

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144591.1 - phase: 0
(167 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
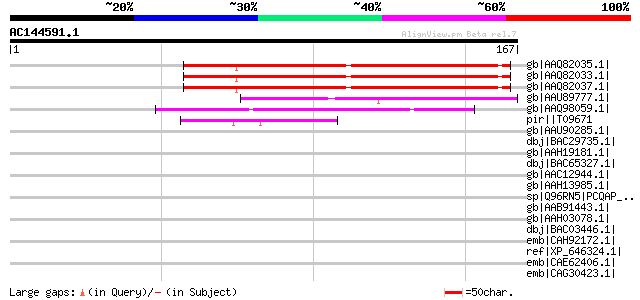
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ82035.1| gag/pol polyprotein [Pisum sativum] 100 2e-20
gb|AAQ82033.1| gag/pol polyprotein [Pisum sativum] 100 2e-20
gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum] 94 1e-18
gb|AAU89777.1| gag-pol polyprotein-like [Solanum tuberosum] 52 6e-06
gb|AAQ98059.1| unknown [Vitis cinerea x Vitis rupestris] 50 3e-05
pir||T09671 RPE15 protein - alfalfa (fragment) gi|840619|gb|AAB5... 49 5e-05
gb|AAU90285.1| putative gag/pol polyprotein, 3'-partial [Solanum... 44 0.002
dbj|BAC29735.1| unnamed protein product [Mus musculus] 44 0.002
gb|AAH19181.1| PHD finger protein 21A [Mus musculus] gi|20270291... 43 0.003
dbj|BAC65327.1| PFTF1 [Mus musculus] 43 0.003
gb|AAC12944.1| TPA inducible protein [Homo sapiens] 43 0.004
gb|AAH13985.1| Positive cofactor 2, glutamine/Q-rich-associated ... 43 0.004
sp|Q96RN5|PCQAP_HUMAN Positive cofactor 2 glutamine/Q-rich-assoc... 43 0.004
gb|AAB91443.1| CTG7a [Homo sapiens] 43 0.004
gb|AAH03078.1| PCQAP protein [Homo sapiens] 43 0.004
dbj|BAC03446.1| FLJ00386 protein [Homo sapiens] 43 0.004
emb|CAH92172.1| hypothetical protein [Pongo pygmaeus] 41 0.014
ref|XP_646324.1| hypothetical protein DDB0190596 [Dictyostelium ... 41 0.014
emb|CAE62406.1| Hypothetical protein CBG06493 [Caenorhabditis br... 41 0.014
emb|CAG30423.1| PCQAP [Homo sapiens] 40 0.019
>gb|AAQ82035.1| gag/pol polyprotein [Pisum sativum]
Length = 1105
Score = 100 bits (249), Expect = 2e-20
Identities = 54/111 (48%), Positives = 71/111 (63%), Gaps = 5/111 (4%)
Query: 58 QFQQRPQPPRQPSFTQ---QPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPE 114
Q QQR Q P+Q Q QPRQ+ P RFD +PM YAE LP LL+ V+ R P P
Sbjct: 385 QQQQRVQQPQQQQQQQRPYQPRQRMPDRRFDSLPMSYAELLPELLRLGMVELRTMAP-PT 443
Query: 115 KLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTDLNPNL 165
LP G+ A++ C FH GAPGH E+C AL++ VQDL++AK ++F + PN+
Sbjct: 444 VLPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDAKAINFAPV-PNV 493
>gb|AAQ82033.1| gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 100 bits (249), Expect = 2e-20
Identities = 54/111 (48%), Positives = 71/111 (63%), Gaps = 5/111 (4%)
Query: 58 QFQQRPQPPRQPSFTQ---QPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPE 114
Q QQR Q P+Q Q QPRQ+ P RFD +PM YAE LP LL+ V+ R P P
Sbjct: 385 QQQQRVQQPQQQQQQQRPYQPRQRMPDRRFDSLPMSYAELLPELLRLGMVELRTMAP-PT 443
Query: 115 KLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTDLNPNL 165
LP G+ A++ C FH GAPGH E+C AL++ VQDL++AK ++F + PN+
Sbjct: 444 VLPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDAKAINFAPV-PNV 493
>gb|AAQ82037.1| gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 94.4 bits (233), Expect = 1e-18
Identities = 51/110 (46%), Positives = 69/110 (62%), Gaps = 4/110 (3%)
Query: 58 QFQQRPQPPRQPSFTQ--QPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEK 115
Q QQR Q P+Q + QPRQ+ P RFD +PM YAE LP LL+ V+ P P
Sbjct: 384 QQQQRVQQPQQQQQQRPYQPRQRMPDRRFDSLPMSYAELLPELLRLGLVELCTMAP-PTV 442
Query: 116 LPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTDLNPNL 165
LP G+ A++ C FH GAPGH E+C AL++ VQDL++A ++F + PN+
Sbjct: 443 LPPGYDANVRCDFHSGAPGHHTEKCRALQHKVQDLIDANAINFAPV-PNV 491
>gb|AAU89777.1| gag-pol polyprotein-like [Solanum tuberosum]
Length = 1716
Score = 52.0 bits (123), Expect = 6e-06
Identities = 26/92 (28%), Positives = 43/92 (46%), Gaps = 3/92 (3%)
Query: 77 QQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLPAGF-RADLSCVFHQGAPGH 135
++ P F P+ + L Y+ P P+P + F R D C +H + GH
Sbjct: 416 EKKPARNFTPLIKSRTKLFERLTVAGYIH--PVGPKPVDTSSKFYRPDQRCAYHSNSVGH 473
Query: 136 DVERCYALRNAVQDLVEAKILSFTDLNPNLQS 167
D E C L++ +QDL+ ++S + PN+ S
Sbjct: 474 DTEDCINLKHKIQDLINQNVVSLQTVAPNVNS 505
>gb|AAQ98059.1| unknown [Vitis cinerea x Vitis rupestris]
