

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144515.13 + phase: 0
(381 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
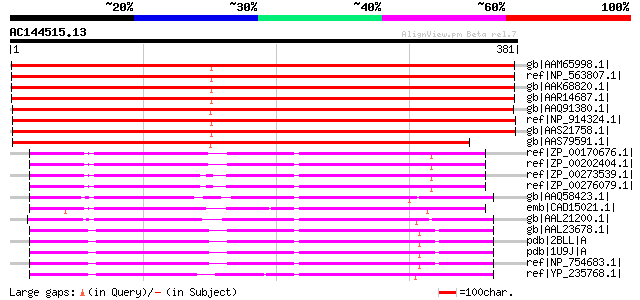
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM65998.1| putative dTDP-glucose 4-6-dehydratase [Arabidopsi... 603 e-171
ref|NP_563807.1| expressed protein [Arabidopsis thaliana] gi|254... 603 e-171
gb|AAK68820.1| similar to dihydroflavonol reductase [Arabidopsis... 597 e-169
gb|AAR14687.1| UDP-D-apiose/UDP-D-xylose synthase [Arabidopsis t... 593 e-168
gb|AAQ91380.1| putative nucleoside-diphosphate-sugar epimerase/d... 587 e-166
ref|NP_914324.1| OJ1656_A11.18 [Oryza sativa (japonica cultivar-... 586 e-166
gb|AAS21758.1| dTDP-glucose 4,6-dehydratase [Zea mays] 572 e-162
gb|AAS79591.1| putative dihydroflavonol reductase [Ipomoea trifida] 553 e-156
ref|ZP_00170676.1| COG0451: Nucleoside-diphosphate-sugar epimera... 233 7e-60
ref|ZP_00202404.1| COG0451: Nucleoside-diphosphate-sugar epimera... 233 7e-60
ref|ZP_00273539.1| COG0451: Nucleoside-diphosphate-sugar epimera... 231 2e-59
ref|ZP_00276079.1| COG0451: Nucleoside-diphosphate-sugar epimera... 231 3e-59
gb|AAQ58423.1| probable transformylase [Chromobacterium violaceu... 224 4e-57
emb|CAD15021.1| PUTATIVE OXIDOREDUCTASE PROTEIN [Ralstonia solan... 218 2e-55
gb|AAL21200.1| putative transformylase [Salmonella typhimurium L... 218 3e-55
gb|AAL23678.1| UDP-D-glucuronate dehydrogenase [Escherichia coli... 216 7e-55
pdb|2BLL|A Chain A, Apo-Structure Of The C-Terminal Decarboxylas... 216 7e-55
pdb|1U9J|A Chain A, Crystal Structure Of E. Coli Arna (Pmri) Dec... 216 7e-55
ref|NP_754683.1| Hypothetical protein yfbG [Escherichia coli CFT... 216 9e-55
ref|YP_235768.1| Formyl transferase, N-terminal:Formyl transfera... 216 1e-54
>gb|AAM65998.1| putative dTDP-glucose 4-6-dehydratase [Arabidopsis thaliana]
Length = 389
Score = 603 bits (1554), Expect = e-171
Identities = 283/381 (74%), Positives = 339/381 (88%), Gaps = 3/381 (0%)
Query: 2 DRVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-P 60
DR++LDGKPI P++IC+IG GGFIGSHL EKLM+ET HK + +DV ++K+ HLL+
Sbjct: 6 DRLDLDGKPIKPMTICMIGAGGFIGSHLCEKLMTETPHKVLALDVYNDKIKHLLEPDTVQ 65
Query: 61 WANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK 120
WA RI+FH++NIK+DSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K
Sbjct: 66 WAGRIQFHRINIKHDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVK 125
Query: 121 FCTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSY 178
+C+ENNKRLIHFSTCEV+GKTIGSFLP+++ R++P++Y LKED+SPCIFG + KQRWSY
Sbjct: 126 YCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRQDPEFYVLKEDISPCIFGSIEKQRWSY 185
Query: 179 ACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLL 238
ACAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLL
Sbjct: 186 ACAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLL 245
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELM 298
R EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP+RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 246 RREPLKLVDGGESQRTFIYIKDAIEAVLLMIENPERANGHIFNVGNPNNEVTVRQLAEMM 305
Query: 299 IKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VYAKV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QLGW PKTSL DLL+
Sbjct: 306 TEVYAKVSGETAIDSPTIDVSSKEFYGEGYDDSDKRIPDMTIINRQLGWNPKTSLWDLLE 365
Query: 359 STLQYQHQTYSHAIKKELSKP 379
STL YQH TY+ AIKK SKP
Sbjct: 366 STLTYQHTTYAEAIKKATSKP 386
>ref|NP_563807.1| expressed protein [Arabidopsis thaliana] gi|25407031|pir||C86216
protein T23G18.6 [imported] - Arabidopsis thaliana
gi|6579211|gb|AAF18254.1| T23G18.6 [Arabidopsis
thaliana] gi|24899785|gb|AAN65107.1| similar to
dihydroflavonol reductase [Arabidopsis thaliana]
Length = 389
Score = 603 bits (1554), Expect = e-171
Identities = 283/381 (74%), Positives = 339/381 (88%), Gaps = 3/381 (0%)
Query: 2 DRVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-P 60
DR++LDGKPI P++IC+IG GGFIGSHL EKLM+ET HK + +DV ++K+ HLL+
Sbjct: 6 DRLDLDGKPIKPMTICMIGAGGFIGSHLCEKLMTETPHKVLALDVYNDKIKHLLEPDTVQ 65
Query: 61 WANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK 120
WA RI+FH++NIK+DSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K
Sbjct: 66 WAGRIQFHRINIKHDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVK 125
Query: 121 FCTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSY 178
