

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144729.7 - phase: 0
(226 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
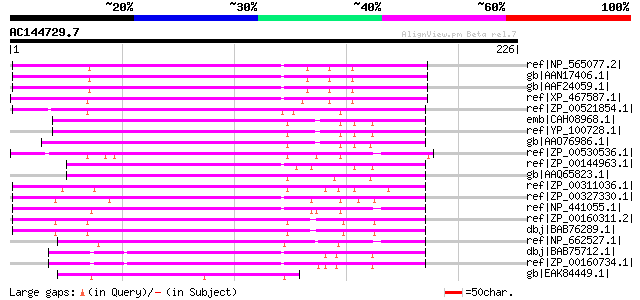
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_565077.2| peptidase U7 family protein [Arabidopsis thalia... 160 3e-38
gb|AAN17406.1| Unknown protein [Arabidopsis thaliana] gi|2805891... 160 3e-38
gb|AAF24059.1| putative protease SppA [Arabidopsis thaliana] 160 3e-38
ref|XP_467587.1| putative protease IV [Oryza sativa (japonica cu... 154 2e-36
ref|ZP_00521854.1| Peptidase S49, SppA 67 kDa type:Peptidase S49... 105 1e-21
emb|CAH08968.1| putative protease IV [Bacteroides fragilis NCTC ... 98 2e-19
ref|YP_100728.1| protease IV [Bacteroides fragilis YCH46] gi|522... 98 2e-19
gb|AAO76986.1| protease IV [Bacteroides thetaiotaomicron VPI-548... 94 2e-18
ref|ZP_00530536.1| Peptidase S49, SppA 67 kDa type:Peptidase S49... 92 1e-17
ref|ZP_00144963.1| Putative signal peptide peptidase sppA [Fusob... 85 2e-15
gb|AAQ65823.1| signal peptide peptidase SppA, 67K type [Porphyro... 84 2e-15
ref|ZP_00311036.1| COG0616: Periplasmic serine proteases (ClpP c... 83 7e-15
ref|ZP_00327330.1| COG0616: Periplasmic serine proteases (ClpP c... 82 9e-15
ref|NP_441055.1| protease IV [Synechocystis sp. PCC 6803] gi|165... 82 9e-15
ref|ZP_00160311.2| COG0616: Periplasmic serine proteases (ClpP c... 82 1e-14
dbj|BAB76289.1| protease IV [Nostoc sp. PCC 7120] gi|17232082|re... 81 2e-14
ref|NP_662527.1| protease IV [Chlorobium tepidum TLS] gi|2164765... 80 4e-14
dbj|BAB75712.1| protease IV [Nostoc sp. PCC 7120] gi|17231505|re... 75 2e-12
ref|ZP_00160734.1| COG0616: Periplasmic serine proteases (ClpP c... 74 2e-12
gb|EAK84449.1| hypothetical protein UM03629.1 [Ustilago maydis 5... 72 9e-12
>ref|NP_565077.2| peptidase U7 family protein [Arabidopsis thaliana]
gi|12325146|gb|AAG52522.1| putative protease IV;
48713-44371 [Arabidopsis thaliana]
gi|25406310|pir||F96767 proteinase IV F2P9.14 [imported]
- Arabidopsis thaliana
Length = 677
Score = 160 bits (405), Expect = 3e-38
Identities = 100/226 (44%), Positives = 130/226 (57%), Gaps = 42/226 (18%)
Query: 2 ISFRKYSKVRKWTVGISEAKEQIAIIRASGTMD----------SDIVTSNFIEKIGMVKD 51
+ ++KYS V+KWT+G++ ++QIAIIRA G++ S I+ IEKI V++
Sbjct: 354 VDYKKYSGVKKWTLGLTGGRDQIAIIRAGGSISRVKGPLSTPGSAIIAEQLIEKIRSVRE 413
Query: 52 SKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIV 111
SKK+KA I+RIDS GGD AS W I+ LA KPVIASMSD A S GYYMAM A AIV
Sbjct: 414 SKKYKAAIIRIDSPGGDALASDLMWREIKLLAETKPVIASMSDVAASGGYYMAMAANAIV 473
Query: 112 AENLTLTGSIRGTVSQNFNL---------DNELPSFIPY-------------DDAKVLDN 149
AENLTLTGSI G V+ F L + E S Y ++A++ +
Sbjct: 474 AENLTLTGSI-GVVTARFTLAKLYEKIGFNKETISRGKYAELLGAEERPLKPEEAELFEK 532
Query: 150 VV---------RLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVDA 186
