

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.12 + phase: 0
(417 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
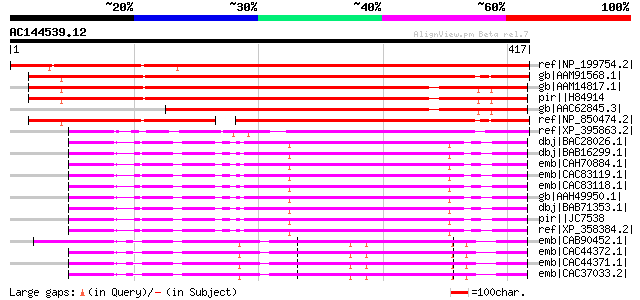
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_199754.2| transducin family protein / WD-40 repeat family... 567 e-160
gb|AAM91568.1| putative WD-40 repeat protein [Arabidopsis thaliana] 540 e-152
gb|AAM14817.1| hypothetical protein [Arabidopsis thaliana] 523 e-147
pir||H84914 probable WD-40 repeat protein [imported] - Arabidops... 523 e-147
gb|AAC62845.3| putative WD-40 repeat protein [Arabidopsis thalia... 410 e-113
ref|NP_850474.2| transducin family protein / WD-40 repeat family... 383 e-105
ref|XP_395863.2| PREDICTED: similar to CG31132-PA [Apis mellifera] 182 2e-44
dbj|BAC28026.1| unnamed protein product [Mus musculus] 179 1e-43
dbj|BAB16299.1| neuronal differentiaion related protein [Mus mus... 179 2e-43
emb|CAH70884.1| OTTHUMP00000016771 [Homo sapiens] gi|55959101|em... 179 2e-43
emb|CAC83119.1| WD repeat domain 11 protein [Mus musculus] 179 2e-43
emb|CAC83118.1| WD repeat domain 11 protein [Homo sapiens] gi|34... 179 2e-43
gb|AAH49950.1| Unknown (protein for IMAGE:4021341) [Mus musculus] 179 2e-43
dbj|BAB71353.1| unnamed protein product [Homo sapiens] 179 2e-43
pir||JC7538 neuronal differentiation-related protein - mouse 179 2e-43
ref|XP_358384.2| PREDICTED: pleckstrin homology domain interacti... 179 2e-43
emb|CAB90452.1| homolog to cAMP response element binding and bet... 178 3e-43
emb|CAC44372.1| WDR9 protein, form B [Homo sapiens] 177 4e-43
emb|CAC44371.1| WDR9 protein, form A [Homo sapiens] 177 4e-43
emb|CAC37033.2| hypothetical protein [Homo sapiens] gi|23831562|... 177 4e-43
>ref|NP_199754.2| transducin family protein / WD-40 repeat family protein
[Arabidopsis thaliana]
Length = 1677
Score = 567 bits (1462), Expect = e-160
Identities = 286/423 (67%), Positives = 338/423 (79%), Gaps = 8/423 (1%)
Query: 1 MALQKYVPSGDAPTVSLKHLS-SSKAPEKTQ---PDVANQNHDMDVDVDIGEVYFLIMRF 56
MAL+K P GD+ ++ +K L+ S K P Q PD+ Q+ ++D+D+ EVYFL++
Sbjct: 1 MALRKNTPKGDSVSLPMKPLNFSRKLPGNVQIPDPDIV-QSVAPNIDLDLREVYFLMLHL 59
Query: 57 LSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERY 116
LS+GPC KT + L +ELLE++LLPRRYHAWYSRSG SG +DDG SFPL +Y +LA+RY
Sbjct: 60 LSSGPCQKTYALLRHELLEHELLPRRYHAWYSRSGLPSGDENDDGNSFPL-NYTELAKRY 118
Query: 117 PHIEKDHLVKLLKQLLL--NKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNE 174
H++KDHLV+LLKQL+ N+ + S G+ GN A VPTLLG GSFSLLS D + V
Sbjct: 119 SHVKKDHLVELLKQLVFVSNRPNPSRGIGDGNKMIGAGVPTLLGTGSFSLLSSDKEIVGS 178
Query: 175 EVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNI 234
++KPPP MRWPH A+QV G+ LREIGGG RHHRAPSIRAACY IAKPS+MVQKMQNI
Sbjct: 179 DLKPPPIGMRWPHMHADQVRGISLREIGGGFARHHRAPSIRAACYVIAKPSSMVQKMQNI 238
Query: 235 KRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSN 294
KR+RGH NAVYCAI DRSGRYVITGSDDRLVK+WSM+TAY LASCRGH GDITDLAVSSN
Sbjct: 239 KRLRGHRNAVYCAILDRSGRYVITGSDDRLVKVWSMDTAYCLASCRGHEGDITDLAVSSN 298
Query: 295 NALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRI 354
N +AS+SND +IRVWRLPDGLP+SVLRGHTGAVTAIAFSPRP + YQLLSSSDDGTCRI
Sbjct: 299 NIFIASASNDCVIRVWRLPDGLPVSVLRGHTGAVTAIAFSPRPGSPYQLLSSSDDGTCRI 358
Query: 355 WDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLAR 414
WDAR PR+YVP+P G++SGPSS+ QSHQIFCCAFNA+G+VFVTGSSD LAR
Sbjct: 359 WDARGAQFAPRIYVPRPPSPDGKNSGPSSSNAQQSHQIFCCAFNASGSVFVTGSSDTLAR 418
Query: 415 VYT 417
VY+
Sbjct: 419 VYS 421
>gb|AAM91568.1| putative WD-40 repeat protein [Arabidopsis thaliana]
Length = 1519
Score = 540 bits (1391), Expect = e-152
