

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144513.4 - phase: 0 /pseudo
(1015 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
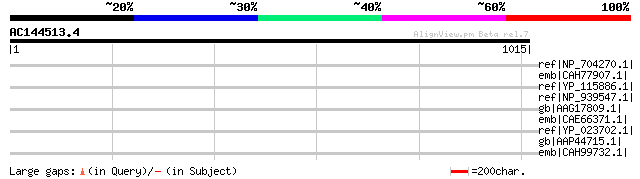
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_704270.1| integral membrane protein, conserved [Plasmodiu... 41 0.23
emb|CAH77907.1| integral membrane protein, conserved, putative [... 39 1.2
ref|YP_115886.1| putative membrane lipoprotein [Mycoplasma hyopn... 38 1.5
ref|NP_939547.1| Putative peptide transport protein [Corynebacte... 38 1.5
gb|AAG17809.1| NADH dehydrogenase subunit 4 [Naegleria gruberi] ... 38 2.0
emb|CAE66371.1| Hypothetical protein CBG11630 [Caenorhabditis br... 37 2.6
ref|YP_023702.1| signal peptidase I [Picrophilus torridus DSM 97... 37 3.4
gb|AAP44715.1| NADH dehydrogenase subunit 2 [lepidopsocid RS-200... 36 5.7
emb|CAH99732.1| integral membrane protein, conserved, putative [... 36 5.7
>ref|NP_704270.1| integral membrane protein, conserved [Plasmodium falciparum 3D7]
gi|23499009|emb|CAD51089.1| integral membrane protein,
conserved [Plasmodium falciparum 3D7]
Length = 505
Score = 40.8 bits (94), Expect = 0.23
Identities = 42/150 (28%), Positives = 63/150 (42%), Gaps = 27/150 (18%)
Query: 611 YSPEASDKGTLFLLIYSFYVLKCYLV*FRKVGIRVTFMVSKLLLMPPALLTFFMQMAIFS 670
+SP K +F + SF + Y++ F K+ IR + S +++ P F
Sbjct: 292 FSPNLLGKMAMFQSLASFISIISYMLFFTKIDIRKLLLYSTIIITP------------FC 339
Query: 671 FVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSLAQIPVLVL-----------KIYSE 719
+ L +++ + Y F NT LF +LI I +P+LVL IYS
Sbjct: 340 LLPLVVIKKVNYFLFIPNT-LFFITDTVLIEFIAEFQTMPILVLCSRLIPEGFESTIYSL 398
Query: 720 LISLSTSRIIFLSIWGFLLSLEDLKLKILS 749
L+S + I S FL SL L I S
Sbjct: 399 LLSSNNFASIISS---FLSSLLTYSLNITS 425
>emb|CAH77907.1| integral membrane protein, conserved, putative [Plasmodium
chabaudi]
Length = 448
Score = 38.5 bits (88), Expect = 1.2
Identities = 35/145 (24%), Positives = 61/145 (41%), Gaps = 29/145 (20%)
Query: 611 YSPEASDKGTLFLLIYSFYVLKCYLV*FRKVGIRVTFMVSKLLLMPPALLTFFMQMAIFS 670
++P+ K ++F + S + Y++ K+ I+ + S +++ P +LL I
Sbjct: 292 FTPKLLGKISMFQSLASLLAILLYMLILNKINIKKLLLYSTIVIAPFSLLPLLAVQKI-- 349
Query: 671 FVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSLAQIPVLVLKIYSELISLSTSRII- 729
F+ LF+ IL+ I L IP+LV SRII
Sbjct: 350 -------------NIFIPNTLFIITDTILVEFIGELQSIPILV----------ECSRIIP 386
Query: 730 ---FLSIWGFLLSLEDLKLKILSLL 751
+I+ FLLS+ + + I SL+
Sbjct: 387 EGFESTIYSFLLSVNNFAIMISSLI 411
>ref|YP_115886.1| putative membrane lipoprotein [Mycoplasma hyopneumoniae 232]
gi|53987339|gb|AAV27540.1| putative membrane lipoprotein
[Mycoplasma hyopneumoniae 232]
Length = 411
Score = 38.1 bits (87), Expect = 1.5
Identities = 42/150 (28%), Positives = 70/150 (46%), Gaps = 20/150 (13%)
Query: 613 PEASD--KGTLFLLIYSFYVLKCYLV*-FRKV--GIRVTFMVSKLLLMPPALLTFFMQMA 667