+C+ENNKRLIHFSTCEV+GKTIGSFLP+++ R++P++Y LKED+SPCIFG + KQRWSY
Sbjct: 126 YCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRQDPEFYVLKEDISPCIFGSIEKQRWSY 185
Query: 179 ACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLL 238
ACAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLL
Sbjct: 186 ACAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLL 245
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELM 298
R EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP+RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 246 RREPLKLVDGGESQRTFIYIKDAIEAVLLMIENPERANGHIFNVGNPNNEVTVRQLAEMM 305
Query: 299 IKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VYAKV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QLGW PKTSL DLL+
Sbjct: 306 TEVYAKVSGETAIESPTIDVSSKEFYGEGYDDSDKRIPDMTIINRQLGWNPKTSLWDLLE 365
Query: 359 STLQYQHQTYSHAIKKELSKP 379
STL YQH TY+ AIKK SKP
Sbjct: 366 STLTYQHTTYAEAIKKATSKP 386
>gb|AAK68820.1| similar to dihydroflavonol reductase [Arabidopsis thaliana]
Length = 389
Score = 597 bits (1540), Expect = e-169
Identities = 282/381 (74%), Positives = 338/381 (88%), Gaps = 3/381 (0%)
Query: 2 DRVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-P 60
DR++LDGKPI P++IC+IG GGFIGSHL EKLM+ET HK + +DV ++K+ HLL+
Sbjct: 6 DRLDLDGKPIKPMTICMIGAGGFIGSHLCEKLMTETPHKVLALDVYNDKIKHLLEPDTVQ 65
Query: 61 WANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK 120
WA RI+FH++NIK+DSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K
Sbjct: 66 WAGRIQFHRINIKHDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVK 125
Query: 121 FCTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSY 178
+C+ENNKRLIHFSTCEV+GKTIGSFLP+++ R++P++Y LKED+SPCIFG + KQRWSY
Sbjct: 126 YCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRQDPEFYVLKEDISPCIFGSIEKQRWSY 185
Query: 179 ACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLL 238
ACAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLL
Sbjct: 186 ACAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLL 245
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELM 298
R EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP+RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 246 RREPLKLVDGGESQRTFIYIKDAIEAVLLMIENPERANGHIFNVGNPNNEVTVRQLAEMM 305
Query: 299 IKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VYAKV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QLG PKTSL DLL+
Sbjct: 306 TEVYAKVSGETAIESPTIDVSSKEFYGEGYDDSDKRIPDMTIINRQLGCTPKTSLWDLLE 365
Query: 359 STLQYQHQTYSHAIKKELSKP 379
STL YQH TY+ AIKK SKP
Sbjct: 366 STLTYQHTTYAEAIKKATSKP 386
>gb|AAR14687.1| UDP-D-apiose/UDP-D-xylose synthase [Arabidopsis thaliana]
gi|24111293|gb|AAN46770.1| At2g27860/F15K20.4
[Arabidopsis thaliana] gi|21555529|gb|AAM63878.1|
putative dTDP-glucose 4-6-dehydratase [Arabidopsis
thaliana] gi|52354277|gb|AAU44459.1| hypothetical
protein AT2G27860 [Arabidopsis thaliana]
gi|3860247|gb|AAC73015.1| putative dTDP-glucose
4-6-dehydratase [Arabidopsis thaliana]
gi|60547725|gb|AAX23826.1| hypothetical protein
At2g27860 [Arabidopsis thaliana]
gi|13605497|gb|AAK32742.1| At2g27860/F15K20.4
[Arabidopsis thaliana] gi|25407929|pir||G84677 probable
dTDP-glucose 4-6-dehydratase [imported] - Arabidopsis
thaliana gi|15226264|ref|NP_180353.1| expressed protein
[Arabidopsis thaliana]
Length = 389
Score = 593 bits (1530), Expect = e-168
Identities = 279/381 (73%), Positives = 336/381 (87%), Gaps = 3/381 (0%)
Query: 2 DRVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-P 60
+RV+LDGKPI P++IC+IG GGFIGSHL EKL++ET HK + +DV ++K+ HLL+
Sbjct: 6 NRVDLDGKPIQPLTICMIGAGGFIGSHLCEKLLTETPHKVLALDVYNDKIKHLLEPDTVE 65
Query: 61 WANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIK 120
W+ RI+FH++NIK+DSRLE LVK +DL INLAAICTPADYNTRPLDTIYSNFIDA+PV+K
Sbjct: 66 WSGRIQFHRINIKHDSRLEGLVKMADLIINLAAICTPADYNTRPLDTIYSNFIDALPVVK 125
Query: 121 FCTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSY 178
+C+ENNKRLIHFSTCEV+GKTIGSFLP+++ R +P +Y LKED+SPCIFG + KQRWSY
Sbjct: 126 YCSENNKRLIHFSTCEVYGKTIGSFLPKDHPLRDDPAFYVLKEDISPCIFGSIEKQRWSY 185
Query: 179 ACAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLL 238
ACAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLL
Sbjct: 186 ACAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLL 245
Query: 239 RGEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELM 298