Identities = 275/407 (67%), Positives = 320/407 (78%), Gaps = 10/407 (2%)
Query: 16 SLKHLSSSKAPEKTQPDVANQNHDM-----DVDVDIGEVYFLIMRFLSAGPCHKTCSHLW 70
S + S + Q +V Q HD D+D+D+ EVYFLI+ FLS GPC +T HL
Sbjct: 7 STPFMEPSNLAKLVQGNVPLQPHDSHSSLTDLDMDLREVYFLILHFLSIGPCERTFGHLR 66
Query: 71 NELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERYPHIEKDHLVKLLKQ 130
+E+LE LLPRRYH+W+SRSG SG DDG S PL Y+ L ERYPHIEKDHLVKLLKQ
Sbjct: 67 DEILEKGLLPRRYHSWWSRSGIYSGRADDDGISLPLS-YDNLIERYPHIEKDHLVKLLKQ 125
Query: 131 LLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNEEVKPPPPYMRWPHTKA 190
L+LN + S GN PNAADVPTLLG G+FSL+ + +++ + Y+RWPH A
Sbjct: 126 LILNPSFPSHMRVEGNAPNAADVPTLLGSGTFSLVDRSNNIESQKARHVASYLRWPHMHA 185
Query: 191 NQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFD 250
+QV GL LREIGGG +HHRAPSI +AC+AIAKPSTMVQKMQNIK++RGH NAVYCAIFD
Sbjct: 186 DQVRGLSLREIGGGFRKHHRAPSILSACHAIAKPSTMVQKMQNIKKLRGHRNAVYCAIFD 245
Query: 251 RSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVW 310
RSGRYVITGSDDRLVKIWSMETA LASCRGH GDITDLAVSSNNALVAS+SND++IRVW
Sbjct: 246 RSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASASNDFVIRVW 305
Query: 311 RLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPK 370
RLPDG+PISVLRGHTGAVTAIAFSPR +VYQLLSSSDDGTCRIWDARY+ PR+YVP
Sbjct: 306 RLPDGMPISVLRGHTGAVTAIAFSPRQASVYQLLSSSDDGTCRIWDARYSQWLPRIYVPS 365
Query: 371 PSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVYT 417
PSD+ ++G +SN QSHQI CCA+NANGT+FVTGSSD+ ARV++
Sbjct: 366 PSDA---NTGSTSNA-SQSHQILCCAYNANGTIFVTGSSDSNARVWS 408
>gb|AAM14817.1| hypothetical protein [Arabidopsis thaliana]
Length = 544
Score = 523 bits (1348), Expect = e-147
Identities = 273/420 (65%), Positives = 314/420 (74%), Gaps = 27/420 (6%)
Query: 16 SLKHLSSSKAPEKTQPDVANQNHDM-----DVDVDIGEVYFLIMRFLSAGPCHKTCSHLW 70
S + S + Q +V Q HD D+D+D+ EVYFLI+ FLS GPC +T HL
Sbjct: 7 STPFMEPSNLAKLVQGNVPLQPHDSHSSLTDLDMDLREVYFLILHFLSIGPCERTFGHLR 66
Query: 71 NELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERYPHIEKDHLVKLLKQ 130
+E+LE LLPRRYH+W+SRSG SG DDG S PL Y+ L ERYPHIEKDHLVKLLKQ
Sbjct: 67 DEILEKGLLPRRYHSWWSRSGIYSGRADDDGISLPLS-YDNLIERYPHIEKDHLVKLLKQ 125
Query: 131 LLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNEEVKPPPPYMRWPHTKA 190
L+LN + S GN PNAADVPTLLG G+FSL+ + +++ + Y+RWPH A
Sbjct: 126 LILNPSFPSHMRVEGNAPNAADVPTLLGSGTFSLVDRSNNIESQKARHVASYLRWPHMHA 185
Query: 191 NQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFD 250
+QV GL LREIGGG +HHRAPSI +AC+AIAKPSTMVQKMQNIK++RGH NAVYCAIFD
Sbjct: 186 DQVRGLSLREIGGGFRKHHRAPSILSACHAIAKPSTMVQKMQNIKKLRGHRNAVYCAIFD 245
Query: 251 RSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVW 310
RSGRYVITGSDDRLVKIWSMETA LASCRGH GDITDLAVSSNNALVAS+SND++IRVW
Sbjct: 246 RSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASASNDFVIRVW 305
Query: 311 RLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPK 370
RLPDG+PISVLRGHTGAVTAIAFSPR +SSDDGTCRIWDARY+ PR+YVP
Sbjct: 306 RLPDGMPISVLRGHTGAVTAIAFSPRQ-------ASSDDGTCRIWDARYSQWLPRIYVPS 358
Query: 371 PSDST---------GRSSGPSSNT-----MPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
PSD+ G S+ SSNT QSHQI CCA+NANGT+FVTGSSD+ ARVY
Sbjct: 359 PSDANTGSLEVSLLGESTFTSSNTGSTSNASQSHQILCCAYNANGTIFVTGSSDSNARVY 418
>pir||H84914 probable WD-40 repeat protein [imported] - Arabidopsis thaliana
Length = 1389
Score = 523 bits (1348), Expect = e-147
Identities = 273/420 (65%), Positives = 314/420 (74%), Gaps = 27/420 (6%)
Query: 16 SLKHLSSSKAPEKTQPDVANQNHDM-----DVDVDIGEVYFLIMRFLSAGPCHKTCSHLW 70
S + S + Q +V Q HD D+D+D+ EVYFLI+ FLS GPC +T HL
Sbjct: 7 STPFMEPSNLAKLVQGNVPLQPHDSHSSLTDLDMDLREVYFLILHFLSIGPCERTFGHLR 66