+ L +DKM++VA GR+WTGKDA S GL+DA
Sbjct: 533 SAQHAYQLFRDKAALSRSMPVDKMEEVAQGRVWTGKDAHSRGLIDA 578
>gb|AAN17406.1| Unknown protein [Arabidopsis thaliana] gi|28058917|gb|AAO29968.1|
Unknown protein [Arabidopsis thaliana]
Length = 388
Score = 160 bits (405), Expect = 3e-38
Identities = 100/226 (44%), Positives = 130/226 (57%), Gaps = 42/226 (18%)
Query: 2 ISFRKYSKVRKWTVGISEAKEQIAIIRASGTMD----------SDIVTSNFIEKIGMVKD 51
+ ++KYS V+KWT+G++ ++QIAIIRA G++ S I+ IEKI V++
Sbjct: 65 VDYKKYSGVKKWTLGLTGGRDQIAIIRAGGSISRVKGPLSTPGSAIIAEQLIEKIRSVRE 124
Query: 52 SKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIV 111
SKK+KA I+RIDS GGD AS W I+ LA KPVIASMSD A S GYYMAM A AIV
Sbjct: 125 SKKYKAAIIRIDSPGGDALASDLMWREIKLLAETKPVIASMSDVAASGGYYMAMAANAIV 184
Query: 112 AENLTLTGSIRGTVSQNFNL---------DNELPSFIPY-------------DDAKVLDN 149
AENLTLTGSI G V+ F L + E S Y ++A++ +
Sbjct: 185 AENLTLTGSI-GVVTARFTLAKLYEKIGFNKETISRGKYAELLGAEERPLKPEEAELFEK 243
Query: 150 VV---------RLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVDA 186
+ L +DKM++VA GR+WTGKDA S GL+DA
Sbjct: 244 SAQHAYQLFRDKAALSRSMPVDKMEEVAQGRVWTGKDAHSRGLIDA 289
>gb|AAF24059.1| putative protease SppA [Arabidopsis thaliana]
Length = 680
Score = 160 bits (405), Expect = 3e-38
Identities = 100/226 (44%), Positives = 130/226 (57%), Gaps = 42/226 (18%)
Query: 2 ISFRKYSKVRKWTVGISEAKEQIAIIRASGTMD----------SDIVTSNFIEKIGMVKD 51
+ ++KYS V+KWT+G++ ++QIAIIRA G++ S I+ IEKI V++
Sbjct: 357 VDYKKYSGVKKWTLGLTGGRDQIAIIRAGGSISRVKGPLSTPGSAIIAEQLIEKIRSVRE 416
Query: 52 SKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIV 111
SKK+KA I+RIDS GGD AS W I+ LA KPVIASMSD A S GYYMAM A AIV
Sbjct: 417 SKKYKAAIIRIDSPGGDALASDLMWREIKLLAETKPVIASMSDVAASGGYYMAMAANAIV 476
Query: 112 AENLTLTGSIRGTVSQNFNL---------DNELPSFIPY-------------DDAKVLDN 149
AENLTLTGSI G V+ F L + E S Y ++A++ +
Sbjct: 477 AENLTLTGSI-GVVTARFTLAKLYEKIGFNKETISRGKYAELLGAEERPLKPEEAELFEK 535
Query: 150 VV---------RLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVDA 186
+ L +DKM++VA GR+WTGKDA S GL+DA
Sbjct: 536 SAQHAYQLFRDKAALSRSMPVDKMEEVAQGRVWTGKDAHSRGLIDA 581
>ref|XP_467587.1| putative protease IV [Oryza sativa (japonica cultivar-group)]
gi|46390611|dbj|BAD16095.1| putative protease IV [Oryza
sativa (japonica cultivar-group)]
Length = 674
Score = 154 bits (389), Expect = 2e-36
Identities = 97/227 (42%), Positives = 129/227 (56%), Gaps = 42/227 (18%)
Query: 1 MISFRKYSKVRKWTVGISEAKEQIAIIRASGTM----------DSDIVTSNFIEKIGMVK 50
M+ + KYS+V KWT+G+ EQIA+IRASG++ S I+ IEKI V+
Sbjct: 350 MVDYSKYSRVSKWTLGLQGGGEQIAVIRASGSITRTRSPLSVPSSGIIAEQLIEKIRTVR 409
Query: 51 DSKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAI 110
+S+K+KAVI+RIDS GGD AS W IR LA KPV+ASMSD A S GYYMAM A I
Sbjct: 410 ESEKYKAVILRIDSPGGDALASDLMWREIRLLADTKPVVASMSDVAASGGYYMAMAAPVI 469