PEA+D K + L IY +V+ + + F+K G+ F + + P Q+
Sbjct: 152 PEANDFRKWLVNLGIYQIFVISGFHINLFKKAIFGVSKAFKIKPFIYKPIFFAFLLFQLF 211
Query: 668 IFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSLAQIPVLVLKIYSELISLSTSR 727
+F+F ++ +R ++ + KLFL +KL I L S ++ LST+
Sbjct: 212 LFNFA-VSFLRGFIFWSLIEINKLFLNKKLNKIE------------LLSISGIVILSTNP 258
Query: 728 IIFLSIWGFLLSLEDLKLKILSLLWIEFGK 757
+IF SI GF+L+ + IL + I F K
Sbjct: 259 LIFYSI-GFVLTF-TISFVILIINQISFSK 286
>ref|NP_939547.1| Putative peptide transport protein [Corynebacterium diphtheriae
NCTC 13129] gi|38200041|emb|CAE49717.1| Putative peptide
transport protein [Corynebacterium diphtheriae]
Length = 498
Score = 38.1 bits (87), Expect = 1.5
Identities = 40/134 (29%), Positives = 64/134 (46%), Gaps = 22/134 (16%)
Query: 646 TFMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQI-- 703
TF V+ L+ P L TF + + + + + LLY + F ++++ LA K+ L+ I
Sbjct: 249 TFCVATGLIKPVVLATFLLALTLGAAL-------LLYTSMFRSSEVTLAEKIQLLRYIPI 301
Query: 704 -------WSLAQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSL----LW 752
W++ VL +YS+ + L+ S FL +L SL + ILSL LW
Sbjct: 302 FLASVAFWAIMNQTYGVLAVYSD-VRLNRSVGDFLIPAAWLQSLNPFYVFILSLPLAYLW 360
Query: 753 IEFG-KS*RALIKL 765
+ G +S R KL
Sbjct: 361 LRLGDRSPRPATKL 374
>gb|AAG17809.1| NADH dehydrogenase subunit 4 [Naegleria gruberi]
gi|11466208|ref|NP_066531.1| NADH dehydrogenase subunit
4 [Naegleria gruberi]
Length = 474
Score = 37.7 bits (86), Expect = 2.0
Identities = 33/134 (24%), Positives = 57/134 (41%), Gaps = 9/134 (6%)
Query: 581 FSYFHGIYYYEMY*ISLFLYSYQWQSY*SIYSPEASDKGTLFLLIYSFYVLKCYLV*FRK 640
FSY +Y+++ I F+++ W + S + P + L YSF F
Sbjct: 10 FSYNSKYFYFDISLIIFFIFNIVWVLFDS-FEPSFQFIQYINLFEYSFL--------FGI 60
Query: 641 VGIRVTFMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILI 700
GI + F+ L+P LL F M + + + L+ T FL +F A +++
Sbjct: 61 DGISIFFIYLSTFLIPLCLLFSFYNMKQKGVSEVKMYQSFLFLTLFLLIFVFSALDILVF 120
Query: 701 SQIWSLAQIPVLVL 714
++ + IP VL
Sbjct: 121 YIMFEIILIPFFVL 134
>emb|CAE66371.1| Hypothetical protein CBG11630 [Caenorhabditis briggsae]
Length = 273
Score = 37.4 bits (85), Expect = 2.6
Identities = 49/165 (29%), Positives = 73/165 (43%), Gaps = 22/165 (13%)
Query: 618 KGTLFLLIYSFYVLKCYLV*FRKVGIRVTFM--VSKLLLMPPALLTFFMQMAIFSFVRLT 675
K T FLL F V L F IR++ + +S + +P L I + L
Sbjct: 13 KKTYFLL---FLVFVDVLGIFSSTAIRISLIGTISTICFIPIFLCLHTATQVIQLLISLV 69
Query: 676 LMRPLLYPTF-FLNTKLFLARKLILISQIWSLAQIPVL---------VLKIYSELISLST 725
+R L F L K+ A+K ILI++IW L + + ++ + IS+S
Sbjct: 70 AIRKFLMHFFPSLIPKILNAQK-ILINKIWILYLLLFINDIVGYAYHIISSRNIYISVSI 128
Query: 726 SRIIFLSIWGFLLSLEDLKLKILSLLWIEFGKS*RALIKLWKEKK 770
R+ LSI+ +LL + L I SLL+ LI LW +KK
Sbjct: 129 LRVSILSIFQYLLISMNGVLLISSLLYFP------VLISLWCKKK 167
>ref|YP_023702.1| signal peptidase I [Picrophilus torridus DSM 9790]