Query: 71 NELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERYPHIEKDHLVKLLKQ 130
+E+LE LLPRRYH+W+SRSG SG DDG S PL Y+ L ERYPHIEKDHLVKLLKQ
Sbjct: 67 DEILEKGLLPRRYHSWWSRSGIYSGRADDDGISLPLS-YDNLIERYPHIEKDHLVKLLKQ 125
Query: 131 LLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNEEVKPPPPYMRWPHTKA 190
L+LN + S GN PNAADVPTLLG G+FSL+ + +++ + Y+RWPH A
Sbjct: 126 LILNPSFPSHMRVEGNAPNAADVPTLLGSGTFSLVDRSNNIESQKARHVASYLRWPHMHA 185
Query: 191 NQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFD 250
+QV GL LREIGGG +HHRAPSI +AC+AIAKPSTMVQKMQNIK++RGH NAVYCAIFD
Sbjct: 186 DQVRGLSLREIGGGFRKHHRAPSILSACHAIAKPSTMVQKMQNIKKLRGHRNAVYCAIFD 245
Query: 251 RSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVW 310
RSGRYVITGSDDRLVKIWSMETA LASCRGH GDITDLAVSSNNALVAS+SND++IRVW
Sbjct: 246 RSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASASNDFVIRVW 305
Query: 311 RLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPK 370
RLPDG+PISVLRGHTGAVTAIAFSPR +SSDDGTCRIWDARY+ PR+YVP
Sbjct: 306 RLPDGMPISVLRGHTGAVTAIAFSPRQ-------ASSDDGTCRIWDARYSQWLPRIYVPS 358
Query: 371 PSDST---------GRSSGPSSNT-----MPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
PSD+ G S+ SSNT QSHQI CCA+NANGT+FVTGSSD+ ARVY
Sbjct: 359 PSDANTGSLEVSLLGESTFTSSNTGSTSNASQSHQILCCAYNANGTIFVTGSSDSNARVY 418
>gb|AAC62845.3| putative WD-40 repeat protein [Arabidopsis thaliana]
gi|7487572|pir||T00443 hypothetical protein T30B22.29 -
Arabidopsis thaliana (fragment)
Length = 1269
Score = 410 bits (1053), Expect = e-113
Identities = 211/305 (69%), Positives = 239/305 (78%), Gaps = 21/305 (6%)
Query: 126 KLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNEEVKPPPPYMRW 185
KLLKQL+LN + S GN PNAADVPTLLG G+FSL+ + +++ + Y+RW
Sbjct: 1 KLLKQLILNPSFPSHMRVEGNAPNAADVPTLLGSGTFSLVDRSNNIESQKARHVASYLRW 60
Query: 186 PHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHCNAVY 245
PH A+QV GL LREIGGG +HHRAPSI +AC+AIAKPSTMVQKMQNIK++RGH NAVY
Sbjct: 61 PHMHADQVRGLSLREIGGGFRKHHRAPSILSACHAIAKPSTMVQKMQNIKKLRGHRNAVY 120
Query: 246 CAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDY 305
CAIFDRSGRYVITGSDDRLVKIWSMETA LASCRGH GDITDLAVSSNNALVAS+SND+
Sbjct: 121 CAIFDRSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASASNDF 180
Query: 306 IIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTHSTPR 365
+IRVWRLPDG+PISVLRGHTGAVTAIAFSPR +SSDDGTCRIWDARY+ PR
Sbjct: 181 VIRVWRLPDGMPISVLRGHTGAVTAIAFSPRQ-------ASSDDGTCRIWDARYSQWLPR 233
Query: 366 LYVPKPSDST---------GRSSGPSSNT-----MPQSHQIFCCAFNANGTVFVTGSSDN 411
+YVP PSD+ G S+ SSNT QSHQI CCA+NANGT+FVTGSSD+
Sbjct: 234 IYVPSPSDANTGSLEVSLLGESTFTSSNTGSTSNASQSHQILCCAYNANGTIFVTGSSDS 293
Query: 412 LARVY 416
ARVY
Sbjct: 294 NARVY 298
>ref|NP_850474.2| transducin family protein / WD-40 repeat family protein
[Arabidopsis thaliana]
Length = 1589
Score = 383 bits (984), Expect = e-105
Identities = 187/236 (79%), Positives = 210/236 (88%), Gaps = 4/236 (1%)
Query: 182 YMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAKPSTMVQKMQNIKRIRGHC 241
Y+RWPH A+QV GL LREIGGG +HHRAPSI +AC+AIAKPSTMVQKMQNIK++RGH
Sbjct: 246 YLRWPHMHADQVRGLSLREIGGGFRKHHRAPSILSACHAIAKPSTMVQKMQNIKKLRGHR 305
Query: 242 NAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASS 301
NAVYCAIFDRSGRYVITGSDDRLVKIWSMETA LASCRGH GDITDLAVSSNNALVAS+
Sbjct: 306 NAVYCAIFDRSGRYVITGSDDRLVKIWSMETALCLASCRGHEGDITDLAVSSNNALVASA 365
Query: 302 SNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDDGTCRIWDARYTH 361
SND++IRVWRLPDG+PISVLRGHTGAVTAIAFSPR +VYQLLSSSDDGTCRIWDARY+
Sbjct: 366 SNDFVIRVWRLPDGMPISVLRGHTGAVTAIAFSPRQASVYQLLSSSDDGTCRIWDARYSQ 425
Query: 362 STPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVYT 417