Query: 111 VAENLTLTGSIRGTVSQNF---------NLDNELPSFIPY-------------DDAKVLD 148
VAE LTLTGSI G V+ F + + E+ S Y D+A++ +
Sbjct: 470 VAEKLTLTGSI-GVVTGKFILQKLYERIDFNKEIISKGRYAELNAADQRPLRPDEAELFE 528
Query: 149 NVV---------RLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVDA 186
+ + +D+M+ VA GR+W+G+DAAS GLVD+
Sbjct: 529 KSAQNAYALFRDKAAMSRSMNVDQMETVAQGRVWSGQDAASRGLVDS 575
>ref|ZP_00521854.1| Peptidase S49, SppA 67 kDa type:Peptidase S49, SppA [Solibacter
usitatus Ellin6076] gi|67864233|gb|EAM59240.1| Peptidase
S49, SppA 67 kDa type:Peptidase S49, SppA [Solibacter
usitatus Ellin6076]
Length = 574
Score = 105 bits (261), Expect = 1e-21
Identities = 75/224 (33%), Positives = 107/224 (47%), Gaps = 41/224 (18%)
Query: 2 ISFRKYSKVRKWTVGISEAKEQIAIIRASGTM-----------DSDIVTSNFIEKIGMVK 50
+S KYSKV +VG+ + K Q+A++ +G + +S++ + F + + V
Sbjct: 272 VSMDKYSKVTAESVGL-QGKSQVALVVGAGDIVRGDANDDGSGESNLTSYGFNKILRRVG 330
Query: 51 DSKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAI 110
+ KAVIVRIDS GG+ AS W + LA KKP++ SMSD A S GYYMAM I
Sbjct: 331 SDSQIKAVIVRIDSPGGEVTASDEIWREMNLLAKKKPMVISMSDTAASGGYYMAMTGDPI 390
Query: 111 VAENLTLTGS---------IRGTV------------SQNFNLDNELPSFIPYDDAKV--- 146
VA T TGS IRG +N ++D++ P AK+
Sbjct: 391 VAYPATFTGSIGVVFGKPNIRGLYEKLGISKDAIQRGKNADIDSDYTPLTPEQRAKLRAG 450
Query: 147 -----LDNVVRLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVD 185
D V ++ D++ +VA GR+W G A GLVD
Sbjct: 451 IDESYQDFVTKVANARHRTFDQIDQVAQGRVWLGSQAKPRGLVD 494
>emb|CAH08968.1| putative protease IV [Bacteroides fragilis NCTC 9343]
gi|60682742|ref|YP_212886.1| putative protease IV
[Bacteroides fragilis NCTC 9343]
Length = 592
Score = 97.8 bits (242), Expect = 2e-19
Identities = 67/197 (34%), Positives = 97/197 (49%), Gaps = 33/197 (16%)
Query: 20 AKEQIAIIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWEAI 79
A +I S D I+ S I + +K+ KAV++R++S GG AS+ W A+
Sbjct: 314 ASGEITDYAGSAASDEGIIGSKMIRDLRKLKEDDDVKAVVLRVNSPGGSAFASEQIWHAV 373
Query: 80 RSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIR--GTVSQNFNLDNELPS 137
+ L +KKPVI SM D A S GYY++ A +I+AE TLTGSI G + L ++
Sbjct: 374 KELKAKKPVIVSMGDYAASGGYYISCAADSIIAEPTTLTGSIGIFGMIPNVKGLTEKIG- 432
Query: 138 FIPYDDAKV-----LDNVVR--------LRLMTLGK----------------LDKMKKVA 168
+ YD K N++R L M +G+ DK++K+A
Sbjct: 433 -LTYDVVKTNQFSDFGNLMRPVNSDERALLQMMIGQGYDLFVSRCAEGRHMSKDKIEKIA 491
Query: 169 HGRIWTGKDAASHGLVD 185
GR+WTG+ A GLVD
Sbjct: 492 EGRVWTGEMAKKIGLVD 508
>ref|YP_100728.1| protease IV [Bacteroides fragilis YCH46]
gi|52217601|dbj|BAD50194.1| protease IV [Bacteroides
fragilis YCH46]
Length = 592
Score = 97.8 bits (242), Expect = 2e-19
Identities = 67/197 (34%), Positives = 97/197 (49%), Gaps = 33/197 (16%)
Query: 20 AKEQIAIIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWEAI 79
A +I S D I+ S I + +K+ KAV++R++S GG AS+ W A+