R EPLKLVDGG SQRTF+YI DAIEAV+LMI+NP+RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 246 RREPLKLVDGGESQRTFVYINDAIEAVLLMIENPERANGHIFNVGNPNNEVTVRQLAEMM 305
Query: 299 IKVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VYAKV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QLGW PKTSL DLL+
Sbjct: 306 TEVYAKVSGEGAIESPTVDVSSKEFYGEGYDDSDKRIPDMTIINRQLGWNPKTSLWDLLE 365
Query: 359 STLQYQHQTYSHAIKKELSKP 379
STL YQH+TY+ A+KK SKP
Sbjct: 366 STLTYQHRTYAEAVKKATSKP 386
>gb|AAQ91380.1| putative nucleoside-diphosphate-sugar epimerase/dehydratase
[Nicotiana benthamiana]
Length = 387
Score = 587 bits (1512), Expect = e-166
Identities = 276/379 (72%), Positives = 329/379 (85%), Gaps = 3/379 (0%)
Query: 3 RVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSH-PW 61
RV+LDG I P+ IC+IG GGFIGSHL EKLM+ET H + +DV S+K+ HLL+ + PW
Sbjct: 5 RVDLDGNVIKPMKICMIGAGGFIGSHLCEKLMNETPHTVLAVDVYSDKIKHLLEPADLPW 64
Query: 62 ANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKF 121
RI+FH++NIKNDSRLE L+K +DL +NLAAICTPADYNTRPLDTIYSNFIDA+PV+K+
Sbjct: 65 TGRIQFHRINIKNDSRLEGLIKMADLVVNLAAICTPADYNTRPLDTIYSNFIDALPVVKY 124
Query: 122 CTENNKRLIHFSTCEVFGKTIGSFLPEE--YRKEPQYYKLKEDVSPCIFGPVHKQRWSYA 179
C+EN KRLIHFSTCEV+GKTIG+FLPE R++P YY LKEDVSPCIFG + KQRWSYA
Sbjct: 125 CSENGKRLIHFSTCEVYGKTIGAFLPEXSPLRQDPAYYVLKEDVSPCIFGSIEKQRWSYA 184
Query: 180 CAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLR 239
CAKQ+ +RLIYAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLLR
Sbjct: 185 CAKQLIERLIYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLLR 244
Query: 240 GEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
EPLKLVDGGHSQRTF+YIKDAIEAV+LMI+NP RANG IFNVGNP+NEV+V+QLAE+M
Sbjct: 245 HEPLKLVDGGHSQRTFIYIKDAIEAVLLMIENPARANGQIFNVGNPNNEVTVRQLAEMMT 304
Query: 300 KVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDS 359
+VY+KV+G T+DVSS+ FYG+GYDDSD+RIPDMT+I +QLGW PKTSL DLL+S
Sbjct: 305 QVYSKVSGESPPETPTIDVSSKEFYGEGYDDSDKRIPDMTLINRQLGWNPKTSLWDLLES 364
Query: 360 TLQYQHQTYSHAIKKELSK 378
L YQH+TY+ A+K+ +SK
Sbjct: 365 XLTYQHRTYAEAVKQAMSK 383
>ref|NP_914324.1| OJ1656_A11.18 [Oryza sativa (japonica cultivar-group)]
gi|18844860|dbj|BAB85329.1| putative dTDP-glucose
4,6-dehydratase [Oryza sativa (japonica cultivar-group)]
Length = 398
Score = 586 bits (1510), Expect = e-166
Identities = 279/381 (73%), Positives = 331/381 (86%), Gaps = 3/381 (0%)
Query: 3 RVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWA 62
R++LDG PI P++IC+IG GGFIGSHL EKLM+ET+H +DV +K+ HL+D + P
Sbjct: 15 RLDLDGNPIAPLTICMIGAGGFIGSHLCEKLMAETAHVVYAVDVYCDKIRHLVDPAPPHL 74
Query: 63 N-RIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKF 121
+ RI FH++NIKNDSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K+
Sbjct: 75 HGRISFHRLNIKNDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVKY 134
Query: 122 CTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSYA 179
C+ENNKRLIHFSTCEV+GKTIGSFLP ++ RKEP++Y LKED SPCIFGP+ KQRWSYA
Sbjct: 135 CSENNKRLIHFSTCEVYGKTIGSFLPTDHPLRKEPEFYVLKEDESPCIFGPIVKQRWSYA 194
Query: 180 CAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLR 239
CAKQ+ +RLI+AE AENGL+FTIVRP+NWIGPRMDFIPGVDGPS+GVPRVLACFSNNLLR
Sbjct: 195 CAKQLIERLIFAEGAENGLEFTIVRPFNWIGPRMDFIPGVDGPSEGVPRVLACFSNNLLR 254
Query: 240 GEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
EPLKLVDGG SQRTF+YIKDAIEAV LMI+NP RANG IFNVGNP+NEV+V+QLAE+M
Sbjct: 255 REPLKLVDGGQSQRTFVYIKDAIEAVHLMIENPARANGQIFNVGNPNNEVTVRQLAEMMT 314
Query: 300 KVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDS 359
+VYA V+G P +DVSS+ FYG+GYDDSD+RIPDMTII KQLGW PKT L DLL++
Sbjct: 315 EVYANVSGEPPLDEPMIDVSSKQFYGEGYDDSDKRIPDMTIINKQLGWNPKTPLKDLLET 374
Query: 360 TLQYQHQTYSHAIKKELSKPS 380
TL YQH+TY AIK+++S+ S
Sbjct: 375 TLTYQHKTYKEAIKRQMSQAS 395
>gb|AAS21758.1| dTDP-glucose 4,6-dehydratase [Zea mays]
Length = 395
Score = 572 bits (1475), Expect = e-162
Identities = 272/382 (71%), Positives = 328/382 (85%), Gaps = 4/382 (1%)
Query: 3 RVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPW- 61
R++LDG PI P++IC+IG GGFIGSHL EKLM+ET H + +DV +K+ HL+D + P
Sbjct: 11 RLDLDGNPIAPLTICMIGAGGFIGSHLCEKLMAETHHTVLAVDVYCDKIRHLVDPAPPHL 70
Query: 62 ANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKF 121
A RI FH++NIKNDSRLE L+K +DLTINLAAICTPADYNTRPLDTIYSNFIDA+PV+K+
Sbjct: 71 AGRISFHRLNIKNDSRLEGLIKMADLTINLAAICTPADYNTRPLDTIYSNFIDALPVVKY 130