gi|48430644|gb|AAT43509.1| signal peptidase I
[Picrophilus torridus DSM 9790]
Length = 399
Score = 37.0 bits (84), Expect = 3.4
Identities = 34/122 (27%), Positives = 60/122 (48%), Gaps = 15/122 (12%)
Query: 616 SDKGTLFLLIYSFYVLKCYLV*FRKVGIRVTFMVSKLLLMPPALLTFFMQMAIFSFVRLT 675
S KG L+ LI F +L+ + F+K I++ F+++ + P +L + + +A FSF
Sbjct: 52 SYKGILYALILIFIILEPAIA-FKK--IKLIFIINAI----PNILLYSLILA-FSF---- 99
Query: 676 LMRPLLYPTFFLNTKLFLARKLILISQIWSLAQIPVLVLKIYSELISLSTSRIIFLSIWG 735
+L+ + K R + L S I++L+ +P L SL ++FL + G
Sbjct: 100 ---SMLFLVITRSNKKAAVRNIFLSSVIFALSSVPYSAYMDVKNLFSLILFELLFLILAG 156
Query: 736 FL 737
FL
Sbjct: 157 FL 158
>gb|AAP44715.1| NADH dehydrogenase subunit 2 [lepidopsocid RS-2001]
gi|31324909|ref|NP_852163.1| NADH dehydrogenase subunit
2 [lepidopsocid RS-2001]
Length = 344
Score = 36.2 bits (82), Expect = 5.7
Identities = 37/154 (24%), Positives = 68/154 (44%), Gaps = 18/154 (11%)
Query: 596 SLFLYSYQWQSY*SIYSPEASDKGTLFLLIYSFYVLKCYLV*FRKVGIRVTFMVSKLLLM 655
S+ L Y Y IYS +F + FY+ + Y + ++K+ I + + L +
Sbjct: 195 SMILSKYILILYFLIYSFINFTLIMIFNNFFLFYINQSYFILYKKMNIFLFLSLLSLGGL 254
Query: 656 PPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSLAQIPVLVLK 715
PP LL FF + I + ++Y ++ + + LI I ++ +L+LK
Sbjct: 255 PP-LLGFFPKWLI--------IEKMMYSNLYMLNFILILTSLITIFYYLQISFSSILLLK 305
Query: 716 IYSELISLSTSRII---------FLSIWGFLLSL 740
+ +++I + II F+SI G +L L
Sbjct: 306 LENKIIMNNMKNIINSMFLNICTFISILGLVLFL 339
>emb|CAH99732.1| integral membrane protein, conserved, putative [Plasmodium berghei]
Length = 388
Score = 36.2 bits (82), Expect = 5.7
Identities = 36/145 (24%), Positives = 62/145 (41%), Gaps = 29/145 (20%)
Query: 611 YSPEASDKGTLFLLIYSFYVLKCYLV*FRKVGIRVTFMVSKLLLMPPALLTFFMQMAIFS 670
++P+ K ++F + S + Y++ KV I+ + S +++ P +LL +AI
Sbjct: 232 FTPKLLGKISMFQSLASLLAILLYMLILSKVNIKKLLLYSTIIIAPFSLLPL---LAIHK 288
Query: 671 FVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSLAQIPVLVLKIYSELISLSTSRII- 729
F+ LF+ IL+ I L IP+LV SRII
Sbjct: 289 I------------NIFIPNTLFIITDTILVEFIGELQSIPILV----------QCSRIIP 326
Query: 730 ---FLSIWGFLLSLEDLKLKILSLL 751
+I+ FLLS+ + + I S +
Sbjct: 327 EGFESTIYSFLLSVNNFAVMISSFI 351
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.368 0.167 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,391,805,012
Number of Sequences: 2540612
Number of extensions: 49075119
Number of successful extensions: 313427
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 313412
Number of HSP's gapped (non-prelim): 19
length of query: 1015
length of database: 863,360,394
effective HSP length: 138
effective length of query: 877
effective length of database: 512,755,938
effective search space: 449686957626
effective search space used: 449686957626
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC144513.4