Sbjct: 314 ASGEITDYAGSAASDEGIIGSKMIRDLRKLKEDDDVKAVVLRVNSPGGSAFASEQIWHAV 373
Query: 80 RSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIR--GTVSQNFNLDNELPS 137
+ L +KKPVI SM D A S GYY++ A +I+AE TLTGSI G + L ++
Sbjct: 374 KELKAKKPVIVSMGDYAASGGYYISCAADSIIAEPTTLTGSIGIFGMIPNVKGLTEKIG- 432
Query: 138 FIPYDDAKV-----LDNVVR--------LRLMTLGK----------------LDKMKKVA 168
+ YD K N++R L M +G+ DK++K+A
Sbjct: 433 -LTYDVVKTNQFSDFGNLMRPVNSDERALLQMMIGQGYDLFVSRCAEGRHMSKDKIEKIA 491
Query: 169 HGRIWTGKDAASHGLVD 185
GR+WTG+ A GLVD
Sbjct: 492 EGRVWTGEMAKKIGLVD 508
>gb|AAO76986.1| protease IV [Bacteroides thetaiotaomicron VPI-5482]
gi|29347289|ref|NP_810792.1| protease IV [Bacteroides
thetaiotaomicron VPI-5482]
Length = 592
Score = 94.4 bits (233), Expect = 2e-18
Identities = 67/202 (33%), Positives = 98/202 (48%), Gaps = 33/202 (16%)
Query: 15 VGISEAKEQIAIIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQS 74
V + A +I S T + IV + I + +KD + KAV++R++S GG AS+
Sbjct: 309 VAVYYASGEITDYSGSSTSEEGIVGTKVIRDLRKLKDDEDVKAVVLRVNSPGGSAFASEQ 368
Query: 75 TWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIR--GTVSQNFNLD 132
W A++ L ++KPVI SM D A S GYY++ A IVAE TLTGSI G V L
Sbjct: 369 IWHAVKELKTEKPVIVSMGDYAASGGYYISCVADTIVAEPTTLTGSIGIFGMVPNVKELS 428
Query: 133 NELPSFIPYDDAKV-----LDNVVR----------LRLMTLG--------------KLDK 163
++ + YD K N++R ++T G +
Sbjct: 429 EKIG--LTYDVVKTNKFSDFGNIMRPFNQDEKTLMQMMITQGYDTFVNRCAEGRHMSKEA 486
Query: 164 MKKVAHGRIWTGKDAASHGLVD 185
++K+A GR+WTG+ A GLVD
Sbjct: 487 IEKIAEGRVWTGEAAKELGLVD 508
>ref|ZP_00530536.1| Peptidase S49, SppA 67 kDa type:Peptidase S49, SppA [Chlorobium
phaeobacteroides BS1] gi|67915867|gb|EAM65185.1|
Peptidase S49, SppA 67 kDa type:Peptidase S49, SppA
[Chlorobium phaeobacteroides BS1]
Length = 596
Score = 92.0 bits (227), Expect = 1e-17
Identities = 78/234 (33%), Positives = 112/234 (47%), Gaps = 49/234 (20%)
Query: 1 MISFRKYSKVRKWTVGISEAKEQIAIIRASGTM--DSDIVTSN----FIEK-----IGMV 49
++S +Y W E KE IA+I ASG + SD + + F E+ +
Sbjct: 279 LVSGVRYKAAVPWPYK-PETKEHIAVITASGPIVRSSDDMAAGTEQGFDEETLRSSVQAA 337
Query: 50 KDSKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGA 109
D + KA+++RIDS GGD AS + + + S KKP++ SMS A S GY +A+ A +
Sbjct: 338 LDDESVKAIVLRIDSPGGDALASANMLQVLDSARVKKPIVTSMSSVAASGGYMIALAADS 397
Query: 110 IVAENLTLTGSIR--------GTVSQNFNLDNEL----------PSFIPYDDAKV----- 146
I AE LT+TGSI + + L+ E+ F P D+A +
Sbjct: 398 IFAEPLTVTGSIGVYALKPEISKLQEKIALNREVFTRGKNADAYTVFKPLDEAGMAKFME 457
Query: 147 ---------LDNVVRLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVD--AGMP 189
LD VVR R MT ++D VA GR+W G+ A +GLVD G+P
Sbjct: 458 TTGWIYDDFLDKVVRSRKMTREEVD---AVAGGRVWMGEAAVKNGLVDRIGGLP 508
>ref|ZP_00144963.1| Putative signal peptide peptidase sppA [Fusobacterium nucleatum
subsp. vincentii ATCC 49256] gi|27886138|gb|EAA23435.1|