Query: 122 CTENNKRLIHFSTCEVFGKTIGSFLPEEY--RKEPQYYKLKEDVSPCIFGPVHKQRWSYA 179
C+EN+KRLIHF TCEV+GKTIGSFLP+++ RKEP++Y LKED SPCIFGP+ KQRWSYA
Sbjct: 131 CSENSKRLIHFPTCEVYGKTIGSFLPKDHPLRKEPEFYVLKEDESPCIFGPIVKQRWSYA 190
Query: 180 CAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLR 239
CAKQ+ +RL++AE AENGL FTIVRP+NWIGPRMDFIPGVDGPS+GVPRVLACFSNNLLR
Sbjct: 191 CAKQLIERLVFAEGAENGLDFTIVRPFNWIGPRMDFIPGVDGPSEGVPRVLACFSNNLLR 250
Query: 240 GEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP RANGHIFNVGNP+NEV+V++LA +M
Sbjct: 251 REPLKLVDGGQSQRTFVYIKDAIEAVVLMIENPARANGHIFNVGNPNNEVTVRELAPMMT 310
Query: 300 KVYAKVA-GVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLD 358
+VY +++ G +DVSS FYG+GYDDSD+RIPDMTII KQLGW PKT L DLL+
Sbjct: 311 EVYTQMSQGEAPLDEPMIDVSSSQFYGEGYDDSDKRIPDMTIINKQLGWNPKTPLKDLLE 370
Query: 359 STLQYQHQTYSHAIKKELSKPS 380
+TL YQH+TY A K+++S+ S
Sbjct: 371 TTLTYQHKTYKEAAKRQMSQAS 392
>gb|AAS79591.1| putative dihydroflavonol reductase [Ipomoea trifida]
Length = 407
Score = 553 bits (1425), Expect = e-156
Identities = 260/346 (75%), Positives = 308/346 (88%), Gaps = 3/346 (0%)
Query: 3 RVNLDGKPIVPISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDK-SHPW 61
RV+LDG PI PI+IC+IG GGFIGSHL EKLMSET HK + +DV ++K+ HLL+ S PW
Sbjct: 4 RVDLDGNPIKPITICMIGAGGFIGSHLCEKLMSETQHKVLAVDVYNDKIKHLLEPASLPW 63
Query: 62 ANRIEFHQMNIKNDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKF 121
A+RI+FH++NIKNDSRLE L+K +DLT+NLAAICTPADYNTRPLDTIYSNFIDA+PV+K+
Sbjct: 64 ADRIQFHRLNIKNDSRLEGLIKMADLTVNLAAICTPADYNTRPLDTIYSNFIDALPVVKY 123
Query: 122 CTENNKRLIHFSTCEVFGKTIGSFLPEE--YRKEPQYYKLKEDVSPCIFGPVHKQRWSYA 179
C+EN KRLIHFSTCEV+GKTIG FLP++ R++P YY LKED SPCIFGP+ KQRWSYA
Sbjct: 124 CSENGKRLIHFSTCEVYGKTIGCFLPKDSPLRQDPAYYVLKEDASPCIFGPIEKQRWSYA 183
Query: 180 CAKQMTDRLIYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLR 239
CAKQ+ +RL+YAE AENGL+FTIVRP+NWIGPRMDFIPG+DGPS+GVPRVLACFSNNLLR
Sbjct: 184 CAKQLIERLVYAEGAENGLEFTIVRPFNWIGPRMDFIPGIDGPSEGVPRVLACFSNNLLR 243
Query: 240 GEPLKLVDGGHSQRTFLYIKDAIEAVMLMIDNPDRANGHIFNVGNPDNEVSVKQLAELMI 299
EPLKLVDGG SQRTF+YIKDAIEAV+LMI+NP RANGHIFNVGNP+NEV+V+QLAE+M
Sbjct: 244 REPLKLVDGGQSQRTFVYIKDAIEAVVLMIENPARANGHIFNVGNPNNEVTVRQLAEMMT 303
Query: 300 KVYAKVAGVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQL 345
+VY+KV+G T+DVSS+ FYG+GYDDSD+RIPDMTII +QL
Sbjct: 304 QVYSKVSGEVSLETPTIDVSSKEFYGEGYDDSDKRIPDMTIINRQL 349
>ref|ZP_00170676.1| COG0451: Nucleoside-diphosphate-sugar epimerases [Ralstonia
eutropha JMP134]
Length = 349
Score = 233 bits (594), Expect = 7e-60
Identities = 132/347 (38%), Positives = 202/347 (58%), Gaps = 27/347 (7%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK-N 74
+ ++G GFIG HLT +++ TS + +D+SS+++ L+D HP R+ F + +I N
Sbjct: 6 VLILGVNGFIGHHLTRRILETTSWEVYGMDMSSDRLGDLVD--HP---RMHFFEGDITIN 60
Query: 75 DSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
+E ++ D+ + L AI TPA Y +PL +F +P+++ + K L+ ST
Sbjct: 61 KEWIEYNIRKCDVVLPLVAIATPATYVRQPLRVFELDFEANLPIVRAAVKYGKHLVFPST 120
Query: 135 CEVFGKTIGS-FLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEH 193
EV+G F PE SP I+GP++K RW YAC+KQ+ DR+I+A
Sbjct: 121 SEVYGMCADDEFDPES--------------SPLIYGPINKPRWIYACSKQLMDRVIHAYG 166
Query: 194 AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQR 253
E GL +T+ RP+NWIG +D + +G RV+ F +++RGEP+KLVDGG +R
Sbjct: 167 MEQGLNYTLFRPFNWIGAGLD---SIFESKEGSSRVVTQFLGHIVRGEPIKLVDGGEQKR 223
Query: 254 TFLYIKDAIEAVMLMIDNPDR-ANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESS 312
F I D I A+M +I+NP+ A G IFN+GNP N SV++LAE+M+K+ A E +
Sbjct: 224 AFADISDGISALMRIIENPNGIATGKIFNIGNPSNIHSVRELAEMMLKMAADYPEYAEEA 283
Query: 313 LST--LDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLL 357
T ++ SS FYGKGY D R+P + ++LGWKP+ +++ L
Sbjct: 284 RKTQIVETSSGDFYGKGYQDVQHRVPKIDNTMQELGWKPEVTMEQAL 330
>ref|ZP_00202404.1| COG0451: Nucleoside-diphosphate-sugar epimerases [Ralstonia
eutropha JMP134]
Length = 350
Score = 233 bits (594), Expect = 7e-60
Identities = 132/347 (38%), Positives = 202/347 (58%), Gaps = 27/347 (7%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK-N 74
+ ++G GFIG HLT +++ TS + +D+SS+++ L+D HP R+ F + +I N
Sbjct: 4 VLILGVNGFIGHHLTRRILETTSWEVYGMDMSSDRLGDLVD--HP---RMHFFEGDITIN 58
Query: 75 DSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
+E ++ D+ + L AI TPA Y +PL +F +P+++ + K L+ ST
Sbjct: 59 KEWIEYNIRKCDVVLPLVAIATPATYVRQPLRVFELDFEANLPIVRAAVKYGKHLVFPST 118