Length = 248
Score = 49.7 bits (117), Expect = 3e-05
Identities = 28/105 (26%), Positives = 50/105 (46%), Gaps = 2/105 (1%)
Query: 49 SLMVTKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRP 108
SL ++ P + P P P ++ R F + + + + ++
Sbjct: 109 SLAMSTRPPSRVRGPPVVTVPQGRPHPHRRC-RRHFSDLSIPLSRAFETFQAMGFLAPLA 167
Query: 109 PPPRPEKLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEA 153
P P+ +P FR DL C++HQ GH +RC ALR+A+QD++++
Sbjct: 168 PRALPDPVPPQFRLDLYCMYHQSV-GHHTDRCTALRHAIQDIIDS 211
>pir||T09671 RPE15 protein - alfalfa (fragment) gi|840619|gb|AAB50037.1| RPE15
gene product
Length = 219
Score = 48.9 bits (115), Expect = 5e-05
Identities = 30/70 (42%), Positives = 35/70 (49%), Gaps = 18/70 (25%)
Query: 57 PQFQQRPQPPRQPSFT----------------QQPRQQAPR--TRFDPIPMKYAEFLPSL 98
P +QQ P PP QP QQ +QQ PR T F PIPM+YAE LP+L
Sbjct: 130 PFYQQYPLPPGQPPVPVNAMAQQVQHQPPAQQQQQQQQGPRPKTYFPPIPMRYAELLPTL 189
Query: 99 LKGNYVQTRP 108
L + TRP
Sbjct: 190 LAKGHCVTRP 199
>gb|AAU90285.1| putative gag/pol polyprotein, 3'-partial [Solanum demissum]
Length = 1784
Score = 43.9 bits (102), Expect = 0.002
Identities = 28/88 (31%), Positives = 42/88 (46%), Gaps = 6/88 (6%)
Query: 81 RTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLPAGFRADLS--CVFHQGAPGHDVE 138
+ F PI YA L N + P R P D S C + APGH++E
Sbjct: 889 KDEFTPIGESYASLFQKLRTLNVLS---PIERKMSNPPPRNLDYSQHCAYCSDAPGHNIE 945
Query: 139 RCYALRNAVQDLVEA-KILSFTDLNPNL 165
RC+ L+ A+QDL++ +I+ + PN+
Sbjct: 946 RCWYLKKAIQDLIDTRRIIVESPNGPNI 973
>dbj|BAC29735.1| unnamed protein product [Mus musculus]
Length = 604
Score = 43.5 bits (101), Expect = 0.002
Identities = 37/124 (29%), Positives = 61/124 (48%), Gaps = 18/124 (14%)
Query: 6 LSAHMQTLAQTHKMDQ---LEQELHELRKEMTALRAKVEELTNRV-SSLMVTKDLP-QFQ 60
+ H + +AQ + Q L+++LHEL+ ++TAL K + + ++ +L+V ++ P +FQ
Sbjct: 14 IQVHQKLVAQMKQDPQNADLKKQLHELQAKITALSEKQKRVVEQLRKNLIVKQEQPDKFQ 73
Query: 61 QRP------------QPPRQPSFTQQPRQ-QAPRTRFDPIPMKYAEFLPSLLKGNYVQTR 107
+P Q P QP QQP+Q Q + + + A PSL VQ
Sbjct: 74 IQPLSQSENKLQTAQQQPLQPLQQQQPQQPQQQQQQQQQHAQQSAAAPPSLTASQQVQAT 133
Query: 108 PPPP 111
PP P
Sbjct: 134 PPQP 137
>gb|AAH19181.1| PHD finger protein 21A [Mus musculus] gi|20270291|ref|NP_620094.1|
PHD finger protein 21A [Mus musculus]
Length = 556
Score = 43.1 bits (100), Expect = 0.003
Identities = 34/123 (27%), Positives = 60/123 (48%), Gaps = 17/123 (13%)
Query: 6 LSAHMQTLAQTHKMDQ---LEQELHELRKEMTALRAKVEELTNRV-SSLMVTKDLP-QFQ 60
+ H + +AQ + Q L+++LHEL+ ++TAL K + + ++ +L+V ++ P +FQ
Sbjct: 14 IQVHQKLVAQMKQDPQNADLKKQLHELQAKITALSEKQKRVVEQLRKNLIVKQEQPDKFQ 73
Query: 61 QRP------------QPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRP 108
+P Q P QP QQP+Q + + + + P L + VQ P
Sbjct: 74 IQPLSQSENKLQTAQQQPLQPLQQQQPQQPQQQQQQQQQHAQQSAAAPPSLTASQVQATP 133
Query: 109 PPP 111
P P
Sbjct: 134 PQP 136
>dbj|BAC65327.1| PFTF1 [Mus musculus]
Length = 556
Score = 43.1 bits (100), Expect = 0.003
Identities = 34/123 (27%), Positives = 60/123 (48%), Gaps = 17/123 (13%)
Query: 6 LSAHMQTLAQTHKMDQ---LEQELHELRKEMTALRAKVEELTNRV-SSLMVTKDLP-QFQ 60
+ H + +AQ + Q L+++LHEL+ ++TAL K + + ++ +L+V ++ P +FQ
Sbjct: 14 IQVHQKLVAQMKQDPQNADLKKQLHELQAKITALSEKQKRVVEQLRKNLIVKQEQPDKFQ 73
Query: 61 QRP------------QPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRP 108
+P Q P QP QQP+Q + + + + P L + VQ P
Sbjct: 74 IQPLSQSENKLQTAQQQPLQPLQQQQPQQPQQQQQQQQQHAQQSAAAPPSLTASQVQATP 133
Query: 109 PPP 111
P P
Sbjct: 134 PQP 136
>gb|AAC12944.1| TPA inducible protein [Homo sapiens]
Length = 579
Score = 42.7 bits (99), Expect = 0.004
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 176 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 235
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 236 QALEAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 283
Score = 34.7 bits (78), Expect = 1.0
Identities = 28/112 (25%), Positives = 53/112 (47%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL A+ +++ + ++ LPQ Q+