Putative signal peptide peptidase sppA [Fusobacterium
nucleatum subsp. vincentii ATCC 49256]
Length = 263
Score = 84.7 bits (208), Expect = 2e-15
Identities = 61/189 (32%), Positives = 91/189 (47%), Gaps = 30/189 (15%)
Query: 26 IIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASK 85
+IR ++ + S +EK+ + K++ K KAV++R++S GG S E ++ LA +
Sbjct: 2 LIRKEVSLGEEYNVSETLEKLNIAKENDKIKAVVLRVNSPGGSALTSDIIAEKVKELAEE 61
Query: 86 KPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIRGTVS-----QNFNLDN----ELP 136
KPV SMS A S GYY++ A I + T+TGSI G VS DN E
Sbjct: 62 KPVYVSMSSVAASGGYYISANANKIFVDRNTITGSI-GVVSILPDFSKLITDNGVNIEKI 120
Query: 137 SFIPYDDAKVLDNVVR-----------------LRLMTLGK---LDKMKKVAHGRIWTGK 176
S Y D +D+ L +++ G+ +K+K +A GRIWTG
Sbjct: 121 SDGEYSDLYSVDSFTEKKYNKIYNSNLKVYEDFLNVVSKGRKIDKEKLKTIAEGRIWTGD 180
Query: 177 DAASHGLVD 185
+A GL D
Sbjct: 181 EAIKIGLAD 189
>gb|AAQ65823.1| signal peptide peptidase SppA, 67K type [Porphyromonas gingivalis
W83] gi|34540445|ref|NP_904924.1| signal peptide
peptidase SppA, 67K type [Porphyromonas gingivalis W83]
Length = 595
Score = 84.3 bits (207), Expect = 2e-15
Identities = 56/189 (29%), Positives = 86/189 (44%), Gaps = 29/189 (15%)
Query: 26 IIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWEAIRSLASK 85
II+ D +T ++I D KAV++R++S GG S+ W+ + L +K
Sbjct: 321 IIKKPFDTDGSSITQELAKEIKAAADDDDIKAVVLRVNSPGGSAFTSEQIWKQVADLKAK 380
Query: 86 KPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIR--------GTVSQNFNLDNELPS 137
KP++ SM D A S GYY+A A +IVAE+ TLTGSI V++ ++ ++
Sbjct: 381 KPIVVSMGDVAASGGYYIACAANSIVAEHTTLTGSIGIFGMFPNFAGVAKKIGVNMDVVQ 440
Query: 138 FIPYDD-------AKVLDNVVRLRLMTLG--------------KLDKMKKVAHGRIWTGK 176
Y D V D + R + G ++ +A GR+W G
Sbjct: 441 TSKYADLGNTFAPMTVEDRALIQRYIEQGYDLFLTRVSEGRNRTKAQIDSIAQGRVWLGD 500
Query: 177 DAASHGLVD 185
A + GLVD
Sbjct: 501 KALALGLVD 509
>ref|ZP_00311036.1| COG0616: Periplasmic serine proteases (ClpP class) [Cytophaga
hutchinsonii]
Length = 583
Score = 82.8 bits (203), Expect = 7e-15
Identities = 67/221 (30%), Positives = 101/221 (45%), Gaps = 37/221 (16%)
Query: 2 ISFRKYSKVRKWTVGISEAKE-QIAIIRASGTMDSD------IVTSNFIEKIGMVKDSKK 54
I F Y K+ K + + E IA++ A+G + S I + F + +++ K
Sbjct: 280 IDFVSYKKIIKGKPEKTTSSEPHIAVLFANGEIQSGKGDNETIGSETFCTDLRRLREDKN 339
Query: 55 FKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAEN 114
KA+++R++S GG AS W I KPVIASM + A S GYY+AM IVA
Sbjct: 340 VKAIVLRVNSPGGSALASDLIWREIMLAREVKPVIASMGNVAASGGYYIAMACDTIVASP 399
Query: 115 LTLTGSIR------------------GTVSQNFNLDNELPSFI-PYDDAK--VLDNVVR- 152
T+TGSI T + L ++L S P DA+ ++ N V
Sbjct: 400 ATITGSIGVFGLLMNTEDLLNNKLGISTDREKTGLYSDLGSLTRPVTDAERMIIQNEVNA 459
Query: 153 -----LRLMTLGKLDKMKKV---AHGRIWTGKDAASHGLVD 185
+R G+ ++ + A GR+W GKDA + L+D
Sbjct: 460 IYATFIRKAAEGRHTSVETIEVHASGRVWAGKDAKENNLID 500