Query: 135 CEVFGKTIGS-FLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEH 193
EV+G F PE SP I+GP++K RW YAC+KQ+ DR+I+A
Sbjct: 119 SEVYGMCADDEFDPES--------------SPLIYGPINKPRWIYACSKQLMDRVIHAYG 164
Query: 194 AENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQR 253
E GL +T+ RP+NWIG +D + +G RV+ F +++RGEP+KLVDGG +R
Sbjct: 165 MEQGLNYTLFRPFNWIGAGLD---SIFESKEGSSRVVTQFLGHIVRGEPIKLVDGGEQKR 221
Query: 254 TFLYIKDAIEAVMLMIDNPDR-ANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESS 312
F I D I A+M +I+NP+ A G IFN+GNP N SV++LAE+M+K+ A E +
Sbjct: 222 AFADISDGISALMRIIENPNGIATGKIFNIGNPSNIHSVRELAEMMLKMAADYPEYAEEA 281
Query: 313 LST--LDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLL 357
T ++ SS FYGKGY D R+P + ++LGWKP+ +++ L
Sbjct: 282 RKTQIVETSSGDFYGKGYQDVQHRVPKIDNTMQELGWKPEVTMEQAL 328
>ref|ZP_00273539.1| COG0451: Nucleoside-diphosphate-sugar epimerases [Ralstonia
metallidurans CH34]
Length = 350
Score = 231 bits (590), Expect = 2e-59
Identities = 132/346 (38%), Positives = 201/346 (57%), Gaps = 25/346 (7%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK-N 74
+ ++G GFIG HLT +++ T + +D+SS+++ L++ HP R+ F + +I N
Sbjct: 4 VLILGVNGFIGHHLTRRILETTQWEVYGMDMSSDRLGDLVN--HP---RMHFFEGDITIN 58
Query: 75 DSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
+E ++ D+ + L AI TPA Y PL +F +P+++ + K L+ ST
Sbjct: 59 KEWIEYNIRKCDVVLPLVAIATPATYVREPLRVFELDFEANLPIVRAAVKYGKHLVFPST 118
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G EE+ E SP I+GP++K RW YAC+KQ+ DR+I+A
Sbjct: 119 SEVYGMCAD----EEFDPE---------ASPLIYGPINKPRWIYACSKQLMDRVIHAYGM 165
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL +T+ RP+NWIG +D + +G RV+ F +++RGEP+KLVDGG QR
Sbjct: 166 QEGLNYTLFRPFNWIGAGLD---SIFESKEGSSRVVTQFLGHIVRGEPIKLVDGGAQQRA 222
Query: 255 FLYIKDAIEAVMLMIDNPDR-ANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESSL 313
F I D I A+M +I+N D ANG IFN+GNP N SV++LAE+M+K+ A+ E +
Sbjct: 223 FADISDGISALMRIIENKDGVANGKIFNIGNPGNIHSVRELAEMMLKMAAEYPEYAEEAR 282
Query: 314 ST--LDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLL 357
T ++ SS FYGKGY D R+P + +LGWKP+ S++ L
Sbjct: 283 KTKIVETSSGDFYGKGYQDVQHRVPKIDNTIGELGWKPEVSMEQAL 328
>ref|ZP_00276079.1| COG0451: Nucleoside-diphosphate-sugar epimerases [Ralstonia
metallidurans CH34]
Length = 352
Score = 231 bits (589), Expect = 3e-59
Identities = 132/346 (38%), Positives = 201/346 (57%), Gaps = 25/346 (7%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK-N 74
+ ++G GFIG HLT +++ T + +D+SS+++ L++ HP R+ F + +I N
Sbjct: 6 VLILGINGFIGHHLTRRILETTQWEVYGMDMSSDRLGDLVN--HP---RMHFFEGDITIN 60
Query: 75 DSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
+E ++ D+ + L AI TPA Y PL +F +P+++ + K L+ ST
Sbjct: 61 KEWIEYNIRKCDVVLPLVAIATPATYVREPLRVFELDFEANLPIVRAAVKYGKHLVFPST 120
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G EE+ E SP I+GP++K RW YAC+KQ+ DR+I+A
Sbjct: 121 SEVYGMCAD----EEFDPE---------ASPLIYGPINKPRWIYACSKQLMDRVIHAYGM 167
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL +T+ RP+NWIG +D + +G RV+ F +++RGEP+KLVDGG QR
Sbjct: 168 QEGLNYTLFRPFNWIGAGLD---SIFESKEGSSRVVTQFLGHIVRGEPIKLVDGGAQQRA 224
Query: 255 FLYIKDAIEAVMLMIDNPDR-ANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAGVPESSL 313
F I D I A+M +I+N D ANG IFN+GNP N SV++LAE+M+K+ A+ E +
Sbjct: 225 FADISDGISALMRIIENKDGVANGKIFNIGNPGNIHSVRELAEMMLKMAAEYPEYAEEAR 284
Query: 314 ST--LDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLL 357
T ++ SS FYGKGY D R+P + +LGWKP+ S++ L
Sbjct: 285 KTKIVETSSGDFYGKGYQDVQHRVPKIDNTIGELGWKPEVSMEQAL 330
>gb|AAQ58423.1| probable transformylase [Chromobacterium violaceum ATCC 12472]
gi|34496202|ref|NP_900417.1| probable transformylase
[Chromobacterium violaceum ATCC 12472]
Length = 347
Score = 224 bits (570), Expect = 4e-57
Identities = 123/353 (34%), Positives = 202/353 (56%), Gaps = 27/353 (7%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK-N 74
+ ++G GFIG HLT++++ T + +D+ +++V K HP R F + +I N
Sbjct: 4 VLILGVNGFIGHHLTKRIIETTDWEIYGMDMHADRVAEW--KDHP---RFHFFEGDITIN 58
Query: 75 DSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
+E VK D+ + L AI TP+ Y PL +F +P+++ C + K L+ ST
Sbjct: 59 KEWIEYHVKKCDVVLPLVAIATPSTYVNNPLRVFELDFEANLPIVRQCVKYKKHLVFPST 118
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G + ++ +P+ +L I+GP++K RW YAC+KQ+ DR+I+A
Sbjct: 119 SEVYG------MSQDAEFDPENSQL-------IYGPINKPRWIYACSKQLMDRVIHAYAM 165
