH-------SSPVEHLSCRSSRKTLIYLVLTLSHMYPDYDFSAVRAHLFFKEE-EWES 112
Query: 113 FSQRNHGTVLSRFLKPTCLKPQ--SLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPER 170
F + T LS K + SLLD++ +DEV+ + + +IY Y PD + DP E+
Sbjct: 113 FKEMTD-TYLSEASKQWAATNEGTSLLDSMTNVIDEVIKIGESDIYSYNPDQDGDPFLEK 171
Query: 171 GAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
G IWS NF FYNRKLKR+V+FR SC S + D F D +E+ DMD
Sbjct: 172 GVIWSINFFFYNRKLKRVVSFRCSCISKISGDDFLTSAPSDGEEEDALIDMD 223
>ref|XP_624527.1| PREDICTED: similar to ENSANGP00000019317 [Apis mellifera]
Length = 232
Score = 87.0 bits (214), Expect = 3e-16
Identities = 73/252 (28%), Positives = 111/252 (43%), Gaps = 59/252 (23%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE--------- 51
MK LE + ++ LS + G+ I G IE+YSCK +G DK+L + E
Sbjct: 1 MKLLESTRFEAINSALS-IKTGDSKIIGRIESYSCKMAGNDKQLYKRFNAEQGFTPHDLQ 59
Query: 52 -------------ILDYLGKSSDNDSPSPVDSLSSP-----VDSLSARTSP------AGR 87
Y S D D P D++S + +L++ P A
Sbjct: 60 ALSPPQTSLGTSPAQGYFSISGDEDGPL-CDTISRKTLFYLIATLNSAFHPDYDFSDAKS 118
Query: 88 HWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEV 147
H F+ PS+ + + + L T+ ++L AL+ A+D+
Sbjct: 119 HEFS-KEPSLQWVMNAVDSNLSATAGDHY-----------------RTLRTALWAAIDDE 160
Query: 148 VSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNL------IA 201
+SL++C+IY Y PD +DP E G +WSFN+ FYN+KLKRIV F + L +
Sbjct: 161 ISLNECDIYSYNPDFASDPFGEDGCLWSFNYFFYNKKLKRIVFFPCRAINPLYVLDSGVG 220
Query: 202 DGFSLDEICDEY 213
F++DE D Y
Sbjct: 221 SDFAMDEDEDRY 232
>emb|CAF94595.1| unnamed protein product [Tetraodon nigroviridis]
Length = 243
Score = 86.7 bits (213), Expect = 4e-16
Identities = 61/204 (29%), Positives = 97/204 (46%), Gaps = 23/204 (11%)
Query: 30 IEAYSCKHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSARTSPAGRHW 89
IE+YSCK +G DK + E G+ ++ SP S S SL ++S G +
Sbjct: 27 IESYSCKMAGDDKHMFKQFCQE-----GEPHVLEALSPPQSSSGASPSLLGKSSEDGENP 81
Query: 90 FT*FLPSITCIQIMIS-----------AQLKLTSFSQRNH-----GTVLSRFLKPTCLKP 133
+ T ++ + + + FS+ + V S +
Sbjct: 82 LSDKCCRKTLFYLITTLNESFRPDYDFSAARAHEFSRESSLNWVANAVNSSLFSAVGEEF 141
Query: 134 QSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRL 193
SL L+ A+D+ ++L C+IY Y PD ++DP E G++WSFN+ FYN+KLKRIV F
Sbjct: 142 NSLRPELWNAIDQEINLQGCDIYSYNPDLDSDPFGEEGSLWSFNYFFYNKKLKRIVFF-- 199
Query: 194 SCFSNLIADGFSLDEICDEYDEEI 217
+C S + G+ D + +E D E+
Sbjct: 200 TCRSVSVLSGYGCDSLDNELDMEL 223
>gb|AAH56831.1| Zgc:63803 [Danio rerio] gi|51701652|sp|Q6PGU2|MAF1_BRARE Repressor
of RNA polymerase III transcription MAF1 homolog
gi|47087413|ref|NP_998606.1| repressor of RNA polymerase
III transcription MAF1 [Danio rerio]
Length = 247
Score = 85.1 bits (209), Expect = 1e-15
Identities = 77/257 (29%), Positives = 118/257 (44%), Gaps = 54/257 (21%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK++ E +L+ L
Sbjct: 1 MKLLENSSFEAINSLLT-IETGDCKIIGRIESYSCKMAGDDKQMFKQFCQEGQPHVLEAL 59
Query: 57 GKS-----SDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT* 92
S N +D P+ D S +T S H F+
Sbjct: 60 SPPQSSGISPNKLSHSLDDGEGPLSDKCSRKTLFYLIATLNESFRPDYDFSRTKGHDFS- 118
Query: 93 FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDD 152
PS+ + +++ L + +H L+PQ L++A+D+ +SL +
Sbjct: 119 KEPSVNWVFNAVNSSLSAAAGEAYSH------------LQPQ-----LWEALDKEISLAE 161