>ref|ZP_00327330.1| COG0616: Periplasmic serine proteases (ClpP class) [Trichodesmium
erythraeum IMS101]
Length = 608
Score = 82.4 bits (202), Expect = 9e-15
Identities = 67/223 (30%), Positives = 113/223 (50%), Gaps = 40/223 (17%)
Query: 2 ISFRKYSKVRKWTVGISE---AKEQIAIIRASGTMDSDIVTSNFI------EKIGMVKDS 52
IS YSKV + +S+ + +IA++ A G + + TS I +K+ ++
Sbjct: 297 ISLNNYSKVPRVAKTLSKNIKSNNKIAVVYAQGEVVNGSGTSRQIGGDRLAKKLRQLRLD 356
Query: 53 KKFKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVA 112
+K KAV++R++S GG AS+ ++ ++ +KPVI SM + A S GY+++M A IVA
Sbjct: 357 EKVKAVVLRVNSPGGSASASEVISREVKLMSEEKPVIVSMGNIAASGGYWISMNADRIVA 416
Query: 113 ENLTLTGSIRGTVSQNFNLDN-ELPSFIPYDDAKV-----LDNVVRLR----LMTLGKL- 161
E T+TGSI G FN+ + I +D K+ L+ R + L+ + K+
Sbjct: 417 EVNTITGSI-GVFGVLFNIQEIANQNGITWDVVKIGKFADLNTTSRPKTEDELVLIQKMV 475
Query: 162 -------------------DKMKKVAHGRIWTGKDAASHGLVD 185
+K++++A GR+W+G +A GLVD
Sbjct: 476 DSIYERFIKNVATARNLAPEKVEEIAQGRVWSGANAQKLGLVD 518
>ref|NP_441055.1| protease IV [Synechocystis sp. PCC 6803] gi|1652816|dbj|BAA17735.1|
protease IV [Synechocystis sp. PCC 6803]
gi|2499882|sp|P73689|SPPA_SYNY3 Protease IV homolog
(Endopeptidase IV) (Signal peptide peptidase)
Length = 610
Score = 82.4 bits (202), Expect = 9e-15
Identities = 68/223 (30%), Positives = 103/223 (45%), Gaps = 43/223 (19%)
Query: 2 ISFRKYSKVRKWTVGISEAKEQIAIIRASGTMDS------DIVTSNFIEKIGMVKDSKKF 55
IS +Y +++ W + +IAI+ G++ + +I + E + ++
Sbjct: 299 ISLAEYHRLQNWETENHDQDPKIAIVYLEGSIVNGRGTWENIGGDRYGELLRTIRQDDDI 358
Query: 56 KAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENL 115
KAV++RI+S GG A+ W + L ++KPVI SM + A S GY++A IVA+
Sbjct: 359 KAVVLRINSPGGSASAADIIWREVELLQAQKPVIISMGNVAASGGYWIATAGEKIVAQPN 418
Query: 116 TLTGSIRGTVSQNFNLDN---------------EL----PSFIPYDDAKV---------- 146
T+TGSI G S FN++N EL S P + ++
Sbjct: 419 TVTGSI-GVFSILFNVENLGDRLGLNWDEVATGELANVGSSIKPKTELELAIFQRSVDQV 477
Query: 147 ----LDNVVRLRLMTLGKLDKMKKVAHGRIWTGKDAASHGLVD 185
LD V R R ++ LD VA GR+WTG A GLVD
Sbjct: 478 YEIFLDKVGRARNLSPTALD---SVAQGRVWTGLAAQKVGLVD 517
>ref|ZP_00160311.2| COG0616: Periplasmic serine proteases (ClpP class) [Anabaena
variabilis ATCC 29413]
Length = 609
Score = 81.6 bits (200), Expect = 1e-14
Identities = 66/222 (29%), Positives = 103/222 (45%), Gaps = 40/222 (18%)
Query: 2 ISFRKYSKVRKWTVGISE-AKEQIAIIRASGTM------DSDIVTSNFIEKIGMVKDSKK 54
IS R+Y++V ++G++ +K +IA++ A G + D I F ++ +
Sbjct: 294 ISLRRYAQVPGQSLGVARNSKNKIAVVYAEGDIVDGKGDDGQIGGDRFARIFNKIRQDEN 353
Query: 55 FKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAEN 114
KAV++RI+S GG AS+ IR KPV+ SM D A S GY++A + I AE
Sbjct: 354 VKAVVLRINSPGGSATASEVMQREIRLTRESKPVVVSMGDYAASGGYWIATDSNRIFAEP 413
Query: 115 LTLTGSIR--GTVSQNFNLDNELPSFIPYDDAKVL---------------------DNVV 151
T+TGSI G + L N+ + I +D K +V
Sbjct: 414 NTITGSIGVFGVLFNGQKLAND--NGITWDAVKTARYADSQTVARPKSPQEIAIYQRSVD 471