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
E GL +T+ RP+NWIG +D ++ P +G RV+ F +++RGE +KLVDGGH +R
Sbjct: 166 EEGLNYTLFRPFNWIGGGLD---NINTPKEGSSRVITQFLGHIVRGETIKLVDGGHQKRA 222
Query: 255 FLYIKDAIEAVMLMIDNPD-RANGHIFNVGNPDNEVSVKQLAELMI---KVYAKVAGVPE 310
F Y+ D I A+M +I+N D +A+G I+N+GNP N S+++LA++M+ +VY + +
Sbjct: 223 FTYVDDGISALMKIIENKDGKASGQIYNIGNPANNYSIRELAQMMLDLARVYPEYQ-LNA 281
Query: 311 SSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+ ++ +S +YGKGY D R+P + L WKP ++ D L Y
Sbjct: 282 DKVQVVETTSGQYYGKGYQDVQNRVPKIANTMADLDWKPGVTMADALRGIYDY 334
>emb|CAD15021.1| PUTATIVE OXIDOREDUCTASE PROTEIN [Ralstonia solanacearum]
gi|17546038|ref|NP_519440.1| PUTATIVE OXIDOREDUCTASE
PROTEIN [Ralstonia solanacearum GMI1000]
Length = 351
Score = 218 bits (555), Expect = 2e-55
Identities = 124/352 (35%), Positives = 203/352 (57%), Gaps = 33/352 (9%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHK-----AIVIDVSSEKVNHLLDKSHPWANRIEFHQM 70
+ ++G GFIG HL+++++ T + +D+ +E++ L++ HP R+ F +
Sbjct: 4 VLILGVNGFIGHHLSKRILESTDPEISQWEVYGMDMQTERLGDLVN--HP---RMHFFEG 58
Query: 71 NIK-NDSRLETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRL 129
+I N +E V+ D+ + L AI TP+ Y PL +F +P+++ + K L
Sbjct: 59 DITINKEWVEYHVRKCDVILPLVAIATPSTYVKAPLRVFELDFEANLPIVRSAAKYGKHL 118
Query: 130 IHFSTCEVFGKT-IGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRL 188
+ ST EV+G F PE SP ++GP++K RW YAC+KQ+ DR+
Sbjct: 119 VFPSTSEVYGMCGDDEFDPE--------------ASPLVYGPINKPRWIYACSKQLMDRV 164
Query: 189 IYAEHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDG 248
I+ E GL FT+ RP+NWIGP +D + P +G RV+ F +++RGE ++LVDG
Sbjct: 165 IWGYGME-GLNFTLFRPFNWIGPGLD---SIHTPKEGSSRVVTQFLGHIVRGENIQLVDG 220
Query: 249 GHSQRTFLYIKDAIEAVMLMIDNPDR-ANGHIFNVGNPDNEVSVKQLAELMIKVYAKVAG 307
G +R F Y+ D I+A++ +I N D A+G I+N+GNP N SV++LAE+M+K +A
Sbjct: 221 GQQKRAFTYVDDGIDALVRIIANKDGVASGKIYNIGNPSNNYSVRELAEMMLKKAGTIAE 280
Query: 308 VPESS--LSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLL 357
E++ + ++ +S +YGKGY D R+P + ++LGWKP T+++D L
Sbjct: 281 YKENAQKVKLVETTSGAYYGKGYQDVQNRVPKIANTMEELGWKPTTTMEDTL 332
>gb|AAL21200.1| putative transformylase [Salmonella typhimurium LT2]
gi|6136698|sp|O52325|ARNA_SALTY Bifunctional polymyxin
resistance arnA protein (Polymyxin resistance protein
pmrI) [Includes: UDP-glucuronic acid decarboxylase
(UDP-GlcUA decarboxylase) (ArnAFT);
UDP-4-amino-4-deoxy-L-arabinose formyltransferase
(UDP-L-Ara4N formyltransferase) (ArnADH)]
gi|2921421|gb|AAC04772.1| unknown [Salmonella
typhimurium] gi|16765626|ref|NP_461241.1| putative
transformylase [Salmonella typhimurium LT2]
Length = 660
Score = 218 bits (554), Expect = 3e-55
Identities = 127/357 (35%), Positives = 201/357 (55%), Gaps = 31/357 (8%)
Query: 14 ISICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIK 73
I + ++G GFIG+HLTE+L++E +++ +D+ S ++ L HP R F + +I
Sbjct: 316 IRVLILGVNGFIGNHLTERLLNEENYEVYGMDIGSNAISRFL--LHP---RFHFVEGDIS 370
Query: 74 NDSR-LETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHF 132
S +E VK D+ + L AI TP +Y PL +F + + +I++C + KR++
Sbjct: 371 IHSEWIEYHVKKCDVVLPLVAIATPIEYTRNPLRVFELDFEENLRIIRYCVKYRKRVVFP 430
Query: 133 STCEVFGK-TIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYA 191
ST EV+G T SF ED S I GPV+K RW Y+ +KQ+ DR+I+A
Sbjct: 431 STSEVYGMCTDASF--------------DEDKSNLIVGPVNKPRWIYSVSKQLLDRVIWA 476
Query: 192 EHAENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHS 251
+ GL+FT+ RP+NW+GPR+D ++ G R + NL+ G P+KL+DGG
Sbjct: 477 YGEKEGLRFTLFRPFNWMGPRLD---SLNAARIGSSRAITQLILNLVEGTPIKLIDGGQQ 533
Query: 252 QRTFLYIKDAIEAVM-LMIDNPDRANGHIFNVGNPDNEVSVKQLAELMIKVYAK----VA 306
+R F I+D IEA+ +++++ DR +G I N+GNPDNE S+++LA L++ + K
Sbjct: 534 KRCFTDIRDGIEALFRIIVNDGDRCDGKIINIGNPDNEASIQELATLLLDSFDKHPLRCH 593
Query: 307 GVPESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
P + V S +YGKGY D R P + + LGW+P ++ D ++ TL +
Sbjct: 594 FPPFAGFQV--VESRSYYGKGYQDVAHRKPSIDNARRCLGWEPSIAMRDTVEETLDF 648
>gb|AAL23678.1| UDP-D-glucuronate dehydrogenase [Escherichia coli]
gi|1788589|gb|AAC75315.1| putative transformylase;
putative formyltransferase [Escherichia coli K12]
gi|16130190|ref|NP_416758.1| putative formyltransferase
[Escherichia coli K12] gi|6176575|sp|P77398|YFBG_ECOLI
Hypothetical protein yfbG gi|1799612|dbj|BAA16082.1|
METHIONYL-TRNA FORMYLTRANSFERASE (EC 2.1.2.9).