Query: 153 CEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD-----GFSLD 207
C+IY Y PD ++DP E G +WSFN+ FYN++LKRIV F S +A G LD
Sbjct: 162 CDIYSYNPDLDSDPYGEEGNLWSFNYFFYNKRLKRIVFFTCRSVSLFMAPRDSGIGTELD 221
Query: 208 EICDE--YDEEIFADMD 222
DE Y+EE +MD
Sbjct: 222 LELDEGSYEEEEEGNMD 238
>gb|AAH96891.1| Unknown (protein for MGC:112356) [Danio rerio]
Length = 245
Score = 84.0 bits (206), Expect = 3e-15
Identities = 69/246 (28%), Positives = 108/246 (43%), Gaps = 50/246 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + LS L + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSRFEALSSQLC-VETGDAQIIGRIESYSCKMAGDDKHMFKQFCQEGEPHVLEAL 59
Query: 57 GKSSDNDSPSPV------DSLSSPVDSLSART-------------------SPAGRHWFT 91
+ +PSP + +P+ R S A H F+
Sbjct: 60 SPPQSSSAPSPNLLGKSGEDAENPLSDKCCRKTLFYLITTLNESFRPDYDFSAARAHEFS 119
Query: 92 *FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLD 151
PS + +++ L Q N +L L+ A+D+ ++L
Sbjct: 120 -REPSANWVANSVNSSLYSAVGEQFN-----------------TLGPELWTAIDQEINLQ 161
Query: 152 DCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICD 211
C+IY Y PD ++DP E G++WSFN+ FYN+KLKRI+ F +C S + + + +
Sbjct: 162 GCDIYSYNPDLDSDPFGEEGSLWSFNYFFYNKKLKRILFF--TCRSVSVLSSYGRSGLDN 219
Query: 212 EYDEEI 217
E D E+
Sbjct: 220 ELDMEL 225
>gb|EAA46258.1| CG40196-PB.3 [Drosophila melanogaster] gi|30923780|gb|EAA46257.1|
CG40196-PA.3 [Drosophila melanogaster]
gi|46409204|gb|AAS93759.1| LD17963p [Drosophila
melanogaster] gi|62862040|ref|NP_001015167.1|
CG40196-PA.3 [Drosophila melanogaster]
gi|62862042|ref|NP_001015168.1| CG40196-PB.3 [Drosophila
melanogaster]
Length = 226
Score = 81.6 bits (200), Expect = 1e-14
Identities = 73/243 (30%), Positives = 108/243 (44%), Gaps = 55/243 (22%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKL--------------SI 46
MK LE + + +++ LS G TI G IE+YSCK A+K L ++
Sbjct: 1 MKLLESSRFEAINNALSIQTSGI-TIFGRIESYSCKMVAAEKVLYKRFTADSHGHDLQAL 59
Query: 47 SLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSART------------------SPAGRH 88
S + D+ N+S S + ++ D++S +T S A H
Sbjct: 60 SPPQTLADFSPNFRRNNSQSGDEGITL-CDTISRKTLFYLIATLNASFEPDYDFSEAKSH 118
Query: 89 WFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVV 148
F+ PS+ + I A L + Q Q++ L+ AVD+ V
Sbjct: 119 EFS-KEPSLQWVMNSIHANLSALAGDQY-----------------QAIRQPLWSAVDDEV 160
Query: 149 SLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD---GFS 205
L +C+IY Y PD DP E G +WSFN+ FYN+KLKRIV F ++L A FS
Sbjct: 161 ILSECDIYSYNPDLSCDPFGEPGCLWSFNYFFYNKKLKRIVFFTSRAVNSLYAGETLPFS 220
Query: 206 LDE 208
++E
Sbjct: 221 MEE 223
>gb|AAH16260.1| Repressor of RNA polymerase III transcription MAF1 [Mus musculus]
gi|31982640|ref|NP_081135.2| repressor of RNA polymerase
III transcription MAF1 [Mus musculus]
Length = 258
Score = 80.5 bits (197), Expect = 3e-14
Identities = 71/231 (30%), Positives = 102/231 (43%), Gaps = 60/231 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS D SP+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGDDESPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS+ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLRWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS 205
>ref|XP_590024.1| PREDICTED: similar to Repressor of RNA polymerase III transcription