PR+YVP PSD+ ++G +SN QSHQI CCA+NANGT+FVTGSSD+ ARV++
Sbjct: 426 WLPRIYVPSPSDA---NTGSTSNA-SQSHQILCCAYNANGTIFVTGSSDSNARVWS 477
Score = 163 bits (413), Expect = 8e-39
Identities = 88/155 (56%), Positives = 105/155 (66%), Gaps = 6/155 (3%)
Query: 16 SLKHLSSSKAPEKTQPDVANQNHDM-----DVDVDIGEVYFLIMRFLSAGPCHKTCSHLW 70
S + S + Q +V Q HD D+D+D+ EVYFLI+ FLS GPC +T HL
Sbjct: 7 STPFMEPSNLAKLVQGNVPLQPHDSHSSLTDLDMDLREVYFLILHFLSIGPCERTFGHLR 66
Query: 71 NELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERYPHIEKDHLVKLLKQ 130
+E+LE LLPRRYH+W+SRSG SG DDG S PL Y+ L ERYPHIEKDHLVKLLKQ
Sbjct: 67 DEILEKGLLPRRYHSWWSRSGIYSGRADDDGISLPL-SYDNLIERYPHIEKDHLVKLLKQ 125
Query: 131 LLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLL 165
L+LN + S GN PNAADVPTLLG G+FSL+
Sbjct: 126 LILNPSFPSHMRVEGNAPNAADVPTLLGSGTFSLV 160
>ref|XP_395863.2| PREDICTED: similar to CG31132-PA [Apis mellifera]
Length = 1732
Score = 182 bits (461), Expect = 2e-44
Identities = 127/382 (33%), Positives = 186/382 (48%), Gaps = 54/382 (14%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI +FL+AGPC + L EL ++LP+R W H Q+F
Sbjct: 15 ELYFLIAKFLAAGPCREAADVLKRELERAKVLPQRLD-WEG---------HIHNQTF--- 61
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
+L ++Y HI +HL+++ A + P + PP + +LLG G SLL
Sbjct: 62 --EELEKKYSHIGSNHLLQIC-------ARIGPVLEKEVPPCISGAISLLGAGRQSLLRT 112
Query: 168 DGDKVNEEVKP---------PPPYMRWPHTKA--NQVHGLHLREIGGGLPRHHRAPSIRA 216
D V +V P++ P++ N VH L RE G L R
Sbjct: 113 HED-VQRQVHTITSYSGRLGGKPFLDSPYSLLVPNIVHVLQGRENSGPLSRRE------- 164
Query: 217 ACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSL 276
A P+ KMQ + GH +AVYC +FDR+G+Y+ITG+DD LVK+WS L
Sbjct: 165 -----AIPTKFYNKMQLYRHTLGHLSAVYCVLFDRTGKYIITGADDLLVKVWSSIDGRLL 219
Query: 277 ASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP- 335
A+ RG +I D+AV+ +N L+A+ S D I+RVW P++VL GH+G +T++ F P
Sbjct: 220 ATFRGASAEIMDIAVNFDNTLLAAGSLDRILRVWCFQTMSPVAVLSGHSGMITSVNFCPI 279
Query: 336 RPNAVYQLLSSSDDGTCRIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFCC 395
N VY L+S+S DG+ W + ++ PKP P Q+ C
Sbjct: 280 SCNGVYYLVSTSTDGSVAFWSHTKKGNERAIFQPKPIQY-------HEKMRPGQAQMICA 332
Query: 396 AFNANGTVFVTGSSDNLARVYT 417
+F+ G GS+D+ RVY+
Sbjct: 333 SFSPGGAFMAAGSADHHVRVYS 354
Score = 39.3 bits (90), Expect = 0.23
Identities = 31/143 (21%), Positives = 53/143 (36%), Gaps = 18/143 (12%)
Query: 234 IKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSL----------------A 277
+ + H + V + +G I+GS D +W E +
Sbjct: 364 VLEVEAHSDTVDSIQWAHNGLKFISGSKDGTANVWHFEQQQWMYKRLLMTTKLPGESETE 423
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+T ++ ++ V ++ ND ++VW G + +LRGH V + P
Sbjct: 424 DDSNKKAKVTMVSWDVSDEWVITAVNDSCLKVWNAKSGELVKILRGHKDEVFVLESHPID 483
Query: 338 NAVYQLLSSSDDGTCRIWDARYT 360
V +LS+ DG IWD T
Sbjct: 484 PRV--ILSAGHDGQLIIWDVLNT 504
>dbj|BAC28026.1| unnamed protein product [Mus musculus]
Length = 755
Score = 179 bits (455), Expect = 1e-43
Identities = 129/382 (33%), Positives = 189/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+++ LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWAIDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PTKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 40.4 bits (93), Expect = 0.10
Identities = 43/202 (21%), Positives = 72/202 (35%), Gaps = 20/202 (9%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD-ARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIF 393
P P L S+ DG +WD AR + + G + P
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWDLARGVKVRSYFNMVFQIEGQGHGAVFDCKCSPDGQHFA 533
Query: 394 CCAFNANGTVFVTGSSDNLARV 415
C + + +F GSS ++
Sbjct: 534 CTDSHGHLLIFGFGSSSKYDKI 555