Sbjct: 212 LQRIAQLQLQQQQQQQQQQQQQQQQALEAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 271
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 272 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 318
>gb|AAH13985.1| Positive cofactor 2, glutamine/Q-rich-associated protein [Homo
sapiens] gi|21312134|ref|NP_056973.2| positive cofactor
2, glutamine/Q-rich-associated protein isoform b [Homo
sapiens]
Length = 748
Score = 42.7 bits (99), Expect = 0.004
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 202 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 261
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 262 QALQAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 309
Score = 35.0 bits (79), Expect = 0.78
Identities = 28/112 (25%), Positives = 54/112 (48%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + ++ LPQ Q+
Sbjct: 238 LQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 297
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 298 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 344
>sp|Q96RN5|PCQAP_HUMAN Positive cofactor 2 glutamine/Q-rich-associated protein (PC2
glutamine/Q-rich-associated protein) (TPA-inducible
gene-1) (TIG-1) (Activator-recruited cofactor 105 kDa
component) (ARC105) (CTG repeat protein 7a)
gi|51477700|ref|NP_001003891.1| positive cofactor 2,
glutamine/Q-rich-associated protein isoform a [Homo
sapiens]
Length = 788
Score = 42.7 bits (99), Expect = 0.004
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 202 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 261
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 262 QALQAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 309
Score = 35.0 bits (79), Expect = 0.78
Identities = 28/112 (25%), Positives = 54/112 (48%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + ++ LPQ Q+
Sbjct: 238 LQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 297
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 298 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 344
>gb|AAB91443.1| CTG7a [Homo sapiens]
Length = 349
Score = 42.7 bits (99), Expect = 0.004
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 18 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 77
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 78 QALEAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 125
Score = 34.7 bits (78), Expect = 1.0
Identities = 28/112 (25%), Positives = 53/112 (47%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL A+ +++ + ++ LPQ Q+
Sbjct: 54 LQRIAQLQLQQQQQQQQQQQQQQQQALEAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 113
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 114 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 160
>gb|AAH03078.1| PCQAP protein [Homo sapiens]
Length = 767
Score = 42.7 bits (99), Expect = 0.004
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 221 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 280
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 281 QALQAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 328
Score = 35.0 bits (79), Expect = 0.78
Identities = 28/112 (25%), Positives = 54/112 (48%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + ++ LPQ Q+
Sbjct: 257 LQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 316
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 317 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 363
>dbj|BAC03446.1| FLJ00386 protein [Homo sapiens]
Length = 811
Score = 42.7 bits (99), Expect = 0.004
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 225 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 284
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 285 QALQAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 332
Score = 35.0 bits (79), Expect = 0.78
Identities = 28/112 (25%), Positives = 54/112 (48%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + ++ LPQ Q+