MAF1 homolog [Bos taurus]
Length = 260
Score = 79.7 bits (195), Expect = 5e-14
Identities = 73/251 (29%), Positives = 111/251 (44%), Gaps = 61/251 (24%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + P+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEDEGPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS++ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLSWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSL 206
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S + ++
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS-ISGSTYTP 213
Query: 207 DEICDEYDEEI 217
E +E D E+
Sbjct: 214 SEAGNELDMEL 224
>gb|AAH46951.1| Maf1-pending-prov protein [Xenopus laevis]
Length = 245
Score = 79.3 bits (194), Expect = 7e-14
Identities = 75/248 (30%), Positives = 107/248 (42%), Gaps = 51/248 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSHFEAVNSQLC-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 GKSSDNDSPSPV------DSLSSPVDSLSART------------------SPAGRHWFT* 92
SPS + D D S +T S A H F+
Sbjct: 60 SPPQTGISPSRLGKSQGADEEGPLSDKCSRKTLFYLIATLNESFRPDYDFSAARAHEFS- 118
Query: 93 FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDD 152
PS+ + +++ L V + T L+P L+ AVDE ++L +
Sbjct: 119 REPSLNWVVNAVNSSL------------VSALGDDFTSLRPN-----LWDAVDEEINLSE 161
Query: 153 CEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD----GFSLDE 208
C+IY Y PD + DP E G +WSFN+ FYN+KLKRIV F S G LD
Sbjct: 162 CDIYSYNPDLDCDPFGEDGNLWSFNYFFYNKKLKRIVFFSCRSISGYTYTHCDAGAELDM 221
Query: 209 ICDEYDEE 216
+E D+E
Sbjct: 222 DLEEEDDE 229
>emb|CAB66871.1| hypothetical protein [Homo sapiens]
gi|51701694|sp|Q9H063|MAF1_HUMAN Repressor of RNA
polymerase III transcription MAF1 homolog
gi|17511721|gb|AAH18714.1| MAF1 protein [Homo sapiens]
gi|15559429|gb|AAH14082.1| MAF1 protein [Homo sapiens]
gi|21411503|gb|AAH31273.1| MAF1 protein [Homo sapiens]
gi|14150013|ref|NP_115648.1| MAF1 protein [Homo sapiens]
gi|49065352|emb|CAG38494.1| MAF1 [Homo sapiens]
Length = 256
Score = 79.3 bits (194), Expect = 7e-14
Identities = 77/246 (31%), Positives = 113/246 (45%), Gaps = 51/246 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 ------GKSSDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT 91
G S S S P+ D S +T S A H F+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEEEGPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 92 *FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLD 151
PS++ + ++ L V F LKPQ L+ AVDE + L
Sbjct: 120 -REPSLSWVVNAVNCSL---------FSAVREDFKD---LKPQ-----LWNAVDEEICLA 161
Query: 152 DCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICD 211
+C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S + ++ E +
Sbjct: 162 ECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS-ISGSTYTPSEAGN 218
Query: 212 EYDEEI 217
E D E+
Sbjct: 219 ELDMEL 224
>gb|AAH83712.1| Repressor of RNA polymerase III transcription MAF1 [Rattus
norvegicus] gi|62078903|ref|NP_001014107.1| repressor of
RNA polymerase III transcription MAF1 [Rattus
norvegicus]
Length = 260
Score = 79.3 bits (194), Expect = 7e-14
Identities = 73/256 (28%), Positives = 109/256 (42%), Gaps = 64/256 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + SP+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEDESPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS+ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLRWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS------NLI 200