>dbj|BAB16299.1| neuronal differentiaion related protein [Mus musculus]
Length = 1019
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PTKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>emb|CAH70884.1| OTTHUMP00000016771 [Homo sapiens] gi|55959101|emb|CAI14406.1|
OTTHUMP00000016771 [Homo sapiens]
Length = 1821
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PAKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>emb|CAC83119.1| WD repeat domain 11 protein [Mus musculus]
Length = 1779
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PTKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>emb|CAC83118.1| WD repeat domain 11 protein [Homo sapiens]
gi|34996489|ref|NP_060404.3| pleckstrin homology domain
interacting protein [Homo sapiens]
Length = 1821
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PAKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>gb|AAH49950.1| Unknown (protein for IMAGE:4021341) [Mus musculus]
Length = 896
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PTKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>dbj|BAB71353.1| unnamed protein product [Homo sapiens]
Length = 912
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PAKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>pir||JC7538 neuronal differentiation-related protein - mouse
Length = 1019
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PTKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>ref|XP_358384.2| PREDICTED: pleckstrin homology domain interacting protein [Mus
musculus]
Length = 1821
Score = 179 bits (453), Expect = 2e-43
Identities = 129/382 (33%), Positives = 188/382 (48%), Gaps = 60/382 (15%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI RFL GPC + L E+ E +LLPRR W G+ P
Sbjct: 14 ELYFLIARFLEDGPCQQAAQVLIREVAEKELLPRRTD-W-------------TGKEHP-R 58
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y L + Y H+ DHL+++ +L P + P + V TLLG G SLL
Sbjct: 59 TYQNLVKYYRHLAPDHLLQICHRL-------GPLLEQEIPQSVPGVQTLLGAGRQSLL-- 109
Query: 168 DGDKVNEEVKPPPPYMRWPHTKANQVHGLHLREIGGGLPRHHRAPSIRAACYAIAK---- 223
+ N+ K ++ W K + + LH + PSI ++
Sbjct: 110 ---RTNKSCK----HVVW---KGSALAALHCGRPPESPVNYGSPPSIADTLFSRKLNGKY 159
Query: 224 ------PSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLA 277
P+ + Q M+ KRI GH ++VYC FDR+GR + TGSDD LVKIW+ + LA
Sbjct: 160 RLERLVPTAVYQHMKMHKRILGHLSSVYCVTFDRTGRRIFTGSDDCLVKIWATDDGRLLA 219
Query: 278 SCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRP 337
+ RGH +I+D+AV+ N ++A+ S D +IRVW L P++VL+GH+ ++T++ FSP
Sbjct: 220 TLRGHAAEISDMAVNYENTMIAAGSCDKMIRVWCLRTCAPLAVLQGHSASITSLQFSPLC 279
Query: 338 NAVYQLLSSSD-DGTC--RIWDARYTHSTPRLYVPKPSDSTGRSSGPSSNTMPQSHQIFC 394
+ + LSS+ DGT +WDA PR P+ T R Q+ C
Sbjct: 280 SGSKRYLSSTGADGTICFWLWDAGTLKINPR-----PTKFTERPR--------PGVQMIC 326
Query: 395 CAFNANGTVFVTGSSDNLARVY 416
+F+A G TGS+D++ RVY
Sbjct: 327 SSFSAGGMFLATGSTDHIIRVY 348
Score = 43.9 bits (102), Expect = 0.009
Identities = 28/66 (42%), Positives = 36/66 (54%), Gaps = 5/66 (7%)
Query: 292 SSNNALVASSSNDYIIRVWRLPDGLP--ISVLRGHTGAVTAIAFSPRPNAVYQLLSSSDD 349
S+ +A+ S D+IIRV+ G P IS L HT V +I FS N + +S S D
Sbjct: 330 SAGGMFLATGSTDHIIRVYFFGSGQPEKISELEFHTDKVDSIQFS---NTSNRFVSGSRD 386
Query: 350 GTCRIW 355
GT RIW
Sbjct: 387 GTARIW 392
Score = 39.7 bits (91), Expect = 0.17
Identities = 34/142 (23%), Positives = 55/142 (37%), Gaps = 19/142 (13%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWS------------METAYSLASC 279
+ I + H + V F + ++GS D +IW M T + +
Sbjct: 356 EKISELEFHTDKVDSIQFSNTSNRFVSGSRDGTARIWQFKRREWKSILLDMATRPAGQNL 415
