

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
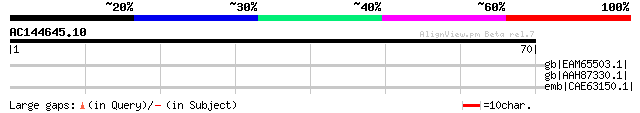
Query= AC144645.10 - phase: 0 /pseudo
(70 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|EAM65503.1| Thiolase [Jannaschia sp. CCS1] 31 7.3
gb|AAH87330.1| LOC496144 protein [Xenopus laevis] 31 7.3
emb|CAE63150.1| Hypothetical protein CBG07465 [Caenorhabditis br... 31 9.6
>gb|EAM65503.1| Thiolase [Jannaschia sp. CCS1]
Length = 403
Score = 31.2 bits (69), Expect = 7.3
Identities = 14/32 (43%), Positives = 19/32 (58%)
Query: 30 HEVDTARFFQTTLSRKIEKNCLNGHVVKQIRW 61
HEV +AR TL+ E+N L GH V+ + W
Sbjct: 24 HEVTSARLSAFTLNTLKERNGLEGHAVEDVIW 55
>gb|AAH87330.1| LOC496144 protein [Xenopus laevis]
Length = 255
Score = 31.2 bits (69), Expect = 7.3
Identities = 13/32 (40%), Positives = 18/32 (55%)
Query: 23 SVLPLQAHEVDTARFFQTTLSRKIEKNCLNGH 54
S+LP AHE+ R + + + K KNCL H
Sbjct: 97 SILPANAHEIAEHRLYVSITNAKSRKNCLVSH 128
>emb|CAE63150.1| Hypothetical protein CBG07465 [Caenorhabditis briggsae]
Length = 576
Score = 30.8 bits (68), Expect = 9.6
Identities = 18/39 (46%), Positives = 24/39 (61%), Gaps = 2/39 (5%)
Query: 27 LQAHEVDTARFFQTTLSR-KIEKNCLNGHVVKQIRWDLQ 64
L H + T RFFQ T S+ I+KN ++ V Q RWDL+
Sbjct: 212 LSGHPMATCRFFQDTFSKFSIKKNPMSKSVRLQ-RWDLE 249
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.130 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 109,536,545
Number of Sequences: 2540612
Number of extensions: 2881065
Number of successful extensions: 5482
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 5480
Number of HSP's gapped (non-prelim): 3
length of query: 70
length of database: 863,360,394
effective HSP length: 46
effective length of query: 24
effective length of database: 746,492,242
effective search space: 17915813808
effective search space used: 17915813808
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144645.10