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F S +
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFFSCRSISGSTYTPSEA 216
Query: 201 ADGFSLDEICDEYDEE 216
+ L+ +E DEE
Sbjct: 217 GNALDLELGAEEVDEE 232
>sp|Q9D0U6|MAF1_MOUSE Repressor of RNA polymerase III transcription MAF1 homolog
gi|26335405|dbj|BAC31403.1| unnamed protein product [Mus
musculus] gi|12835580|dbj|BAB23294.1| unnamed protein
product [Mus musculus]
Length = 258
Score = 79.0 bits (193), Expect = 9e-14
Identities = 70/231 (30%), Positives = 102/231 (43%), Gaps = 60/231 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + SP+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEDESPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS+ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLRWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS 205
>ref|XP_317039.2| ENSANGP00000019317 [Anopheles gambiae str. PEST]
gi|55237278|gb|EAA12877.2| ENSANGP00000019317 [Anopheles
gambiae str. PEST]
Length = 207
Score = 78.6 bits (192), Expect = 1e-13
Identities = 68/221 (30%), Positives = 98/221 (43%), Gaps = 49/221 (22%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEI----LDYL 56
MK LE + +++ L ++ G+ TI G IE+YSCK +G DK L ++E L L
Sbjct: 1 MKLLESTRFEAVNNKL-HIQTGDSTIYGRIESYSCKMAGNDKALYKRFTSEQAPTDLQAL 59
Query: 57 GKSSDNDSPSPV---DSLSSP-----VDSLSART------------------SPAGRHWF 90
SP SLS D++S +T S A H F
Sbjct: 60 SPPQTLQDMSPQILRSSLSGDEGVTLCDTISRKTLFYLIATLNAAFEPDYDFSDARSHEF 119
Query: 91 T*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSL 150
+ PS+ + I L + Q + + AL+ +++ +SL
Sbjct: 120 S-KEPSLQWVLNSIEGNLGPVAGEQYS-----------------KIRHALWTTIEDEISL 161
Query: 151 DDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTF 191
DC+IY Y PD +DP E G +WSFN+ FYN+KLKRIV F
Sbjct: 162 KDCDIYSYNPDLNSDPFGEPGCLWSFNYFFYNKKLKRIVFF 202
>ref|XP_539208.1| PREDICTED: similar to MAF1 homolog [Canis familiaris]
Length = 300
Score = 78.2 bits (191), Expect = 2e-13
Identities = 69/231 (29%), Positives = 102/231 (43%), Gaps = 60/231 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 54 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 112
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + P+ +S D +AR+
Sbjct: 113 SPPQTSGLSPSRLSKSQGGEDEGPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 172
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS++ + ++ L V F LKPQ L+ AVDE
Sbjct: 173 RE------PSLSWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 209
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S
Sbjct: 210 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS 258
>ref|XP_642326.1| repressor of RNA polymerase III transcription [Dictyostelium
discoideum] gi|60470529|gb|EAL68509.1| repressor of RNA
polymerase III transcription [Dictyostelium discoideum]
Length = 278
Score = 77.8 bits (190), Expect = 2e-13
Identities = 72/278 (25%), Positives = 118/278 (41%), Gaps = 71/278 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYL---- 56
MK+++ L L+ +L+++++G+ + G +EAYSCK +G+DKK+ SL E+ +