Query: 280 RGHVGDITDLAVSS-----NNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFS 334
+G IT + V+ ++ V ++ N+ ++VW G I VL GH V
Sbjct: 416 QGIEDKITKMKVTMVAWDRHDNTVITAVNNMTLKVWNSYTGQLIHVLMGHEDEV--FVLE 473
Query: 335 PRPNAVYQLLSSSDDGTCRIWD 356
P P L S+ DG +WD
Sbjct: 474 PHPFDPRVLFSAGHDGNVIVWD 495
>emb|CAB90452.1| homolog to cAMP response element binding and beta transducin family
proteins [Homo sapiens]
Length = 2295
Score = 178 bits (451), Expect = 3e-43
Identities = 133/408 (32%), Positives = 197/408 (47%), Gaps = 55/408 (13%)
Query: 20 LSSSKAPEKTQPDVANQNHDM-DVDVDIGEVYFLIMRFLSAGPCHKTCSHLWNELLENQL 78
L+ S P +P +A + V + E+YFLI R+LSAGPC + L EL + QL
Sbjct: 14 LARSLPPPAPRPAMAEPSSARRPVPLIESELYFLIARYLSAGPCRRAAQVLVQELEQYQL 73
Query: 79 LPRRYHAWYSRSGASSGVPHDDGQSFPLEDYNKLAERYPHIEKDHLVKLLKQLLLNKASL 138
LP+R W G H+ Y +L H+ DHL+++ +++
Sbjct: 74 LPKRLD-W-------EGNEHN-------RSYEELVLSNKHVAPDHLLQICQRI------- 111
Query: 139 SPGMSTGNPPNAADVPTLLGRGSFSLLSYDGDKVNEEVKPPPPYM----RWPHTKANQVH 194
P + PP+ + V +LLG G SLL D + K R P N
Sbjct: 112 GPMLDKEIPPSISRVTSLLGAGRQSLLRTAKDCRHTVWKGSAFAALHRGRPPEMPVNYGS 171
Query: 195 GLHLREIGGGLPRHHRAPSIRA-ACYAIAKPSTMVQKMQNIKRIRGHCNAVYCAIFDRSG 253
+L EI HR + + ++ A P TM Q ++ +RI GH +AVYC FDR+G
Sbjct: 172 PPNLVEI-------HRGKQLTGCSTFSTAFPGTMYQHIKMHRRILGHLSAVYCVAFDRTG 224
Query: 254 RYVITGSDDRLVKIWSMETAYSLASCRGHVGDITDLAVSSNNALVASSSNDYIIRVWRLP 313
+ TGSDD LVKIWS L++ RGH +I+D+AV+ N ++A+ S D IIRVW L
Sbjct: 225 HRIFTGSDDCLVKIWSTHNGRLLSTLRGHSAEISDMAVNYENTMIAAGSCDKIIRVWCLR 284
Query: 314 DGLPISVLRGHTGAVTAIAFSPRPNAVYQ-LLSSSDDGTCRI--WDARYTHSTPR--LYV 368
P++VL+GHTG++T++ FSP + ++S+ DGT WD +PR +
Sbjct: 285 TCAPVAVLQGHTGSITSLQFSPMAKGSQRYMVSTGADGTVCFWQWDLESLKFSPRPLKFT 344
Query: 369 PKPSDSTGRSSGPSSNTMPQSHQIFCCAFNANGTVFVTGSSDNLARVY 416
KP Q+ C +F+ G TGS+D++ R+Y
Sbjct: 345 EKPRPGV---------------QMLCSSFSVGGMFLATGSTDHVIRMY 377
Score = 48.5 bits (114), Expect = 4e-04
Identities = 35/141 (24%), Positives = 62/141 (43%), Gaps = 18/141 (12%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMET----AYSLASCRGHVGD-- 285
+ I + H + V F +G ++GS D +IW E + L GD
Sbjct: 385 EKIAELESHTDKVDSIQFCNNGDRFLSGSRDGTARIWRFEQLEWRSILLDMATRISGDLS 444
Query: 286 ----------ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
+T +A + N+++V ++ ND++++VW G + L GH V + P
Sbjct: 445 SEEERFMKPKVTMIAWNQNDSIVVTAVNDHVLKVWNSYTGQLLHNLMGHADEVFVLETHP 504
Query: 336 RPNAVYQLLSSSDDGTCRIWD 356
+ + +LS+ DG+ IWD
Sbjct: 505 FDSRI--MLSAGHDGSIFIWD 523
Score = 36.6 bits (83), Expect = 1.5
Identities = 17/68 (25%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Query: 246 CAIFDRSGRYVITGSDDRLVKIWSM--ETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
C+ F G ++ TGS D +++++ + E +A H + + +N S S
Sbjct: 355 CSSFSVGGMFLATGSTDHVIRMYFLGFEAPEKIAELESHTDKVDSIQFCNNGDRFLSGSR 414
Query: 304 DYIIRVWR 311
D R+WR
Sbjct: 415 DGTARIWR 422
>emb|CAC44372.1| WDR9 protein, form B [Homo sapiens]
Length = 2269
Score = 177 bits (450), Expect = 4e-43
Identities = 127/379 (33%), Positives = 186/379 (48%), Gaps = 54/379 (14%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI R+LSAGPC + L EL + QLLP+R W G H+
Sbjct: 17 ELYFLIARYLSAGPCRRAAQVLVQELEQYQLLPKRLD-W-------EGNEHN-------R 61
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y +L H+ DHL+++ +++ P + PP+ + V +LLG G SLL
Sbjct: 62 SYEELVLSNKHVAPDHLLQICQRI-------GPMLDKEIPPSISRVTSLLGAGRQSLLRT 114
Query: 168 DGDKVNEEVKPPPPYM----RWPHTKANQVHGLHLREIGGGLPRHHRAPSIRA-ACYAIA 222
D + K R P N +L EI HR + + ++ A
Sbjct: 115 AKDCRHTVWKGSAFAALHRGRPPEMPVNYGSPPNLVEI-------HRGKQLTGCSTFSTA 167
Query: 223 KPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGH 282