Sbjct: 261 LQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 320
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 321 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 367
>emb|CAH92172.1| hypothetical protein [Pongo pygmaeus]
Length = 566
Score = 40.8 bits (94), Expect = 0.014
Identities = 29/104 (27%), Positives = 50/104 (47%), Gaps = 4/104 (3%)
Query: 15 QTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQ 74
Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +Q Q
Sbjct: 126 QQLQQQQQQQHLIKLHHQNQQQIQQQQQQLQRMAQLQLQQQQQQQQQQQQQQQQALQAQP 185
Query: 75 PRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
P QQ P + P P A+ LP L+ ++ Q PPP+P++ P
Sbjct: 186 PIQQPPMQQPQPPP---AQALPQQLQQMHHTQHHQPPPQPQQPP 226
Score = 34.7 bits (78), Expect = 1.0
Identities = 28/112 (25%), Positives = 53/112 (47%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + + LPQ Q+
Sbjct: 155 LQRMAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPAQALPQQLQQMHHTQ 214
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 215 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 261
>ref|XP_646324.1| hypothetical protein DDB0190596 [Dictyostelium discoideum]
gi|60474327|gb|EAL72264.1| hypothetical protein
DDB0190596 [Dictyostelium discoideum]
Length = 418
Score = 40.8 bits (94), Expect = 0.014
Identities = 21/71 (29%), Positives = 39/71 (54%), Gaps = 1/71 (1%)
Query: 45 NRVSSLMVTKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYV 104
N+ + ++ + + + QQ+PQ P+QP QQP+Q R+ PI ++ + + L N
Sbjct: 254 NKTNPIIEKQQIKELQQQPQQPQQPQQPQQPQQNLIRSTNTPIIVENITPIKNTLTNNST 313
Query: 105 QTRPPPPRPEK 115
+ PP P P++
Sbjct: 314 FS-PPYPEPDQ 323
>emb|CAE62406.1| Hypothetical protein CBG06493 [Caenorhabditis briggsae]
Length = 413
Score = 40.8 bits (94), Expect = 0.014
Identities = 32/125 (25%), Positives = 54/125 (42%), Gaps = 12/125 (9%)
Query: 1 MSSFNLSAHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLP--- 57
MS+ +Q +A +M+QL++E + RKE+ + K E+L NR L +++ P
Sbjct: 184 MSNMAYCTSVQQMASVKQMEQLKKERDQFRKELGRYKTKYEKLENRYEQLGTSQNTPLFG 243
Query: 58 ----QFQQRPQPPRQ---PSFTQQPRQQA--PRTRFDPIPMKYAEFLPSLLKGNYVQTRP 108
F+ + QPP Q P F+ A F P+ L ++ +P
Sbjct: 244 GPGGAFKTQFQPPAQFLTPQFSASSHTFAYPNPPNFKPLSSGGPPSLHHRSMDAFIDIKP 303
Query: 109 PPPRP 113
P +P
Sbjct: 304 PRHQP 308
>emb|CAG30423.1| PCQAP [Homo sapiens]
Length = 674
Score = 40.4 bits (93), Expect = 0.019
Identities = 28/104 (26%), Positives = 50/104 (47%), Gaps = 4/104 (3%)
Query: 15 QTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQ 74
Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +Q Q
Sbjct: 135 QQLQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQQALQAQP 194
Query: 75 PRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 195 PIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 235
Score = 35.0 bits (79), Expect = 0.78
Identities = 28/112 (25%), Positives = 54/112 (48%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + ++ LPQ Q+
Sbjct: 164 LQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 223
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 224 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 270
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 306,356,780
Number of Sequences: 2540612
Number of extensions: 14596348
Number of successful extensions: 105673
Number of sequences better than 10.0: 656
Number of HSP's better than 10.0 without gapping: 116
Number of HSP's successfully gapped in prelim test: 577
Number of HSP's that attempted gapping in prelim test: 103440
Number of HSP's gapped (non-prelim): 1879
length of query: 167
length of database: 863,360,394
effective HSP length: 118
effective length of query: 49
effective length of database: 563,568,178
effective search space: 27614840722
effective search space used: 27614840722
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC144591.1