Sbjct: 1 MKYIDSLQLIGLNSYLNSIDVGDALLMGELEAYSCKMAGSDKKIYRSLDKELEELSNTNS 60
Query: 57 ----------------------GKSSDND-----------SPSPVDSLS-SPVDSLSART 82
G S++N+ SPSP ++S SP +S T
Sbjct: 61 GAMNITNSNNNTNNNNNNNSNSGSSNNNNNNNNTLSVSMNSPSPTSNISVSPFGPMSNST 120
Query: 83 SPAGRHW-------------FT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPT 129
S + FT P Q S L + S + G + +L+
Sbjct: 121 SRKTMIYLIQLLNLSFPDYDFTDSKPEQ--FQKEPSLNLVINSINANLAGYIKDYYLEFE 178
Query: 130 CLKPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIV 189
L+ ++ +SL CEIY Y P++ DP E G ++SFN+ FYN+ LK+I+
Sbjct: 179 A--------KLWSTLESEISLQKCEIYLYKPEN-GDPFTENGVLFSFNYFFYNKSLKKII 229
Query: 190 TFRLSCFSNL---------IADGFSLDEICDEYDEEIF 218
F+ S L + D E D+Y E +
Sbjct: 230 FFKCRYISKLQILNDDGSDVKDPLPDQEELDDYYREYY 267
>gb|AAC67397.1| Hypothetical protein C43H8.2 [Caenorhabditis elegans]
gi|17506011|ref|NP_492777.1| homolog of yeast Maf1 like
(1L199) [Caenorhabditis elegans] gi|7497364|pir||T33467
hypothetical protein C43H8.2 - Caenorhabditis elegans
Length = 245
Score = 56.6 bits (135), Expect = 5e-07
Identities = 24/60 (40%), Positives = 34/60 (56%)
Query: 138 DALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L+ +DE + DC+IY + E DP E G IW+ F+FYN+ LKR V + C S
Sbjct: 163 EELWGIIDEAIVPGDCQIYSFKSQFEDDPFTEDGCIWALAFIFYNKGLKRFVLLTIRCLS 222
>emb|CAB75348.2| n150 [Schizosaccharomyces pombe] gi|19114924|ref|NP_594012.1|
hypothetical protein SPAC31G5.12c [Schizosaccharomyces
pombe 972h-] gi|59800467|sp|O14109|YEJC_SCHPO
Hypothetical protein C31G5.12c in chromosome I
Length = 215
Score = 55.1 bits (131), Expect = 1e-06
Identities = 39/173 (22%), Positives = 78/173 (44%), Gaps = 27/173 (15%)
Query: 36 KHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSS-----PVDSLSAR--------- 81
+ + +DKKL ++ N + L S + SP SL+ P+D S+R
Sbjct: 12 RSTNSDKKLFKAIENRCQEDLFALSSSKSPEYAFSLTQQSPFGPLDQSSSRRTFMYIVAT 71
Query: 82 -TSPAGRHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDAL 140
+ H F+ P+ + +S + + + N G + + ++ +
Sbjct: 72 LNASYPDHDFSSLQPTDFYKEPSLSRVVDSVNSTLNNIG------------RGRLSVNGI 119
Query: 141 FKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRL 193
++ +D ++L DC +Y Y PDS++DP + IW ++ F+N+ +KR++ L
Sbjct: 120 WEIIDRHINLSDCSVYSYTPDSDSDPYGDDALIWGMSYFFFNKNMKRMLYLSL 172
>pir||T38629 hypothetical protein SPAC31G5.12c - fission yeast
(Schizosaccharomyces pombe)
Length = 150
Score = 54.7 bits (130), Expect = 2e-06
Identities = 18/57 (31%), Positives = 37/57 (64%)
Query: 137 LDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRL 193
++ +++ +D ++L DC +Y Y PDS++DP + IW ++ F+N+ +KR++ L
Sbjct: 51 VNGIWEIIDRHINLSDCSVYSYTPDSDSDPYGDDALIWGMSYFFFNKNMKRMLYLSL 107
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 368,985,733
Number of Sequences: 2540612
Number of extensions: 14882274
Number of successful extensions: 32913
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 32846
Number of HSP's gapped (non-prelim): 75
length of query: 222
length of database: 863,360,394
effective HSP length: 123
effective length of query: 99
effective length of database: 550,865,118
effective search space: 54535646682
effective search space used: 54535646682
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 73 (32.7 bits)
Medicago: description of AC144731.10