P TM Q ++ +RI GH +AVYC FDR+G + TGSDD LVKIWS L++ RGH
Sbjct: 168 FPGTMYQHIKMHRRILGHLSAVYCVAFDRTGHRIFTGSDDCLVKIWSTHNGRLLSTLRGH 227
Query: 283 VGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQ 342
+I+D+AV+ N ++A+ S D IIRVW L P++VL+GHTG++T++ FSP +
Sbjct: 228 SAEISDMAVNYENTMIAAGSCDKIIRVWCLRTCAPVAVLQGHTGSITSLQFSPMAKGSQR 287
Query: 343 -LLSSSDDGTCRI--WDARYTHSTPR--LYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAF 397
++S+ DGT WD +PR + KP Q+ C +F
Sbjct: 288 YMVSTGADGTVCFWQWDLESLKFSPRPLKFTEKPRPGV---------------QMLCSSF 332
Query: 398 NANGTVFVTGSSDNLARVY 416
+ G TGS+D++ R+Y
Sbjct: 333 SVGGMFLATGSTDHVIRMY 351
Score = 48.5 bits (114), Expect = 4e-04
Identities = 35/141 (24%), Positives = 62/141 (43%), Gaps = 18/141 (12%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMET----AYSLASCRGHVGD-- 285
+ I + H + V F +G ++GS D +IW E + L GD
Sbjct: 359 EKIAELESHTDKVDSIQFCNNGDRFLSGSRDGTARIWRFEQLEWRSILLDMATRISGDLS 418
Query: 286 ----------ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
+T +A + N+++V ++ ND++++VW G + L GH V + P
Sbjct: 419 SEEERFMKPKVTMIAWNQNDSIVVTAVNDHVLKVWNSYTGQLLHNLMGHADEVFVLETHP 478
Query: 336 RPNAVYQLLSSSDDGTCRIWD 356
+ + +LS+ DG+ IWD
Sbjct: 479 FDSRI--MLSAGHDGSIFIWD 497
Score = 36.6 bits (83), Expect = 1.5
Identities = 17/68 (25%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Query: 246 CAIFDRSGRYVITGSDDRLVKIWSM--ETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
C+ F G ++ TGS D +++++ + E +A H + + +N S S
Sbjct: 329 CSSFSVGGMFLATGSTDHVIRMYFLGFEAPEKIAELESHTDKVDSIQFCNNGDRFLSGSR 388
Query: 304 DYIIRVWR 311
D R+WR
Sbjct: 389 DGTARIWR 396
>emb|CAC44371.1| WDR9 protein, form A [Homo sapiens]
Length = 2320
Score = 177 bits (450), Expect = 4e-43
Identities = 127/379 (33%), Positives = 186/379 (48%), Gaps = 54/379 (14%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI R+LSAGPC + L EL + QLLP+R W G H+
Sbjct: 17 ELYFLIARYLSAGPCRRAAQVLVQELEQYQLLPKRLD-W-------EGNEHN-------R 61
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y +L H+ DHL+++ +++ P + PP+ + V +LLG G SLL
Sbjct: 62 SYEELVLSNKHVAPDHLLQICQRI-------GPMLDKEIPPSISRVTSLLGAGRQSLLRT 114
Query: 168 DGDKVNEEVKPPPPYM----RWPHTKANQVHGLHLREIGGGLPRHHRAPSIRA-ACYAIA 222
D + K R P N +L EI HR + + ++ A
Sbjct: 115 AKDCRHTVWKGSAFAALHRGRPPEMPVNYGSPPNLVEI-------HRGKQLTGCSTFSTA 167
Query: 223 KPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGH 282
P TM Q ++ +RI GH +AVYC FDR+G + TGSDD LVKIWS L++ RGH
Sbjct: 168 FPGTMYQHIKMHRRILGHLSAVYCVAFDRTGHRIFTGSDDCLVKIWSTHNGRLLSTLRGH 227
Query: 283 VGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQ 342
+I+D+AV+ N ++A+ S D IIRVW L P++VL+GHTG++T++ FSP +
Sbjct: 228 SAEISDMAVNYENTMIAAGSCDKIIRVWCLRTCAPVAVLQGHTGSITSLQFSPMAKGSQR 287
Query: 343 -LLSSSDDGTCRI--WDARYTHSTPR--LYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAF 397
++S+ DGT WD +PR + KP Q+ C +F
Sbjct: 288 YMVSTGADGTVCFWQWDLESLKFSPRPLKFTEKPRPGV---------------QMLCSSF 332
Query: 398 NANGTVFVTGSSDNLARVY 416
+ G TGS+D++ R+Y
Sbjct: 333 SVGGMFLATGSTDHVIRMY 351
Score = 48.5 bits (114), Expect = 4e-04
Identities = 35/141 (24%), Positives = 62/141 (43%), Gaps = 18/141 (12%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMET----AYSLASCRGHVGD-- 285
+ I + H + V F +G ++GS D +IW E + L GD
Sbjct: 359 EKIAELESHTDKVDSIQFCNNGDRFLSGSRDGTARIWRFEQLEWRSILLDMATRISGDLS 418
Query: 286 ----------ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
+T +A + N+++V ++ ND++++VW G + L GH V + P
Sbjct: 419 SEEERFMKPKVTMIAWNQNDSIVVTAVNDHVLKVWNSYTGQLLHNLMGHADEVFVLETHP 478
Query: 336 RPNAVYQLLSSSDDGTCRIWD 356
+ + +LS+ DG+ IWD
Sbjct: 479 FDSRI--MLSAGHDGSIFIWD 497
Score = 36.6 bits (83), Expect = 1.5