Query: 152 RLRLMTLGKL--------DKMKKVAHGRIWTGKDAASHGLVD 185
R+ M + K+ K+ ++A GR+W+G A GLVD
Sbjct: 472 RIYNMFVNKVAQGRKLPTQKVAQIAQGRVWSGVTAKQIGLVD 513
>dbj|BAB76289.1| protease IV [Nostoc sp. PCC 7120] gi|17232082|ref|NP_488630.1|
protease IV [Nostoc sp. PCC 7120]
gi|25290101|pir||AF2379 proteinase IV [imported] -
Nostoc sp. (strain PCC 7120)
Length = 609
Score = 81.3 bits (199), Expect = 2e-14
Identities = 66/222 (29%), Positives = 103/222 (45%), Gaps = 40/222 (18%)
Query: 2 ISFRKYSKVRKWTVGISE-AKEQIAIIRASGTM------DSDIVTSNFIEKIGMVKDSKK 54
IS R+Y++V ++G+ + +K +IA++ A G + D I F ++ +
Sbjct: 294 ISLRRYAQVPGQSLGLEKNSKNKIAVVYAEGDIVDGKGDDGQIGGDRFARIFNKIRQDEN 353
Query: 55 FKAVIVRIDSVGGDFHASQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAEN 114
KAV++RI+S GG AS+ IR KPV+ SM D A S GY++A + I AE
Sbjct: 354 VKAVVLRINSPGGSATASEVMQREIRLTRESKPVVVSMGDYAASGGYWIATDSNRIFAEP 413
Query: 115 LTLTGSIR--GTVSQNFNLDNELPSFIPYDDAKVL---------------------DNVV 151
T+TGSI G + L N+ + I +D K +V
Sbjct: 414 NTITGSIGVFGVLFNGQKLAND--NGITWDAVKTARYADSQTVARPKSPQEIAIYQRSVD 471
Query: 152 RLRLMTLGKL--------DKMKKVAHGRIWTGKDAASHGLVD 185
R+ M + K+ K+ ++A GR+W+G A GLVD
Sbjct: 472 RIYNMFVNKVAQGRKLPTQKVAEIAQGRVWSGVTAKQIGLVD 513
>ref|NP_662527.1| protease IV [Chlorobium tepidum TLS] gi|21647650|gb|AAM72869.1|
protease IV [Chlorobium tepidum TLS]
Length = 597
Score = 80.1 bits (196), Expect = 4e-14
Identities = 65/207 (31%), Positives = 94/207 (45%), Gaps = 47/207 (22%)
Query: 22 EQIAIIRASGTMDSDIV----------TSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHA 71
++IA+I +G + SD + E + D K KA+++RIDS GGD A
Sbjct: 298 DRIAVINITGMIVSDGAGGMSEGDGTDVATVKEALQTAIDDLKVKAIVLRIDSPGGDALA 357
Query: 72 SQSTWEAIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSI---------- 121
+ + E + +KKP++ASMS A S GY +A+ I AE LT+TGSI
Sbjct: 358 ASTMLELLNEAKAKKPIVASMSGLAASGGYMVALAGDKIFAEPLTITGSIGVFSLKPDLS 417
Query: 122 ---------RGTVSQNFNLDNELPSFIPYDDAK--------------VLDNVVRLRLMTL 158
R + + D E P F +DDA + V + R MT
Sbjct: 418 SLLEKTGIRREVLIRGRFADAETP-FRAFDDASFRKFVELTGTVYEDFIAKVAKGRHMTP 476
Query: 159 GKLDKMKKVAHGRIWTGKDAASHGLVD 185
++D VA GR+W+GK A GL+D
Sbjct: 477 AQVD---AVAGGRVWSGKRALEVGLID 500
>dbj|BAB75712.1| protease IV [Nostoc sp. PCC 7120] gi|17231505|ref|NP_488053.1|
protease IV [Nostoc sp. PCC 7120]
gi|25332287|pir||AF2307 proteinase IV [imported] -
Nostoc sp. (strain PCC 7120)
Length = 273
Score = 74.7 bits (182), Expect = 2e-12
Identities = 60/198 (30%), Positives = 98/198 (49%), Gaps = 33/198 (16%)
Query: 18 SEAKEQIAIIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWE 77
S+ ++QIA I +G + S +E + V++ +KF A+++RIDS GG SQ +
Sbjct: 7 SKFRKQIARIEITGAIASG-TRKRVLEALKTVEE-RKFPALLLRIDSPGGTVGDSQEIYS 64
Query: 78 AIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIRGTVSQNFNLDNELPS 137
