

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144541.14 - phase: 0
(63 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
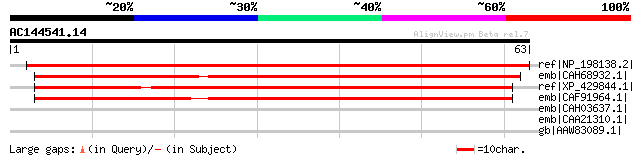
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_198138.2| expressed protein [Arabidopsis thaliana] 100 1e-20
emb|CAH68932.1| novel protein [Danio rerio] gi|66472800|ref|NP_0... 58 6e-08
ref|XP_429844.1| PREDICTED: hypothetical protein XP_429844 [Gall... 55 5e-07
emb|CAF91964.1| unnamed protein product [Tetraodon nigroviridis] 54 1e-06
emb|CAH03637.1| hypothetical protein [Paramecium tetraurelia] gi... 32 3.4
emb|CAA21310.1| SPBC1734.16c [Schizosaccharomyces pombe] gi|1911... 32 4.4
gb|AAW83089.1| YdcA [Neisseria gonorrhoeae] 32 4.4
>ref|NP_198138.2| expressed protein [Arabidopsis thaliana]
Length = 177
Score = 100 bits (249), Expect = 1e-20
Identities = 51/61 (83%), Positives = 55/61 (89%)
Query: 3 GERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASF 62
GE SS PV+LSKFL R+KDDG RRSAVSGKKILLK+DK+KEDK AESKRNELL FLNASF
Sbjct: 117 GEGSSGPVKLSKFLNRDKDDGERRSAVSGKKILLKVDKSKEDKAAESKRNELLKFLNASF 176
Query: 63 D 63
D
Sbjct: 177 D 177
>emb|CAH68932.1| novel protein [Danio rerio] gi|66472800|ref|NP_001018615.1|
hypothetical protein LOC553945 [Danio rerio]
Length = 175
Score = 58.2 bits (139), Expect = 6e-08
Identities = 28/59 (47%), Positives = 43/59 (72%), Gaps = 1/59 (1%)
Query: 4 ERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASF 62
+ SS PVQ+SK+L K G + S +SGKKI +K+ K+K+DK+ + R ELL+FLN+++
Sbjct: 118 DESSGPVQISKYLKEKKKSG-KYSMISGKKIKMKVKKSKKDKQRDKNRAELLDFLNSTY 175
>ref|XP_429844.1| PREDICTED: hypothetical protein XP_429844 [Gallus gallus]
Length = 267
Score = 55.1 bits (131), Expect = 5e-07
Identities = 28/58 (48%), Positives = 39/58 (66%), Gaps = 1/58 (1%)
Query: 4 ERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNAS 61
+ + PVQLSKF +NK S ++GKKI +K+ KTK+DKE + R ELL FLN++
Sbjct: 210 KEKTGPVQLSKFF-KNKKKNENYSMITGKKIKMKIKKTKKDKERDRNRAELLEFLNSA 266
>emb|CAF91964.1| unnamed protein product [Tetraodon nigroviridis]
Length = 160
Score = 53.9 bits (128), Expect = 1e-06
Identities = 28/58 (48%), Positives = 40/58 (68%), Gaps = 2/58 (3%)
Query: 4 ERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNAS 61
E +S PVQ+SK+L K + S +SGKKI +K+ K+KEDK+ + R ELL FLN++
Sbjct: 104 EENSGPVQISKYLKDRKKG--KYSMISGKKIKMKVKKSKEDKKRDKNRAELLEFLNST 159
>emb|CAH03637.1| hypothetical protein [Paramecium tetraurelia]
gi|50405275|ref|YP_054367.1| hypothetical protein
PTMB.436 [Paramecium tetraurelia]
Length = 899
Score = 32.3 bits (72), Expect = 3.4
Identities = 18/49 (36%), Positives = 26/49 (52%), Gaps = 2/49 (4%)
Query: 14 KFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASF 62
K L NK D + S K +L +D+ +D++ S RN L+N N SF
Sbjct: 767 KILKTNKQDS--KILTSDNKRILMVDELDKDQDVSSLRNSLVNLSNISF 813
>emb|CAA21310.1| SPBC1734.16c [Schizosaccharomyces pombe]
gi|19112225|ref|NP_595433.1| hypothetical protein
SPBC1734.16c [Schizosaccharomyces pombe 972h-]
gi|7492269|pir||T39663 paired amphipathic helix,
probable transcription regulator protein - fission
yeast (Schizosaccharomyces pombe)
Length = 1154
Score = 32.0 bits (71), Expect = 4.4
Identities = 17/42 (40%), Positives = 23/42 (54%)
Query: 5 RSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKE 46
+S SPV S G KD+GV V K+IL + T++D E
Sbjct: 35 QSQSPVNTSLHNGDGKDNGVATEPVENKQILSERSVTRDDYE 76
>gb|AAW83089.1| YdcA [Neisseria gonorrhoeae]
Length = 100
Score = 32.0 bits (71), Expect = 4.4
Identities = 18/58 (31%), Positives = 34/58 (58%), Gaps = 1/58 (1%)
Query: 5 RSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDK-EAESKRNELLNFLNAS 61
+++ P + K LG K D +R S + G ++LL+ + T+E++ E + + L+FL S
Sbjct: 13 QTTIPESVRKALGLEKQDKIRFSVLGGGQVLLEKNNTEEEEFEHDPVVGKFLHFLENS 70
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.307 0.129 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 94,508,939
Number of Sequences: 2540612
Number of extensions: 2862170
Number of successful extensions: 8012
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 8005
Number of HSP's gapped (non-prelim): 7
length of query: 63
length of database: 863,360,394
effective HSP length: 39
effective length of query: 24
effective length of database: 764,276,526
effective search space: 18342636624
effective search space used: 18342636624
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144541.14