[Escherichia coli] gi|1799607|dbj|BAA16078.1|
METHIONYL-TRNA FORMYLTRANSFERASE (EC 2.1.2.9).
[Escherichia coli]
Length = 660
Score = 216 bits (551), Expect = 7e-55
Identities = 120/354 (33%), Positives = 198/354 (55%), Gaps = 29/354 (8%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIKND 75
+ ++G GFIG+HLTE+L+ E ++ +D+ S+ ++ L+ H F + +I
Sbjct: 318 VLILGVNGFIGNHLTERLLREDHYEVYGLDIGSDAISRFLNHPH-----FHFVEGDISIH 372
Query: 76 SR-LETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
S +E VK D+ + L AI TP +Y PL +F + + +I++C + KR+I ST
Sbjct: 373 SEWIEYHVKKCDVVLPLVAIATPIEYTRNPLRVFELDFEENLRIIRYCVKYRKRIIFPST 432
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G + E++ S I GPV+K RW Y+ +KQ+ DR+I+A
Sbjct: 433 SEVYGMCSDKYFDEDH-------------SNLIVGPVNKPRWIYSVSKQLLDRVIWAYGE 479
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL+FT+ RP+NW+GPR+D ++ G R + NL+ G P+KL+DGG +R
Sbjct: 480 KEGLQFTLFRPFNWMGPRLD---NLNAARIGSSRAITQLILNLVEGSPIKLIDGGKQKRC 536
Query: 255 FLYIKDAIEAVMLMIDNP-DRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVA----GVP 309
F I+D IEA+ +I+N +R +G I N+GNP+NE S+++L E+++ + K P
Sbjct: 537 FTDIRDGIEALYRIIENAGNRCDGEIINIGNPENEASIEELGEMLLASFEKHPLRHHFPP 596
Query: 310 ESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+ ++ SS +YGKGY D + R P + + L W+PK + + +D TL +
Sbjct: 597 FAGFRVVESSS--YYGKGYQDVEHRKPSIRNAHRCLDWEPKIDMQETIDETLDF 648
>pdb|2BLL|A Chain A, Apo-Structure Of The C-Terminal Decarboxylase Domain Of
Arna
Length = 345
Score = 216 bits (551), Expect = 7e-55
Identities = 120/354 (33%), Positives = 198/354 (55%), Gaps = 29/354 (8%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIKND 75
+ ++G GFIG+HLTE+L+ E ++ +D+ S+ ++ L+ H F + +I
Sbjct: 3 VLILGVNGFIGNHLTERLLREDHYEVYGLDIGSDAISRFLNHPH-----FHFVEGDISIH 57
Query: 76 SR-LETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
S +E VK D+ + L AI TP +Y PL +F + + +I++C + KR+I ST
Sbjct: 58 SEWIEYHVKKCDVVLPLVAIATPIEYTRNPLRVFELDFEENLRIIRYCVKYRKRIIFPST 117
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G + E++ S I GPV+K RW Y+ +KQ+ DR+I+A
Sbjct: 118 SEVYGMCSDKYFDEDH-------------SNLIVGPVNKPRWIYSVSKQLLDRVIWAYGE 164
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL+FT+ RP+NW+GPR+D ++ G R + NL+ G P+KL+DGG +R
Sbjct: 165 KEGLQFTLFRPFNWMGPRLD---NLNAARIGSSRAITQLILNLVEGSPIKLIDGGKQKRC 221
Query: 255 FLYIKDAIEAVMLMIDNP-DRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVA----GVP 309
F I+D IEA+ +I+N +R +G I N+GNP+NE S+++L E+++ + K P
Sbjct: 222 FTDIRDGIEALYRIIENAGNRCDGEIINIGNPENEASIEELGEMLLASFEKHPLRHHFPP 281
Query: 310 ESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+ ++ SS +YGKGY D + R P + + L W+PK + + +D TL +
Sbjct: 282 FAGFRVVESSS--YYGKGYQDVEHRKPSIRNAHRCLDWEPKIDMQETIDETLDF 333
>pdb|1U9J|A Chain A, Crystal Structure Of E. Coli Arna (Pmri) Decarboxylase
Domain
Length = 358
Score = 216 bits (551), Expect = 7e-55
Identities = 120/354 (33%), Positives = 198/354 (55%), Gaps = 29/354 (8%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIKND 75
+ ++G GFIG+HLTE+L+ E ++ +D+ S+ ++ L+ H F + +I
Sbjct: 16 VLILGVNGFIGNHLTERLLREDHYEVYGLDIGSDAISRFLNHPH-----FHFVEGDISIH 70
Query: 76 SR-LETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
S +E VK D+ + L AI TP +Y PL +F + + +I++C + KR+I ST
Sbjct: 71 SEWIEYHVKKCDVVLPLVAIATPIEYTRNPLRVFELDFEENLRIIRYCVKYRKRIIFPST 130
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G + E++ S I GPV+K RW Y+ +KQ+ DR+I+A
Sbjct: 131 SEVYGMCSDKYFDEDH-------------SNLIVGPVNKPRWIYSVSKQLLDRVIWAYGE 177
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL+FT+ RP+NW+GPR+D ++ G R + NL+ G P+KL+DGG +R
Sbjct: 178 KEGLQFTLFRPFNWMGPRLD---NLNAARIGSSRAITQLILNLVEGSPIKLIDGGKQKRC 234
Query: 255 FLYIKDAIEAVMLMIDNP-DRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVA----GVP 309
F I+D IEA+ +I+N +R +G I N+GNP+NE S+++L E+++ + K P
Sbjct: 235 FTDIRDGIEALYRIIENAGNRCDGEIINIGNPENEASIEELGEMLLASFEKHPLRHHFPP 294