Identities = 17/68 (25%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Query: 246 CAIFDRSGRYVITGSDDRLVKIWSM--ETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
C+ F G ++ TGS D +++++ + E +A H + + +N S S
Sbjct: 329 CSSFSVGGMFLATGSTDHVIRMYFLGFEAPEKIAELESHTDKVDSIQFCNNGDRFLSGSR 388
Query: 304 DYIIRVWR 311
D R+WR
Sbjct: 389 DGTARIWR 396
>emb|CAC37033.2| hypothetical protein [Homo sapiens]
gi|23831562|sp|Q9NSI6|WDR9_HUMAN WD-repeat protein 9
Length = 2269
Score = 177 bits (450), Expect = 4e-43
Identities = 127/379 (33%), Positives = 186/379 (48%), Gaps = 54/379 (14%)
Query: 48 EVYFLIMRFLSAGPCHKTCSHLWNELLENQLLPRRYHAWYSRSGASSGVPHDDGQSFPLE 107
E+YFLI R+LSAGPC + L EL + QLLP+R W G H+
Sbjct: 17 ELYFLIARYLSAGPCRRAAQVLVQELEQYQLLPKRLD-W-------EGNEHN-------R 61
Query: 108 DYNKLAERYPHIEKDHLVKLLKQLLLNKASLSPGMSTGNPPNAADVPTLLGRGSFSLLSY 167
Y +L H+ DHL+++ +++ P + PP+ + V +LLG G SLL
Sbjct: 62 SYEELVLSNKHVAPDHLLQICQRI-------GPMLDKEIPPSISRVTSLLGAGRQSLLRT 114
Query: 168 DGDKVNEEVKPPPPYM----RWPHTKANQVHGLHLREIGGGLPRHHRAPSIRA-ACYAIA 222
D + K R P N +L EI HR + + ++ A
Sbjct: 115 AKDCRHTVWKGSAFAALHRGRPPEMPVNYGSPPNLVEI-------HRGKQLTGCSTFSTA 167
Query: 223 KPSTMVQKMQNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMETAYSLASCRGH 282
P TM Q ++ +RI GH +AVYC FDR+G + TGSDD LVKIWS L++ RGH
Sbjct: 168 FPGTMYQHIKMHRRILGHLSAVYCVAFDRTGHRIFTGSDDCLVKIWSTHNGRLLSTLRGH 227
Query: 283 VGDITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSPRPNAVYQ 342
+I+D+AV+ N ++A+ S D IIRVW L P++VL+GHTG++T++ FSP +
Sbjct: 228 SAEISDMAVNYENTMIAAGSCDKIIRVWCLRTCAPVAVLQGHTGSITSLQFSPMAKGSQR 287
Query: 343 -LLSSSDDGTCRI--WDARYTHSTPR--LYVPKPSDSTGRSSGPSSNTMPQSHQIFCCAF 397
++S+ DGT WD +PR + KP Q+ C +F
Sbjct: 288 YMVSTGADGTVCFWQWDLESLKFSPRPLKFTEKPRPGV---------------QMLCSSF 332
Query: 398 NANGTVFVTGSSDNLARVY 416
+ G TGS+D++ R+Y
Sbjct: 333 SVGGMFLATGSTDHVIRMY 351
Score = 48.5 bits (114), Expect = 4e-04
Identities = 35/141 (24%), Positives = 62/141 (43%), Gaps = 18/141 (12%)
Query: 232 QNIKRIRGHCNAVYCAIFDRSGRYVITGSDDRLVKIWSMET----AYSLASCRGHVGD-- 285
+ I + H + V F +G ++GS D +IW E + L GD
Sbjct: 359 EKIAELESHTDKVDSIQFCNNGDRFLSGSRDGTARIWRFEQLEWRSILLDMATRISGDLS 418
Query: 286 ----------ITDLAVSSNNALVASSSNDYIIRVWRLPDGLPISVLRGHTGAVTAIAFSP 335
+T +A + N+++V ++ ND++++VW G + L GH V + P
Sbjct: 419 SEEERFMKPKVTMIAWNQNDSIVVTAVNDHVLKVWNSYTGQLLHNLMGHADEVFVLETHP 478
Query: 336 RPNAVYQLLSSSDDGTCRIWD 356
+ + +LS+ DG+ IWD
Sbjct: 479 FDSRI--MLSAGHDGSIFIWD 497
Score = 36.6 bits (83), Expect = 1.5
Identities = 17/68 (25%), Positives = 31/68 (45%), Gaps = 2/68 (2%)
Query: 246 CAIFDRSGRYVITGSDDRLVKIWSM--ETAYSLASCRGHVGDITDLAVSSNNALVASSSN 303
C+ F G ++ TGS D +++++ + E +A H + + +N S S
Sbjct: 329 CSSFSVGGMFLATGSTDHVIRMYFLGFEAPEKIAELESHTDKVDSIQFCNNGDRFLSGSR 388
Query: 304 DYIIRVWR 311
D R+WR
Sbjct: 389 DGTARIWR 396
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.318 0.134 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 788,855,981
Number of Sequences: 2540612
Number of extensions: 35488880
Number of successful extensions: 123106
Number of sequences better than 10.0: 5359
Number of HSP's better than 10.0 without gapping: 2951
Number of HSP's successfully gapped in prelim test: 2468
Number of HSP's that attempted gapping in prelim test: 88331
Number of HSP's gapped (non-prelim): 22628
length of query: 417
length of database: 863,360,394
effective HSP length: 131
effective length of query: 286
effective length of database: 530,540,222
effective search space: 151734503492
effective search space used: 151734503492
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC144539.12