A++ L K ++AS + + S G Y+ MGA I+A T+TGSI G + + NL+ L
Sbjct: 65 ALKRLREKIKIVASFGNISASGGVYIGMGAEHIMANPGTITGSI-GVILRGNNLERLLDK 123
Query: 138 FI---------PYDDA----------------KVLDNVVRLRLMTLGK-----LDKMKKV 167
PY D +++D + + T+ + +DK+K
Sbjct: 124 IGVSFKVIKSGPYKDILSFDRELTEPEQDILQELIDTSYQQFVQTVAEGRSLAVDKVKSF 183
Query: 168 AHGRIWTGKDAASHGLVD 185
A GRI+TG+ A G+VD
Sbjct: 184 ADGRIFTGQQALELGVVD 201
>ref|ZP_00160734.1| COG0616: Periplasmic serine proteases (ClpP class) [Anabaena
variabilis ATCC 29413]
Length = 273
Score = 74.3 bits (181), Expect = 2e-12
Identities = 62/198 (31%), Positives = 100/198 (50%), Gaps = 33/198 (16%)
Query: 18 SEAKEQIAIIRASGTMDSDIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWE 77
S+ ++QIA I +G + S +E + V++ KKF A+++RIDS GG SQ +
Sbjct: 7 SKFRKQIARIEITGAIASG-TRKRVLEALKTVEE-KKFPALLLRIDSPGGTVGDSQEIYS 64
Query: 78 AIRSLASKKPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSIRGTVSQNFNLDNELP- 136
A++ L K ++AS + + S G Y+ MGA I+A T+TGSI G + + NL+ L
Sbjct: 65 ALKRLREKIKIVASFGNISASGGVYIGMGAEHIMANPGTITGSI-GVILRGNNLERLLDK 123
Query: 137 ---SFI-----PYDDA----------------KVLDNVVRLRLMTLGK-----LDKMKKV 167
SF PY D +++D + + T+ + ++K+K
Sbjct: 124 VGVSFKVIKSGPYKDILSFDRELTEPEQDILQELIDTSYQQFVQTVAEGRSLAVEKVKSF 183
Query: 168 AHGRIWTGKDAASHGLVD 185
A GRI+TG+ A G+VD
Sbjct: 184 ADGRIFTGQQALELGVVD 201
>gb|EAK84449.1| hypothetical protein UM03629.1 [Ustilago maydis 521]
gi|49074184|ref|XP_401244.1| hypothetical protein
UM03629.1 [Ustilago maydis 521]
Length = 949
Score = 72.4 bits (176), Expect = 9e-12
Identities = 43/118 (36%), Positives = 69/118 (58%), Gaps = 10/118 (8%)
Query: 22 EQIAIIRASGTMDS---DIVTSNFIEKIGMVKDSKKFKAVIVRIDSVGGDFHASQSTWEA 78
E++ ++ G + S + TS+ ++ + + K K++++RIDS GGD AS+S W+A
Sbjct: 382 ERVGVVYLLGGISSAPGEFSTSSVLKGLKEAAEHKDIKSIVLRIDSGGGDVVASESIWDA 441
Query: 79 IRSLASK--KPVIASMSDAATSAGYYMAMGAGAIVAENLTLTGSI-----RGTVSQNF 129
+R + KPV+AS + A S GYY A A AI+A T+TGSI R T+++ F
Sbjct: 442 VRRVREDYGKPVVASFGNTAASGGYYAASAADAILACENTVTGSIGVASLRPTITRAF 499
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.134 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 359,914,221
Number of Sequences: 2540612
Number of extensions: 13696503
Number of successful extensions: 34610
Number of sequences better than 10.0: 420
Number of HSP's better than 10.0 without gapping: 245
Number of HSP's successfully gapped in prelim test: 175
Number of HSP's that attempted gapping in prelim test: 33978
Number of HSP's gapped (non-prelim): 624
length of query: 226
length of database: 863,360,394
effective HSP length: 123
effective length of query: 103
effective length of database: 550,865,118
effective search space: 56739107154
effective search space used: 56739107154
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 73 (32.7 bits)
Medicago: description of AC144729.7