Query: 310 ESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+ ++ SS +YGKGY D + R P + + L W+PK + + +D TL +
Sbjct: 295 FAGFRVVESSS--YYGKGYQDVEHRKPSIRNAHRCLDWEPKIDMQETIDETLDF 346
>ref|NP_754683.1| Hypothetical protein yfbG [Escherichia coli CFT073]
gi|26109048|gb|AAN81251.1| Hypothetical protein yfbG
[Escherichia coli CFT073]
Length = 660
Score = 216 bits (550), Expect = 9e-55
Identities = 119/354 (33%), Positives = 198/354 (55%), Gaps = 29/354 (8%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIKND 75
+ ++G GFIG+HLTE+L+ E ++ +D+ S+ ++ ++ H F + +I
Sbjct: 318 VLILGVNGFIGNHLTERLLREDHYEVYGLDIGSDAISRFMNHPH-----FHFVEGDISIH 372
Query: 76 SR-LETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
S +E VK D+ + L AI TP +Y PL +F + + +I++C + KR+I ST
Sbjct: 373 SEWIEYHVKKCDVVLPLVAIATPIEYTRNPLRVFELDFEENLRIIRYCVKYRKRIIFPST 432
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G + E++ S I GPV+K RW Y+ +KQ+ DR+I+A
Sbjct: 433 SEVYGMCSDKYFDEDH-------------SNLIVGPVNKPRWIYSVSKQLLDRVIWAYGE 479
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL+FT+ RP+NW+GPR+D ++ G R + NL+ G P+KL+DGG +R
Sbjct: 480 KEGLQFTLFRPFNWMGPRLD---NLNAARIGSSRAITQLILNLVEGSPIKLIDGGKQKRC 536
Query: 255 FLYIKDAIEAVMLMIDNP-DRANGHIFNVGNPDNEVSVKQLAELMIKVYAKVA----GVP 309
F I+D IEA+ +I+N +R +G I N+GNP+NE S+++L E+++ + K P
Sbjct: 537 FTDIRDGIEALYRIIENAGNRCDGEIINIGNPENEASIEELGEMLLASFEKHPLRHYFPP 596
Query: 310 ESSLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+ ++ SS +YGKGY D + R P + + L W+PK + + +D TL +
Sbjct: 597 FAGFRVVESSS--YYGKGYQDVEHRKPSIRNARRCLDWEPKIDMQETIDETLDF 648
>ref|YP_235768.1| Formyl transferase, N-terminal:Formyl transferase, C-terminal
[Pseudomonas syringae pv. syringae B728a]
gi|63256634|gb|AAY37730.1| Formyl transferase,
N-terminal:Formyl transferase, C-terminal [Pseudomonas
syringae pv. syringae B728a]
Length = 664
Score = 216 bits (549), Expect = 1e-54
Identities = 121/352 (34%), Positives = 194/352 (54%), Gaps = 26/352 (7%)
Query: 16 ICLIGGGGFIGSHLTEKLMSETSHKAIVIDVSSEKVNHLLDKSHPWANRIEFHQMNIKND 75
+ ++G GFIG+HL+E+L+ + ++ +D+ S+ + L K + F + +I
Sbjct: 322 VLILGVNGFIGNHLSERLLQDDRYEIYGMDIGSDAIERLRAKPN-----FHFIEGDISIH 376
Query: 76 SR-LETLVKASDLTINLAAICTPADYNTRPLDTIYSNFIDAIPVIKFCTENNKRLIHFST 134
+ +E +K D+ + L AI TP +Y PL +F + + ++++C + NKR+I ST
Sbjct: 377 TEWIEYHIKKCDVVLPLVAIATPIEYTRNPLRVFELDFEENLKIVRYCVKYNKRVIFPST 436
Query: 135 CEVFGKTIGSFLPEEYRKEPQYYKLKEDVSPCIFGPVHKQRWSYACAKQMTDRLIYAEHA 194
EV+G Q ED S I GP++KQRW Y+ +KQ+ DR+I+A +
Sbjct: 437 SEVYGMC-------------QDANFNEDTSNLIVGPINKQRWIYSVSKQLLDRVIWA-YG 482
Query: 195 ENGLKFTIVRPYNWIGPRMDFIPGVDGPSDGVPRVLACFSNNLLRGEPLKLVDGGHSQRT 254
+ GL+FT+ RP+NW+GPR+D +D G R + +L+ G P++LVDGG +R
Sbjct: 483 QKGLQFTLFRPFNWMGPRLD---RLDSARIGSSRAITQLILHLVEGTPIRLVDGGAQKRC 539
Query: 255 FLYIKDAIEAVMLMIDNPD-RANGHIFNVGNPDNEVSVKQLAELMIKVYA--KVAGVPES 311
F + D IEA+ +I+N D R NG I N+GNPDNE S++QL E +++ + + G
Sbjct: 540 FTDVVDGIEALARIIENRDGRCNGQIINIGNPDNEASIRQLGEELLRQFEAHPLRGHFPP 599
Query: 312 SLSTLDVSSEVFYGKGYDDSDRRIPDMTIITKQLGWKPKTSLDDLLDSTLQY 363
+V S+ FYGKGY D R P + K +GW P L + + TL +
Sbjct: 600 FAGFREVESQSFYGKGYQDVSHRTPSIDNAKKLIGWTPGIELSETIGKTLDF 651
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 669,243,924
Number of Sequences: 2540612
Number of extensions: 28588170
Number of successful extensions: 65904
Number of sequences better than 10.0: 1825
Number of HSP's better than 10.0 without gapping: 280
Number of HSP's successfully gapped in prelim test: 1546
Number of HSP's that attempted gapping in prelim test: 63359
Number of HSP's gapped (non-prelim): 2057
length of query: 381
length of database: 863,360,394
effective HSP length: 130
effective length of query: 251
effective length of database: 533,080,834
effective search space: 133803289334
effective search space used: 133803289334
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC144515.13


